Combined Gene Editing and RNA Sequencing Enhances Understanding of Gene Expression
|
By LabMedica International staff writers Posted on 27 Dec 2016 |

Image: Combining CRISPR with the fine resolution of single-cell RNA sequencing gives researchers new means of controlling cell activities (Photo courtesy of the Weizmann Institute of Science).
A team of Israeli molecular geneticists has combined two powerful genomics tools to study single cell gene expression.
Investigators at the Weizmann Institute of Science (Rehovot, Israel) used CRISPR/Cas9 gene editing to introduce changes into the genome and then profiled the genomic perturbation and transcriptome with single-cell RNA sequencing (RNA-Seq).
CRISPRs (clustered regularly interspaced short palindromic repeats) are segments of prokaryotic DNA containing short repetitions of base sequences. Each repetition is followed by short segments of "spacer DNA" from previous exposures to a bacterial virus or plasmid. CRISPRs are found in approximately 40% of sequenced bacteria genomes and 90% of sequenced archaea. CRISPRs are often associated with cas genes that code for proteins related to CRISPRs. Since 2013, the CRISPR/Cas system has been used in research for gene editing (adding, disrupting, or changing the sequence of specific genes) and gene regulation. By delivering the Cas9 enzyme and appropriate guide RNAs (sgRNAs) into a cell, the organism's genome can be cut at any desired location.
The conventional CRISPR/Cas9 system is composed of two parts: the Cas9 enzyme, which cleaves the DNA molecule and specific RNA guides that shepherd the Cas9 protein to the target gene on a DNA strand. Despite its attractiveness as a gene-editing tool, the technique can inadvertently make excessive or unwanted changes in the genome and create off-target mutations, limiting safety and efficacy in therapeutic applications.
RNA-Seq is used to analyze the continually changing cellular transcriptome. Specifically, RNA-Seq facilitates the ability to look at alternative gene spliced transcripts, post-transcriptional modifications, gene fusion, mutations/SNPs, and changes in gene expression. In addition to mRNA transcripts, RNA-Seq can look at different populations of RNA to include total RNA, small RNA, such as miRNA, tRNA, and ribosomal profiling. RNA-Seq can also be used to determine exon/intron boundaries and verify or amend previously annotated 5' and 3' gene boundaries.
The investigators reported in the December 15, 2016, online edition of the journal Cell that they had applied combined CRISP-Seq methodology to probe regulatory circuits of innate immunity. By sampling tens of thousands of perturbed cells in vitro and in mice, they identified interactions and redundancies between developmental and signaling-dependent factors. These included the opposing effects of Cebpb (CCAAT/enhancer-binding protein beta) and Irf8 (Interferon regulatory factor 8) in regulating the monocyte/macrophage versus dendritic cell lineages and differential functions for Rela (Transcription factor p65) and Stat1/2 (Signal transducer and activator of transcription 1/2) in monocyte versus dendritic cell responses to pathogens.
"The advent of CRISPR presented a true leap in the ability to understand and start editing immune circuits, but CRISPR, on its own, is a blunt research tool, since we often have trouble observing or understanding the outcome of this genomic editing," said senior author Dr. Ido Amit, professor of immunology at the Weizmann Institute of Science. "We are hoping that our approach will be the next leap forward, providing, among other things, the ability to engineer immune cells for immunotherapy."
Related Links:
Weizmann Institute of Science
Investigators at the Weizmann Institute of Science (Rehovot, Israel) used CRISPR/Cas9 gene editing to introduce changes into the genome and then profiled the genomic perturbation and transcriptome with single-cell RNA sequencing (RNA-Seq).
CRISPRs (clustered regularly interspaced short palindromic repeats) are segments of prokaryotic DNA containing short repetitions of base sequences. Each repetition is followed by short segments of "spacer DNA" from previous exposures to a bacterial virus or plasmid. CRISPRs are found in approximately 40% of sequenced bacteria genomes and 90% of sequenced archaea. CRISPRs are often associated with cas genes that code for proteins related to CRISPRs. Since 2013, the CRISPR/Cas system has been used in research for gene editing (adding, disrupting, or changing the sequence of specific genes) and gene regulation. By delivering the Cas9 enzyme and appropriate guide RNAs (sgRNAs) into a cell, the organism's genome can be cut at any desired location.
The conventional CRISPR/Cas9 system is composed of two parts: the Cas9 enzyme, which cleaves the DNA molecule and specific RNA guides that shepherd the Cas9 protein to the target gene on a DNA strand. Despite its attractiveness as a gene-editing tool, the technique can inadvertently make excessive or unwanted changes in the genome and create off-target mutations, limiting safety and efficacy in therapeutic applications.
RNA-Seq is used to analyze the continually changing cellular transcriptome. Specifically, RNA-Seq facilitates the ability to look at alternative gene spliced transcripts, post-transcriptional modifications, gene fusion, mutations/SNPs, and changes in gene expression. In addition to mRNA transcripts, RNA-Seq can look at different populations of RNA to include total RNA, small RNA, such as miRNA, tRNA, and ribosomal profiling. RNA-Seq can also be used to determine exon/intron boundaries and verify or amend previously annotated 5' and 3' gene boundaries.
The investigators reported in the December 15, 2016, online edition of the journal Cell that they had applied combined CRISP-Seq methodology to probe regulatory circuits of innate immunity. By sampling tens of thousands of perturbed cells in vitro and in mice, they identified interactions and redundancies between developmental and signaling-dependent factors. These included the opposing effects of Cebpb (CCAAT/enhancer-binding protein beta) and Irf8 (Interferon regulatory factor 8) in regulating the monocyte/macrophage versus dendritic cell lineages and differential functions for Rela (Transcription factor p65) and Stat1/2 (Signal transducer and activator of transcription 1/2) in monocyte versus dendritic cell responses to pathogens.
"The advent of CRISPR presented a true leap in the ability to understand and start editing immune circuits, but CRISPR, on its own, is a blunt research tool, since we often have trouble observing or understanding the outcome of this genomic editing," said senior author Dr. Ido Amit, professor of immunology at the Weizmann Institute of Science. "We are hoping that our approach will be the next leap forward, providing, among other things, the ability to engineer immune cells for immunotherapy."
Related Links:
Weizmann Institute of Science
Latest BioResearch News
- Study Identifies Protein Changes Driving Immunotherapy Resistance in Multiple Myeloma
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- Innate Immunity Variants Associated With Earlier Breast Cancer in BRCA1 Carriers
- Genetic Cause Identified for Severe Infant Epilepsy
- Study Reveals Diagnostic and Therapeutic Target in Rare Pancreatic Tumors
- Researchers Identify Survival Pathway Undermining Targeted Cancer Drugs
- Large-Scale Study Maps DNA Damage Signatures Across Multiple Cancers
- Study Identifies Distinct Immune Signatures to Early Depression and Psychosis
- Genetic Mutation Behind Aggressive Adult Leukemia Offers Treatment Clues
- Disease Gene Discovery Advances Diagnosis of Rare Movement Disorders
Channels
Clinical Chemistry
view channel
Blood Test Predicts Alzheimer Disease Risk Before Imaging Changes and Symptoms
Alzheimer's disease often advances silently for years, making timely risk stratification difficult in routine practice. Current approaches to detect pathology can involve lumbar puncture or positron emission... Read more
Study Finds ApoB Testing More Effective Than LDL for Guiding Lipid Therapy
Routine blood tests that measure low-density lipoprotein (LDL), commonly known as “bad” cholesterol, are widely used to guide lipid-lowering therapy, but they do not always provide a complete picture of... Read more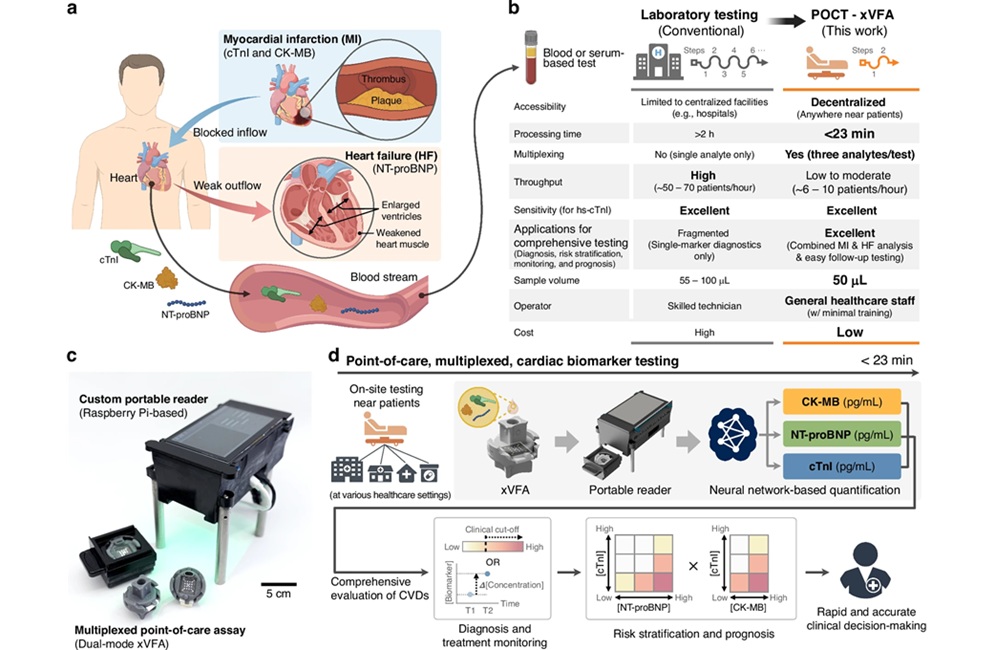
AI-Enabled POC Test Quantifies Multiple Cardiac Biomarkers
Cardiovascular diseases are a leading cause of death, responsible for nearly 20 million deaths each year. Timely triage of myocardial infarction and heart failure hinges on rapid cardiac biomarker measurement,... Read moreNext Generation Automated Analyzers Increase Throughput for Clinical Chemistry and Electrolyte Testing
Clinical laboratories continue to face staffing shortages, limited space, and growing test volumes that pressure chemistry and electrolyte workflows. Maintaining rapid turnaround times increasingly depends... Read moreMolecular Diagnostics
view channel
Blood-Based Epigenetic Signals Enable Osteosarcoma Disease Monitoring
Osteosarcoma is a rare but aggressive pediatric bone cancer where recurrence and metastasis remain difficult to detect early. Imaging-based surveillance can miss small lesions and exposes children to repeated... Read more
Host–Virus Genetic Interactions Drive Nasopharyngeal Cancer Risk
Epstein–Barr virus (EBV) infects more than 95% of adults worldwide, yet only a small fraction develops EBV‑associated cancers such as nasopharyngeal carcinoma. Explaining this divergence requires understanding... Read moreHematology
view channel
Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
Anemia detected in routine care can signal underlying pathology and is frequently encountered in adults. Because it is defined by hemoglobin levels below the normal range, it is often evaluated with red... Read more
Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
Chronic myelomonocytic leukemia (CMML) is a rare blood cancer in which acquired genetic mutations in bone marrow stem cells drive disease. Stem cell transplantation is the only curative option but carries... Read moreImmunology
view channel
Study Finds Influenza Often Undiagnosed in Winter Deaths
Seasonal influenza drives substantial excess mortality, yet its contribution is often obscured when infections go undiagnosed near the time of death. Many deaths occur outside hospitals or in older adults... Read moreCombined Screening Approach Identifies Early Leprosy Cases
Leprosy remains a significant public health concern, with more than 200,000 new cases reported globally each year and early disease often escaping routine laboratory detection. In its initial phase, bacterial... Read moreMicrobiology
view channel
Rapid Color Test Stratifies Virulent and Resistant Staph Strains
Staphylococcus aureus (golden staph) remains a leading cause of infection-related mortality worldwide, responsible for more than a million deaths each year. Rapidly distinguishing highly virulent or a... Read more
Syndromic Panel Enables Rapid Identification of Bloodstream Infections
Bloodstream infections require rapid identification of causative pathogens and resistance determinants to guide therapy, yet laboratories often face pressure to deliver clinically relevant results quickly... Read more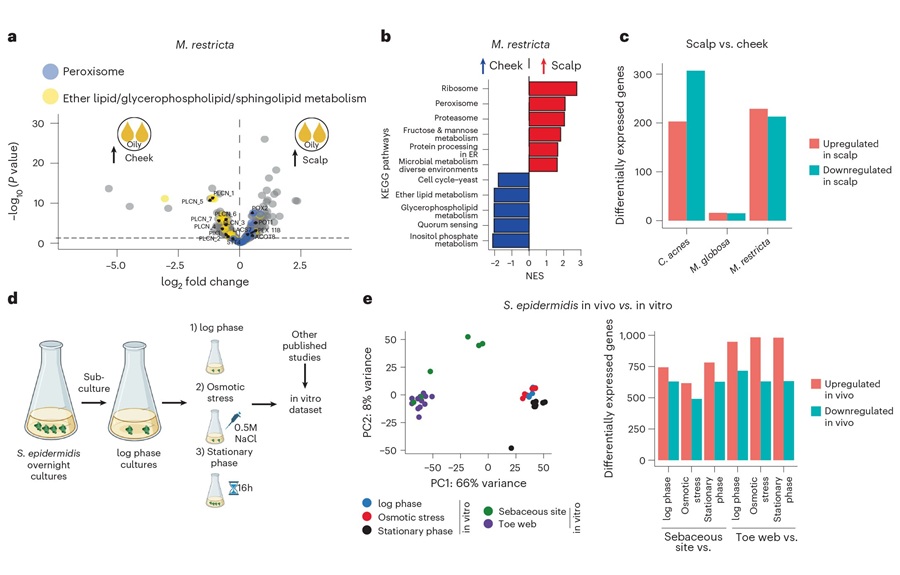
RNA-Based Workflow Identifies Active Skin Microbes for Dermatology Research
Human skin carries diverse microbial communities that influence barrier function and inflammation, yet identifying which organisms are metabolically active has been challenging. DNA-based surveys catalog... Read more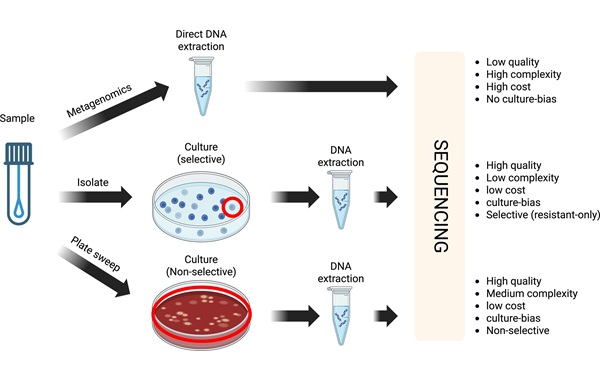
Cost-Effective Sampling and Sequencing Workflow Identifies ICU Infection Hotspots
Intensive care units face persistent threats from hospital-acquired infections, increasingly driven by drug-resistant bacteria. Rapidly pinpointing environmental reservoirs and transmission hotspots remains... Read morePathology
view channel
Biomarker Predicts Immunotherapy Response and Prognosis in Colorectal Cancer
Colorectal cancer is common and often lethal, and therapeutic decision-making is complicated by heterogeneous tumor microenvironments. Immunotherapy benefits only a small subset of patients, around 5%,... Read more
Collaboration Applies AI Pathology to Predict Response to Antibody-Drug Conjugates
Antibody-drug conjugates (ADC) are reshaping oncology, yet scalable biomarkers that reliably predict which patients will benefit remain limited as treatment regimens and combinations grow more complex.... Read moreTechnology
view channel
AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
Advanced bowel cancer remains difficult to treat, and many patients receive targeted therapies that do not help them but still cause harm. Clinicians need reliable ways to identify likely responders before... Read more
Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
Pre-analytical variation remains a leading source of inconsistent molecular test results and added costs, particularly when laboratories rely on multiple instruments and protocols. Standardizing nucleic... Read moreIndustry
view channel
Partnership Expands Ultrasensitive WGS Assay for for Hematologic Malignancies and MRD Monitoring
Tempus AI and Predicta Biosciences announced the commercial expansion of a co-branded whole‑genome sequencing assay GenoPredicta, which is intended for comprehensive genomic characterization of hematologic... Read more