First Molecule Created to Suppress Major Component of Cancer Gene On-Off Switch
|
By LabMedica International staff writers Posted on 21 Oct 2010 |
In the endeavor to block the growth and spread of tumors, there have been many attempts to get cancer genes to ignore their internal regulatory instructions. In a new study, a team of scientists has created the first molecule able to prevent cancer genes from "hearing” those instructions, inhibiting the cancer process at its foundation.
The study, published online in late September 2010 by the journal Nature, demonstrated that proteins delivering stop and start instructions to a cancer gene--known as epigenetic "reader” proteins--can be targeted for future cancer therapies. The research is particularly pertinent to a rare but destructive cancer of children and young adults known as NUT midline carcinoma (NMC)--a disease so inflexible that no potential therapy for it has ever reached the stage of being evaluated in a clinical trial.
"In recent years, it has become clear that being able to control gene activity in cancer-- manipulating which genes are ‘on' or ‘off''--can be a high-impact approach to the disease,” said the study's senior author, James Bradner, M.D., of Dana-Farber Cancer Institute (Boston, MA, USA). "If you can switch off a cancer cell's growth genes, the cell will die. Alternatively, switching on a tissue gene can cause a cancer cell to become a more normal tissue cell.”
In this study, Dr. Bradner's lab synthesized a molecule that has both effects: by blocking a specific abnormal protein in NUT midline carcinoma cells, it stops them from dividing so prolifically and makes them ‘forget' they are cancer cells and start appearing more like normal cells. The assembled molecule affects the cell's multilayered machinery for controlling gene activity--the set of structures collectively known as the epigenome. Large portions of each gene play a regulatory role, dictating whether the gene is active, industriously sending orders for new proteins, or inactive, and temporarily at rest.
The gene's DNA is packaged in a substance called chromatin, which is the slate on which instructions to begin or cease activity are inscribed. The instructions themselves take the form of "bookmarks,” material placed on the chromatin by so-called epigenetic "writer” proteins. Another group of epigenetic proteins, known as "erasers,” is able to remove the bookmarks. Both types of proteins have effectively been disabled by researchers, using molecules generated in the laboratory or taken from nature. Their success has triggered intense interest in the development of anticancer therapies that work by blocking such proteins.
A third kind of epigenetic proteins--potentially the most appealing as therapeutic targets, because they switch genes on or off by "reading” the bookmarks--has received scant scientific attention. Dr. Bradner and his colleagues looked to this little-studied area of biology by focusing on NMC cells.
The disease is caused by a chromosomal translocation, in which two genes from different chromosomes become connected and give rise to an abnormal, fused protein known as BRD4-NUT. A review of the scientific literature suggested that some members of the benzodiazepine family of drugs, which includes Valium, Xanax, and Ativan, are active against "bromodomain” proteins such as BRD4. With that as a clue, Dr. Bradner and his Dana-Farber colleague Jun Qi, Ph.D., created an array of molecules to see if any inhibited a "reader” protein of the BRD4-NUT gene. One did, quite persuasively--a hybrid molecule, which researchers named JQ1, for Qi.
The investigators worked with researchers in the United States and overseas to learn more about the properties of JQ1 and how it works in cells. Stefan Knapp, Ph.D., of Oxford University (U.K.), provided crystal-clear images of the molecule bound to a protein; Olaf Wiest, Ph.D., of the University of Notre Dame (West Bend, IN, USA), showed that the molecule is less flexible in the presence of a protein, clarifying why it so effectively blocks the protein; and Andrew Kung, M.D., Ph.D., of Dana-Farber, modified animal models in which the molecule could be tested against NMC tumors.
The animal studies were especially promising. Investigators transplanted NMC cells from patients into laboratory mice, which were then given the JQ1 molecule. "The activity of the molecule was remarkable,” noted Dr. Bradner. "All the mice that received JQ1 lived; all that did not, died.”
For now, JQ1's primary utility is as a probe for better understanding the biology underlying NUT midline carcinoma. Drs. Bradner, Qi and their colleagues are customizing the molecule to maximize its effectiveness as a BRD4-NUT stopper. Ultimately, it, or a similar molecule, could be the basis for the first effective therapy against NMC.
"The disease tends to arise in the chest, head, or neck, along the vertical centerline of the body, with aggressive tumor growth and metastasis,” Dr. Bradner explained. "Patients may have a brief response to chemotherapy, but they eventually succumb to the spread of the disease.”
Unlike most cancers, NMC's tissue of origin is not known. It is a disease defined entirely by its genetic signature--the presence of the translocated gene BRD4-NUT. Prior to its genetic identification by Christopher French, M.D., of Brigham and Women's Hospital (Boston, MA, USA) and a study coauthor, NMC was not recognized as a definitive disease.
"This research further illustrates the promise of personalized medicine,” Dr. Bradner remarked, "which is the ability to deliver selected molecules to cancer-causing proteins to stop the cancer process while producing a minimum of residual side effects. The development of JQ1 or similar molecule into a drug may produce the first therapy specifically designed for patients with NMC.”
Related Links:
Dana-Farber Cancer Institute
Brigham and Women's Hospital
The study, published online in late September 2010 by the journal Nature, demonstrated that proteins delivering stop and start instructions to a cancer gene--known as epigenetic "reader” proteins--can be targeted for future cancer therapies. The research is particularly pertinent to a rare but destructive cancer of children and young adults known as NUT midline carcinoma (NMC)--a disease so inflexible that no potential therapy for it has ever reached the stage of being evaluated in a clinical trial.
"In recent years, it has become clear that being able to control gene activity in cancer-- manipulating which genes are ‘on' or ‘off''--can be a high-impact approach to the disease,” said the study's senior author, James Bradner, M.D., of Dana-Farber Cancer Institute (Boston, MA, USA). "If you can switch off a cancer cell's growth genes, the cell will die. Alternatively, switching on a tissue gene can cause a cancer cell to become a more normal tissue cell.”
In this study, Dr. Bradner's lab synthesized a molecule that has both effects: by blocking a specific abnormal protein in NUT midline carcinoma cells, it stops them from dividing so prolifically and makes them ‘forget' they are cancer cells and start appearing more like normal cells. The assembled molecule affects the cell's multilayered machinery for controlling gene activity--the set of structures collectively known as the epigenome. Large portions of each gene play a regulatory role, dictating whether the gene is active, industriously sending orders for new proteins, or inactive, and temporarily at rest.
The gene's DNA is packaged in a substance called chromatin, which is the slate on which instructions to begin or cease activity are inscribed. The instructions themselves take the form of "bookmarks,” material placed on the chromatin by so-called epigenetic "writer” proteins. Another group of epigenetic proteins, known as "erasers,” is able to remove the bookmarks. Both types of proteins have effectively been disabled by researchers, using molecules generated in the laboratory or taken from nature. Their success has triggered intense interest in the development of anticancer therapies that work by blocking such proteins.
A third kind of epigenetic proteins--potentially the most appealing as therapeutic targets, because they switch genes on or off by "reading” the bookmarks--has received scant scientific attention. Dr. Bradner and his colleagues looked to this little-studied area of biology by focusing on NMC cells.
The disease is caused by a chromosomal translocation, in which two genes from different chromosomes become connected and give rise to an abnormal, fused protein known as BRD4-NUT. A review of the scientific literature suggested that some members of the benzodiazepine family of drugs, which includes Valium, Xanax, and Ativan, are active against "bromodomain” proteins such as BRD4. With that as a clue, Dr. Bradner and his Dana-Farber colleague Jun Qi, Ph.D., created an array of molecules to see if any inhibited a "reader” protein of the BRD4-NUT gene. One did, quite persuasively--a hybrid molecule, which researchers named JQ1, for Qi.
The investigators worked with researchers in the United States and overseas to learn more about the properties of JQ1 and how it works in cells. Stefan Knapp, Ph.D., of Oxford University (U.K.), provided crystal-clear images of the molecule bound to a protein; Olaf Wiest, Ph.D., of the University of Notre Dame (West Bend, IN, USA), showed that the molecule is less flexible in the presence of a protein, clarifying why it so effectively blocks the protein; and Andrew Kung, M.D., Ph.D., of Dana-Farber, modified animal models in which the molecule could be tested against NMC tumors.
The animal studies were especially promising. Investigators transplanted NMC cells from patients into laboratory mice, which were then given the JQ1 molecule. "The activity of the molecule was remarkable,” noted Dr. Bradner. "All the mice that received JQ1 lived; all that did not, died.”
For now, JQ1's primary utility is as a probe for better understanding the biology underlying NUT midline carcinoma. Drs. Bradner, Qi and their colleagues are customizing the molecule to maximize its effectiveness as a BRD4-NUT stopper. Ultimately, it, or a similar molecule, could be the basis for the first effective therapy against NMC.
"The disease tends to arise in the chest, head, or neck, along the vertical centerline of the body, with aggressive tumor growth and metastasis,” Dr. Bradner explained. "Patients may have a brief response to chemotherapy, but they eventually succumb to the spread of the disease.”
Unlike most cancers, NMC's tissue of origin is not known. It is a disease defined entirely by its genetic signature--the presence of the translocated gene BRD4-NUT. Prior to its genetic identification by Christopher French, M.D., of Brigham and Women's Hospital (Boston, MA, USA) and a study coauthor, NMC was not recognized as a definitive disease.
"This research further illustrates the promise of personalized medicine,” Dr. Bradner remarked, "which is the ability to deliver selected molecules to cancer-causing proteins to stop the cancer process while producing a minimum of residual side effects. The development of JQ1 or similar molecule into a drug may produce the first therapy specifically designed for patients with NMC.”
Related Links:
Dana-Farber Cancer Institute
Brigham and Women's Hospital
Latest BioResearch News
- Lung Cancer Study Reveals Cellular Program Behind Therapy Resistance
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
Channels
Clinical Chemistry
view channel
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read more
Urine-Based Test Shows Promise for Autism Screening in Children
Autism spectrum disorder (ASD) is commonly diagnosed through behavioral assessments, which can involve long waits that delay intervention. Earlier identification is linked to better developmental outcomes,... Read moreMolecular Diagnostics
view channel
Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
Sarcomas remain difficult to monitor during immunotherapy, as low tumor mutation burden can limit traditional circulating tumor DNA approaches and repeat tissue biopsies are often impractical in advanced disease.... Read more
Tumor Mutation Marker Helps Refine Lung Cancer Prognosis and Guide Therapy Selection
Lung cancer remains the leading cause of cancer death worldwide, while heterogeneous tumor genetics continue to complicate treatment decisions. Although molecular testing is increasingly used to match... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more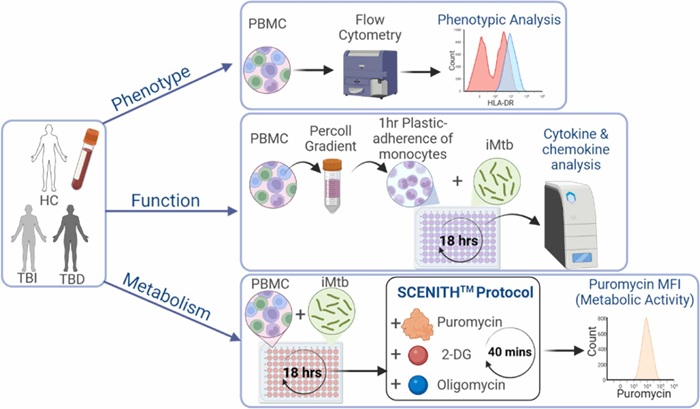
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read moreMicrobiology
view channel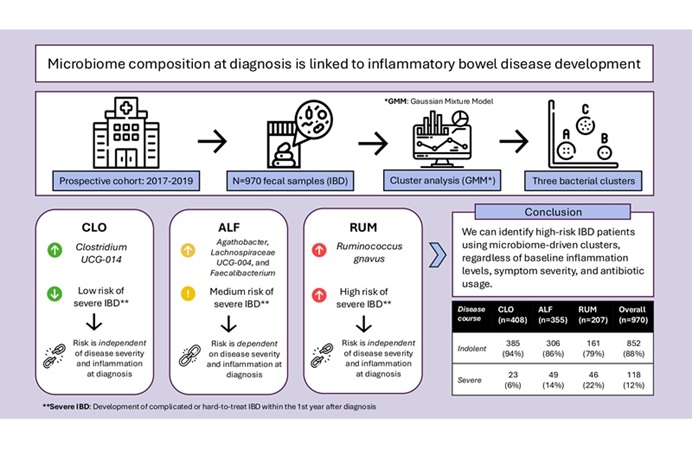
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel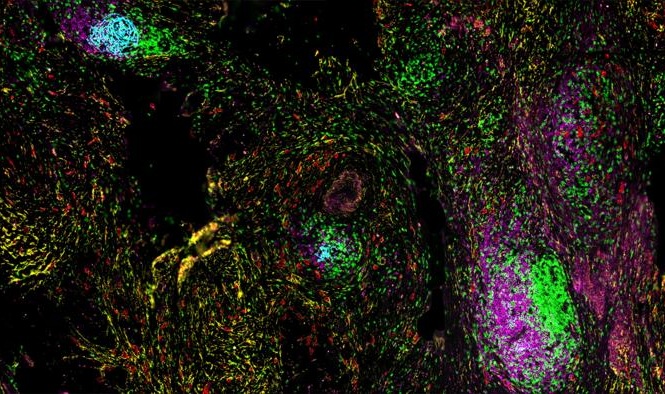
AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
Tertiary lymphoid structures are emerging as important indicators of antitumor immunity, but their heterogeneity and spatial context within tumors remain difficult to capture through routine diagnostics.... Read more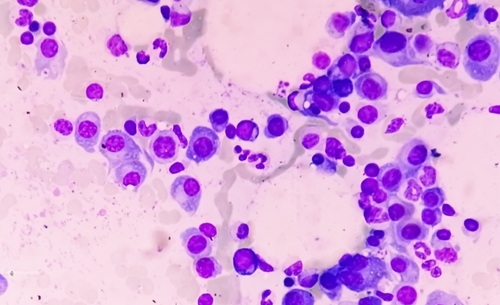
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel