RNA Reference Materials Useful for Standardizing COVID-19 Tests, Finds Study
|
By LabMedica International staff writers Posted on 24 Jan 2022 |
A new study has shown that RNA reference materials are useful for standardizing COVID-19 tests.
The multiorganizational study led by researchers at the National Institute of Standards and Technology (NIST; Gaithersburg, MD, USA) looked at anchoring cycle threshold (Ct) values to a reference sample with known amounts of the virus in an effort to make the COVID-19 tests results more comparable between labs.
Scientists track and monitor the circulation of SARS-CoV-2, the virus that causes COVID-19, using methods based on a laboratory technique called polymerase chain reaction (PCR). Also used as the “gold standard” test to diagnose COVID-19 in individuals, PCR amplifies pieces of DNA by copying them numerous times through a series of chemical reactions. The number of cycles it takes to amplify DNA sequences of interest so that they are detectable by the PCR machine, known as Ct, is what researchers and medical professionals look at to detect the virus. However, not all labs get the same Ct values.
SARS-CoV-2 is an RNA virus: Its genetic material is single-stranded instead of double-stranded like DNA and contains some different molecular building blocks, namely uracil in place of thymine. But the PCR test only works with DNA, and labs first must convert the RNA to DNA to screen for COVID-19. For the test, RNA is isolated from a patient’s sample and combined with other ingredients, including short DNA sequences known as primers, to transform the RNA into DNA. When running a PCR test, labs often use the Ct value to make a positive or negative diagnosis of COVID-19. Labs often set a “cutoff” Ct value above which they interpret and can declare a patient “negative” if the virus is not detected after a certain number of cycles. But even though different labs can accurately detect the virus using their PCR tests, they use their own test methods and instruments, which could lead to different Ct values.
So, for this study, a multiorganizational research team set out to explore how much Ct values could vary among different labs when they ran PCR tests on the same reference samples containing known amounts of the SARS-CoV-2 virus. Two reference materials with carefully measured concentrations of the SARS-CoV-2 RNA were developed by a group of organizations and institutions. RM 1 had an estimated viral load of 10 million (107) copies per milliliter, and RM 2 had an estimated viral load of one million (106) copies per milliliter.
To ensure these values were accurate, metrology institutes measured and validated the reference materials using digital PCR. Digital PCR follows the same steps as traditional PCR but is a more advanced version. In digital PCR, the sample is partitioned into thousands of tiny droplets. A compound is also added so when the targeted DNA is detected it gives off a glow, allowing researchers to confirm if the sample is positive for the coronavirus with the presence of the fluorescent molecules.
The reference materials were sent out to a total of 305 laboratories in Germany, which yielded 1,109 data sets to be analyzed. The Ct values differed between labs depending on the test system or PCR equipment used and the targeted DNA sequences of the virus. For example, PCR assays aiming to detect a key gene (the “N” gene) in the COVID-19 virus had a range of Ct values between 17.6 and 26.9 for RM 1, while for RM 2 the range was between 20.7 and 30.1. The ranges between the two RMs overlap (20.7 to 26.9) even though they have different viral load concentrations, which shows it’s not possible to tell the precise concentration from Ct values alone.
The differences in Ct values among labs showed the usefulness of reference materials as a tool to help labs compare and standardize their results. Some have previously viewed the Ct value as a way to measure the viral load, the amount of virus in a person’s body. But the variation in Ct values across labs underscores that the two variables are merely correlated with one another. For example, if an individual has a higher viral load, then their Ct value would be low because it would take fewer cycles to amplify the virus’s genetic material. However, by using the appropriate reference materials, Ct values can potentially be converted to an actual virus concentration value in copies per microliter, which would be one way to quantify the viral load, though this wasn’t a focus of the study
Some have also suggested using Ct values to determine how infectious a patient is, another practice that the researchers caution against. Even though Ct values alone don’t determine how sick or infected an individual is, understanding these values could help inform decisions on setting criteria for monitoring people affected by the coronavirus. And RNA reference materials can help make these results more reliable and comparable.
“A lower Ct value means more DNA with the coronavirus starting out in the sample. It correlates to the viral load. But Ct can vary depending on the extraction material or the method used. All of these things can affect at which point the DNA is detected,” said NIST researcher Megan Cleveland, a co-author of the study. “People should use well-characterized reference materials along with their test methods instead of just relying on the Ct values. On its own, the Ct values are not easily comparable between testing laboratories, but researchers having access to reference materials and understanding their importance is beneficial.”
Related Links:
NIST
Latest COVID-19 News
- New Immunosensor Paves Way to Rapid POC Testing for COVID-19 and Emerging Infectious Diseases
- Long COVID Etiologies Found in Acute Infection Blood Samples
- Novel Device Detects COVID-19 Antibodies in Five Minutes
- CRISPR-Powered COVID-19 Test Detects SARS-CoV-2 in 30 Minutes Using Gene Scissors
- Gut Microbiome Dysbiosis Linked to COVID-19
- Novel SARS CoV-2 Rapid Antigen Test Validated for Diagnostic Accuracy
- New COVID + Flu + R.S.V. Test to Help Prepare for `Tripledemic`
- AI Takes Guesswork Out Of Lateral Flow Testing
- Fastest Ever SARS-CoV-2 Antigen Test Designed for Non-Invasive COVID-19 Testing in Any Setting
- Rapid Antigen Tests Detect Omicron, Delta SARS-CoV-2 Variants
- Health Care Professionals Showed Increased Interest in POC Technologies During Pandemic, Finds Study
- Set Up Reserve Lab Capacity Now for Faster Response to Next Pandemic, Say Researchers
- Blood Test Performed During Initial Infection Predicts Long COVID Risk
- Low-Cost COVID-19 Testing Platform Combines Sensitivity of PCR and Speed of Antigen Tests
- Finger-Prick Blood Test Identifies Immunity to COVID-19
- Quick Test Kit Determines Immunity Against COVID-19 and Its Variants
Channels
Clinical Chemistry
view channel
New PSA-Based Prognostic Model Improves Prostate Cancer Risk Assessment
Prostate cancer is the second-leading cause of cancer death among American men, and about one in eight will be diagnosed in their lifetime. Screening relies on blood levels of prostate-specific antigen... Read more
Extracellular Vesicles Linked to Heart Failure Risk in CKD Patients
Chronic kidney disease (CKD) affects more than 1 in 7 Americans and is strongly associated with cardiovascular complications, which account for more than half of deaths among people with CKD.... Read moreMolecular Diagnostics
view channel
Diagnostic Device Predicts Treatment Response for Brain Tumors Via Blood Test
Glioblastoma is one of the deadliest forms of brain cancer, largely because doctors have no reliable way to determine whether treatments are working in real time. Assessing therapeutic response currently... Read more
Blood Test Detects Early-Stage Cancers by Measuring Epigenetic Instability
Early-stage cancers are notoriously difficult to detect because molecular changes are subtle and often missed by existing screening tools. Many liquid biopsies rely on measuring absolute DNA methylation... Read more
“Lab-On-A-Disc” Device Paves Way for More Automated Liquid Biopsies
Extracellular vesicles (EVs) are tiny particles released by cells into the bloodstream that carry molecular information about a cell’s condition, including whether it is cancerous. However, EVs are highly... Read more
Blood Test Identifies Inflammatory Breast Cancer Patients at Increased Risk of Brain Metastasis
Brain metastasis is a frequent and devastating complication in patients with inflammatory breast cancer, an aggressive subtype with limited treatment options. Despite its high incidence, the biological... Read moreHematology
view channel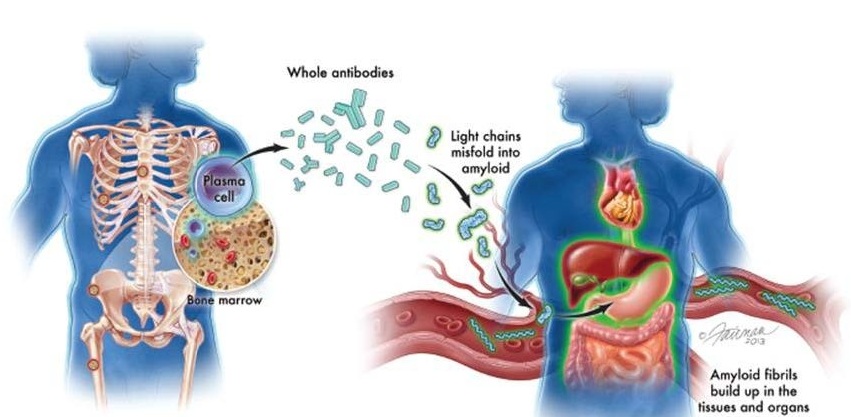
New Guidelines Aim to Improve AL Amyloidosis Diagnosis
Light chain (AL) amyloidosis is a rare, life-threatening bone marrow disorder in which abnormal amyloid proteins accumulate in organs. Approximately 3,260 people in the United States are diagnosed... Read more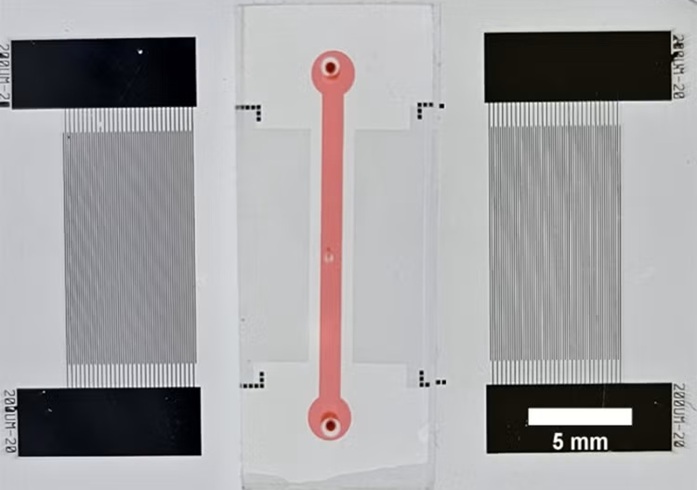
Fast and Easy Test Could Revolutionize Blood Transfusions
Blood transfusions are a cornerstone of modern medicine, yet red blood cells can deteriorate quietly while sitting in cold storage for weeks. Although blood units have a fixed expiration date, cells from... Read more
Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
High-volume hemostasis sections must sustain rapid turnaround while managing reruns and reflex testing. Manual tube handling and preanalytical checks can strain staff time and increase opportunities for error.... Read more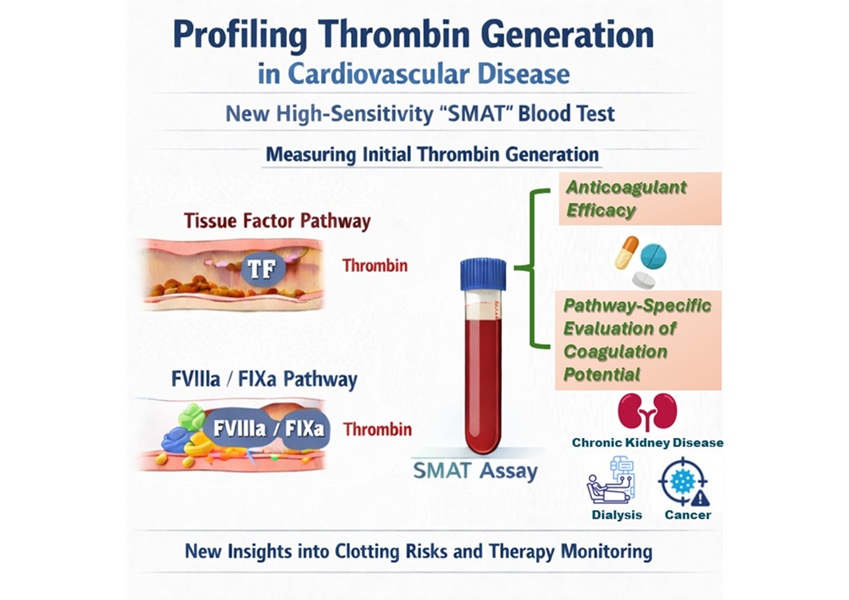
High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
Blood clotting is essential for preventing bleeding, but even small imbalances can lead to serious conditions such as thrombosis or dangerous hemorrhage. In cardiovascular disease, clinicians often struggle... Read moreImmunology
view channelBlood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
Small cell lung cancer (SCLC) is an aggressive disease with limited treatment options, and even newly approved immunotherapies do not benefit all patients. While immunotherapy can extend survival for some,... Read more
Whole-Genome Sequencing Approach Identifies Cancer Patients Benefitting From PARP-Inhibitor Treatment
Targeted cancer therapies such as PARP inhibitors can be highly effective, but only for patients whose tumors carry specific DNA repair defects. Identifying these patients accurately remains challenging,... Read more
Ultrasensitive Liquid Biopsy Demonstrates Efficacy in Predicting Immunotherapy Response
Immunotherapy has transformed cancer treatment, but only a small proportion of patients experience lasting benefit, with response rates often remaining between 10% and 20%. Clinicians currently lack reliable... Read moreMicrobiology
view channel
Comprehensive Review Identifies Gut Microbiome Signatures Associated With Alzheimer’s Disease
Alzheimer’s disease affects approximately 6.7 million people in the United States and nearly 50 million worldwide, yet early cognitive decline remains difficult to characterize. Increasing evidence suggests... Read moreAI-Powered Platform Enables Rapid Detection of Drug-Resistant C. Auris Pathogens
Infections caused by the pathogenic yeast Candida auris pose a significant threat to hospitalized patients, particularly those with weakened immune systems or those who have invasive medical devices.... Read morePathology
view channel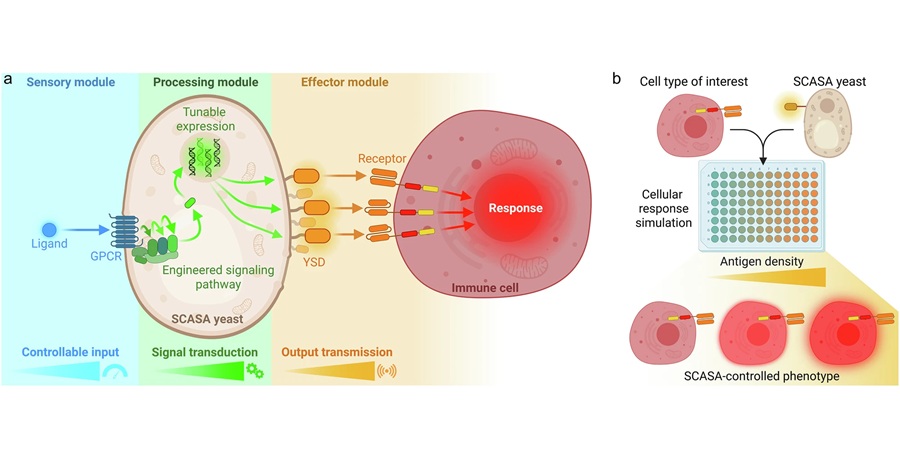
Engineered Yeast Cells Enable Rapid Testing of Cancer Immunotherapy
Developing new cancer immunotherapies is a slow, costly, and high-risk process, particularly for CAR T cell treatments that must precisely recognize cancer-specific antigens. Small differences in tumor... Read more
First-Of-Its-Kind Test Identifies Autism Risk at Birth
Autism spectrum disorder is treatable, and extensive research shows that early intervention can significantly improve cognitive, social, and behavioral outcomes. Yet in the United States, the average age... Read moreTechnology
view channel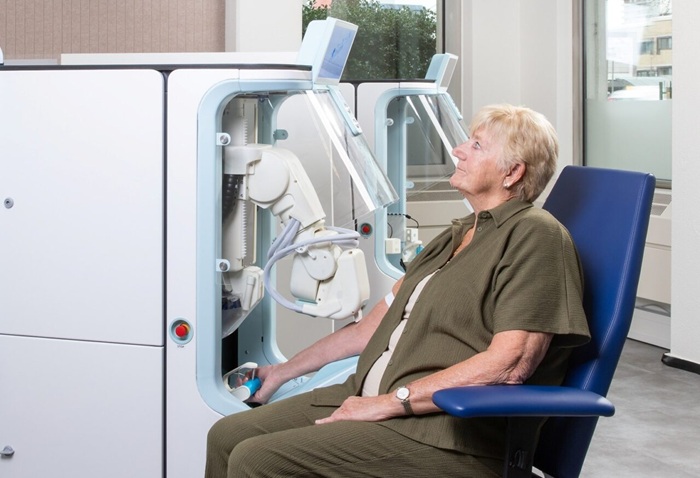
Robotic Technology Unveiled for Automated Diagnostic Blood Draws
Routine diagnostic blood collection is a high‑volume task that can strain staffing and introduce human‑dependent variability, with downstream implications for sample quality and patient experience.... Read more
ADLM Launches First-of-Its-Kind Data Science Program for Laboratory Medicine Professionals
Clinical laboratories generate billions of test results each year, creating a treasure trove of data with the potential to support more personalized testing, improve operational efficiency, and enhance patient care.... Read moreAptamer Biosensor Technology to Transform Virus Detection
Rapid and reliable virus detection is essential for controlling outbreaks, from seasonal influenza to global pandemics such as COVID-19. Conventional diagnostic methods, including cell culture, antigen... Read more
AI Models Could Predict Pre-Eclampsia and Anemia Earlier Using Routine Blood Tests
Pre-eclampsia and anemia are major contributors to maternal and child mortality worldwide, together accounting for more than half a million deaths each year and leaving millions with long-term health complications.... Read moreIndustry
view channelNew Collaboration Brings Automated Mass Spectrometry to Routine Laboratory Testing
Mass spectrometry is a powerful analytical technique that identifies and quantifies molecules based on their mass and electrical charge. Its high selectivity, sensitivity, and accuracy make it indispensable... Read more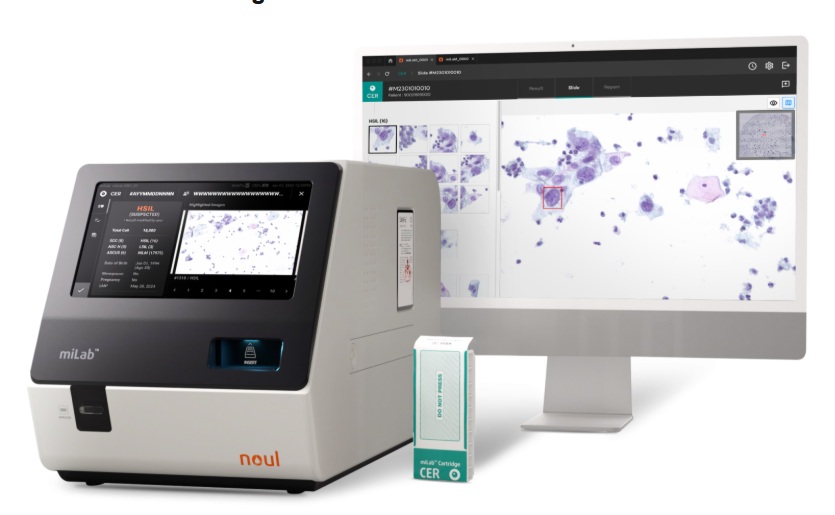
AI-Powered Cervical Cancer Test Set for Major Rollout in Latin America
Noul Co., a Korean company specializing in AI-based blood and cancer diagnostics, announced it will supply its intelligence (AI)-based miLab CER cervical cancer diagnostic solution to Mexico under a multi‑year... Read more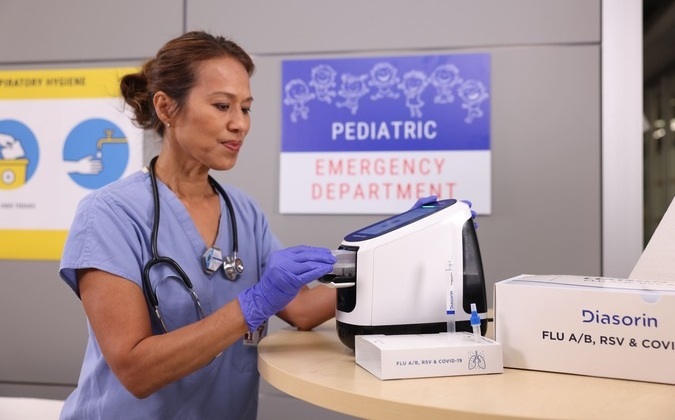
Diasorin and Fisher Scientific Enter into US Distribution Agreement for Molecular POC Platform
Diasorin (Saluggia, Italy) has entered into an exclusive distribution agreement with Fisher Scientific, part of Thermo Fisher Scientific (Waltham, MA, USA), for the LIAISON NES molecular point-of-care... Read more