Analytic Validation and Clinical Utilization of Comprehensive Genomic Profiling Test
|
By LabMedica International staff writers Posted on 02 Jun 2021 |
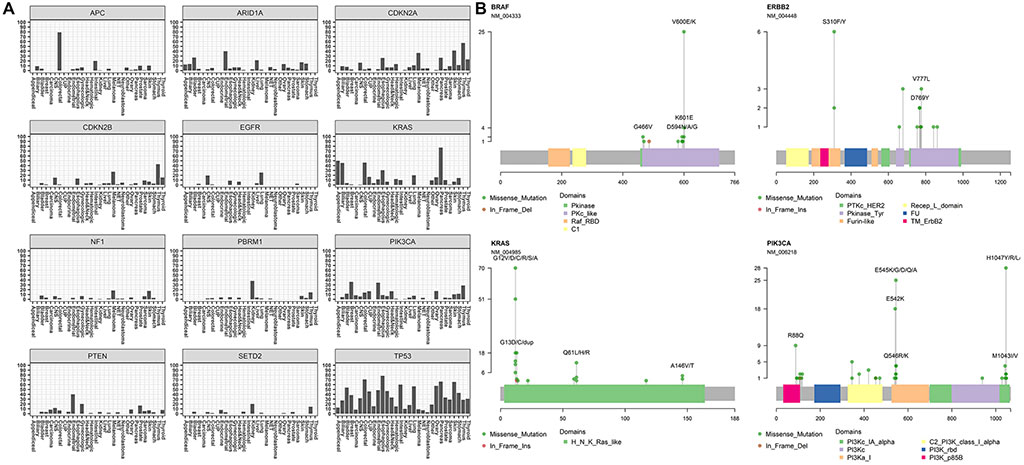
Image: Mutation profiles of clinically actionable genes in GEM ExTra. The twelve most reported genes and their mutation distribution across tumor types (Photo courtesy of Ashion Analytics)
Cancer has a high clinical burden and oncology therapies are expensive. It is estimated that 1,898,160 new cancer cases will be diagnosed and over 608,570 deaths are projected to occur in the USA in 2021. The prevalence of cancer is expected to rise over time, providing an expanding unmet need for genomic tests to help physicians treat patients in a more precise manner.
Identification of genomic alterations by Next Generation Sequencing (NGS) has become an efficient clinical tool, particularly for oncology as molecular markers can guide personalized treatment. Currently available options for tumor profiling include immunohistochemistry (IHC), fluorescence in situ hybridization (FISH), and, more recently, small panel next-generation sequencing (NGS).
Scientists at the Ashion Analytics, LLC, (Phoenix, AZ, USA) developed and analytically validated their comprehensive genomic profiling assay, GEM ExTra. The assay is for patients with advanced solid tumors that and uses Next Generation Sequencing to characterize whole exomes employing a paired tumor-normal subtraction methodology. The test was utilized in over 1,400 patient samples during a period of April 2018 and December 2019 across cancer centers to detect multiple actionable alterations in a variety of cancer types.
The assay detects single nucleotide variants (SNV), indels, focal copy number alterations (CNA), TERT promoter region, as well as tumor mutation burden (TMB) and microsatellite instability (MSI) status. Additionally, the assay incorporates whole transcriptome sequencing of the tumor sample that allows for the detection of gene fusions and selects special transcripts, including AR-V7, EGFR vIII, EGFRvIV, and MET exon 14 skipping events. The assay has a mean target coverage of 180× for the normal (germline) and 400× for tumor DNA including enhanced probe design to facilitate the sequencing of difficult regions.
The team found that found that 83.9% of tumor samples harbored at least one clinically actionable alteration (defined as positive) and the rest defined as negative, with a total of 1,267 positive and 242 negative reports (Detection Rate: 2018 = 76.4%, 2019 = 86.4%). Overall, 3,535 clinically actionable mutations were identified in the cohort (1,864 unique mutations), with a median of two clinically actionable alterations per tumor (mean = 2.93 ± 2.37) showing extensive variation across cancer types. Tumors with highest number of actionable mutations included skin (4.9 ± 2.2), endometrial (4.5 ± 3.9), and colorectal (4.1 ± 3.5). These results generally agree with previous estimates of driver events per patient in these tumor types. This is somewhat lower than previously reported in a pan-cancer study (4.6/tumor) of whole genomes, which also included driver copy number alterations which are not called out as actionable with GEM ExTra. Mean coding SNVs (i.e., missense, nonsense, stop codon) was 1.9 ± 1.4 per tumor which is within the range of predicted driver mutations in cancer.
The scientists concluded that they had developed and analytically validated a comprehensive genomic profiling assay with a 14-day turnaround time that can be adapted to all future tumor profiling needs due to combined DNA and RNA analysis. The GEM ExTra assay not only uses WES for tumor DNA profiling, but also identifies clinically actionable transcript variants and fusion genes through RNA sequencing, both of which ensure that GEM ExTra will be comprehensive in the future. The study was published on April 13, 2021 in the journal Oncotarget.
Related Links:
Ashion Analytics
Identification of genomic alterations by Next Generation Sequencing (NGS) has become an efficient clinical tool, particularly for oncology as molecular markers can guide personalized treatment. Currently available options for tumor profiling include immunohistochemistry (IHC), fluorescence in situ hybridization (FISH), and, more recently, small panel next-generation sequencing (NGS).
Scientists at the Ashion Analytics, LLC, (Phoenix, AZ, USA) developed and analytically validated their comprehensive genomic profiling assay, GEM ExTra. The assay is for patients with advanced solid tumors that and uses Next Generation Sequencing to characterize whole exomes employing a paired tumor-normal subtraction methodology. The test was utilized in over 1,400 patient samples during a period of April 2018 and December 2019 across cancer centers to detect multiple actionable alterations in a variety of cancer types.
The assay detects single nucleotide variants (SNV), indels, focal copy number alterations (CNA), TERT promoter region, as well as tumor mutation burden (TMB) and microsatellite instability (MSI) status. Additionally, the assay incorporates whole transcriptome sequencing of the tumor sample that allows for the detection of gene fusions and selects special transcripts, including AR-V7, EGFR vIII, EGFRvIV, and MET exon 14 skipping events. The assay has a mean target coverage of 180× for the normal (germline) and 400× for tumor DNA including enhanced probe design to facilitate the sequencing of difficult regions.
The team found that found that 83.9% of tumor samples harbored at least one clinically actionable alteration (defined as positive) and the rest defined as negative, with a total of 1,267 positive and 242 negative reports (Detection Rate: 2018 = 76.4%, 2019 = 86.4%). Overall, 3,535 clinically actionable mutations were identified in the cohort (1,864 unique mutations), with a median of two clinically actionable alterations per tumor (mean = 2.93 ± 2.37) showing extensive variation across cancer types. Tumors with highest number of actionable mutations included skin (4.9 ± 2.2), endometrial (4.5 ± 3.9), and colorectal (4.1 ± 3.5). These results generally agree with previous estimates of driver events per patient in these tumor types. This is somewhat lower than previously reported in a pan-cancer study (4.6/tumor) of whole genomes, which also included driver copy number alterations which are not called out as actionable with GEM ExTra. Mean coding SNVs (i.e., missense, nonsense, stop codon) was 1.9 ± 1.4 per tumor which is within the range of predicted driver mutations in cancer.
The scientists concluded that they had developed and analytically validated a comprehensive genomic profiling assay with a 14-day turnaround time that can be adapted to all future tumor profiling needs due to combined DNA and RNA analysis. The GEM ExTra assay not only uses WES for tumor DNA profiling, but also identifies clinically actionable transcript variants and fusion genes through RNA sequencing, both of which ensure that GEM ExTra will be comprehensive in the future. The study was published on April 13, 2021 in the journal Oncotarget.
Related Links:
Ashion Analytics
Latest Molecular Diagnostics News
- Novel Algorithm Improves Detection of B-ALL Gene Fusions
- Rapid Multiplex PCR Test Detects 11 Gastrointestinal Pathogens from Single Sample
- Sensitive Protein Marker Aids Diagnosis of Small Cell Prostate Cancer
- Gene Panel Shows Promise for Predicting Chemotherapy Response in TNBC
- Realistic Mock Samples Aim to Speed Cervical Cancer Test Development
- Blood-Based “Ageing Clock” Helps Predict Dementia Risk and Earlier Onset
- Molecular Marker Identifies Hormone Therapy Resistance Pathway in Prostate Cancer
- Blood Test Refines Biopsy Decisions in Prostate Cancer
- Digital Aging Twin Quantifies Biological Aging Across Multiple Organ Systems
- Emerging Biomarkers Advance Early Detection of MASLD and Liver Cancer Risk
- Urine Test Beats MRI in Identifying Prostate Cancer Upgrading During Active Surveillance
- Finger-Prick Blood Test Aids Early Tuberculosis Detection and Risk Stratification
- Patented Isothermal Amplification Chemistry Advances Decentralized Testing
- Direct-to-Patient Genetic Testing Identifies Hereditary Cancer Risk in Survivors
- Stool DNA Analysis Tracks Disease Activity in IBD
- Blood Test Maps Tumor Microenvironment to Predict Immunotherapy Response
Channels
Clinical Chemistry
view channel
Urine-Based Nanosensor Tracks Lung Cancer and Fibrosis Noninvasively
Lung cancer remains difficult to monitor for early progression and treatment resistance, while pulmonary fibrosis continues to pose major challenges for early diagnosis. Clinicians need repeatable, noninvasive... Read more
Blood-Based Alzheimer’s Test Gains CE Mark for Amyloid Pathology Detection
Alzheimer’s disease is the most common cause of dementia, yet confirmatory testing remains invasive and hard to access. Diagnosis currently takes an average of 3.5 years, and about 75% of people with dementia... Read moreHematology
view channel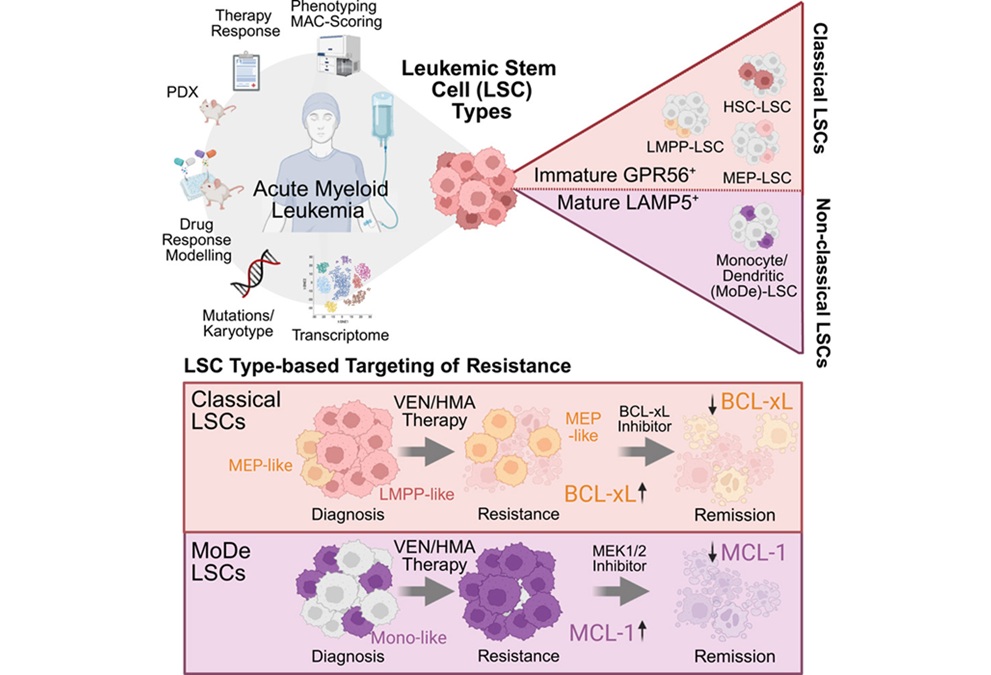
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read more
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read moreImmunology
view channel
Routine TB Screening Test May Reveal Immune Aging and Mortality Risk
Immune aging is associated with weaker responses to vaccination, greater risks of infection, and higher levels of inflammation. Leveraging routinely ordered laboratory tests to quantify that responsiveness... Read more
Biomarkers and Molecular Testing Advance Precision Allergy Care
Allergic diseases often present with similar symptoms but can be driven by distinct biological mechanisms, making standardized care inefficient for many patients. Historically, individuals with pollen... Read moreMicrobiology
view channel
Study Finds Hidden Mpox Infections May Drive Ongoing Spread
Mpox continues to circulate despite vaccination, and many cases show no known link to a symptomatic partner. The role of people without symptoms has remained uncertain, limiting clarity on how transmission persists.... Read more
Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
Antimicrobial resistance (AMR) poses a growing threat to patient safety, with carbapenem-resistant Enterobacterales causing difficult-to-treat infections and leaving clinicians with limited therapeutic options.... Read more
Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
Tuberculosis is the leading cause of death among people living with HIV, and diagnosis in hospital settings remains difficult. Symptoms are often non-specific, disease can be extrapulmonary, and many patients... Read morePathology
view channel
FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
Risk assessment at diagnosis is central to guiding therapy for early-stage, hormone receptor-positive, human epidermal growth factor receptor 2-negative (HR+/HER2-) invasive breast cancer, where overtreatment... Read more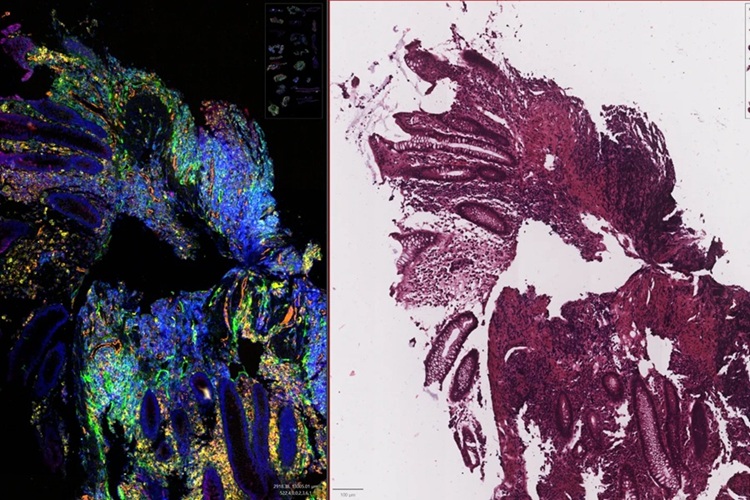
New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
Pathologists worldwide rely on hematoxylin and eosin (H&E) slides to examine tissue architecture, yet these stains do not reveal the underlying molecular activity that often drives disease.... Read moreTechnology
view channel
Point-of-Care Testing Enhances Health Literacy and Self-Management in Chronic Disease
Limited access to general practitioners and pathology services can delay diagnosis and monitoring for people in regional and remote communities. Rapid, on-the-spot testing can shorten turnaround times... Read more
Fully Automated Sample-to-Insight Workflow Advances Latent TB Testing
Latent tuberculosis remains a substantial testing workload for clinical laboratories as screening programs expand. Despite this growth, only about 40% of testing has shifted from traditional skin tests... Read moreIndustry
view channel
AI-Powered Multi-Functional Analyzer Wins German Innovation Award
Hematology services are increasingly delivered across distributed care settings, where limited staffing and complex workflows can extend turnaround times. Advanced morphology review still often depends... Read more