New Technique Detects Breaks in Mitochondrial DNA
|
By LabMedica International staff writers Posted on 22 Apr 2019 |
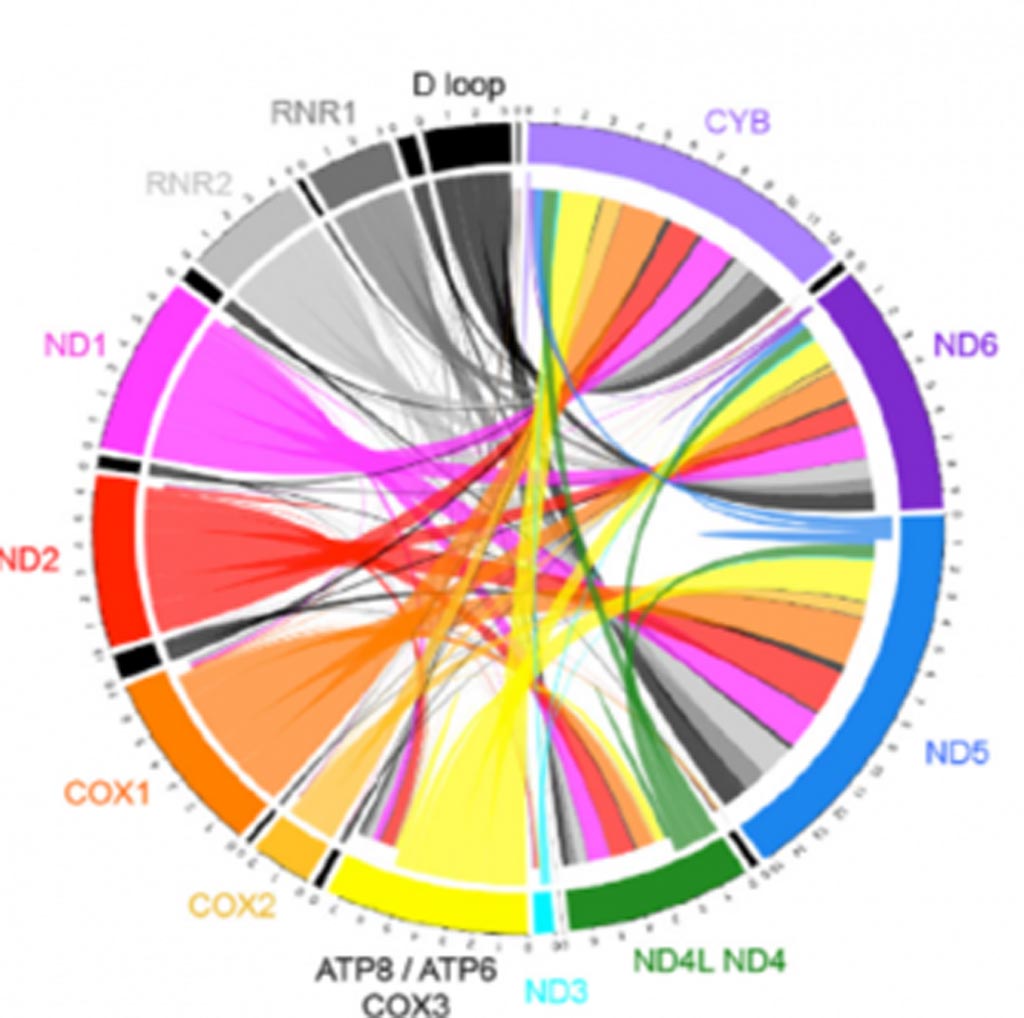
Image: A catalog of deletions (4,489) observed in brain samples derived from both healthy subjects and subjects with psychiatric disorders. The burden of deletions accumulates in various brain regions during aging. Many deletions play a major role in classical mitochondrial disorders, and deletion burden is viewed as an indicator of long lasting mitochondrial oxidative stress. Each colored ribbon is composed of individual lines showing the relative amount of deletions in brain samples in the catalog (Photo courtesy of the University of California, Irvine).
The Splice-Break pipeline is a recently described technique that can detect and quantify mitochondrial DNA (mtDNA) deletions at a high level of resolution.
Deletions in the mitochondrial genome have been implicated in numerous human disorders that often display muscular and/or neurological symptoms due to the high-energy demands of these tissues. Among these "mitochondrial myopathies" are Kearns–Sayre syndrome (KSS), Pearson Syndrome (PS), chronic progressive external ophthalmoplegia (CPEO), Leigh syndrome, and diabetes mellitus.
Investigators at the University of California, Irvine (USA) described a catalogue of 4,489 putative mtDNA deletions, including their frequency and relative read rate. To do this, they employed a combinatorial approach of mitochondria-targeted PCR, next-generation sequencing, bioinformatics, post-hoc filtering, annotation, and validation steps. Their bioinformatics pipeline incorporated MapSplice, an RNA-seq splice junction detection algorithm, to detect and quantify mtDNA deletion breakpoints rather than mRNA splices.
The investigators used their technique to analyze 93 samples from postmortem brain and blood. They found that the 4977-base pairs "common deletion" was neither the most frequent deletion nor the most abundant and that brain contained significantly more deletions than blood.
“Taken together, the pipeline will enable us to look in many brain regions for an accumulation of damage to mitochondria DNA for individuals with various psychiatric symptoms such as depression and psychosis. The ultimate use will be to test other more accessible samples such as blood, saliva, or cerebrospinal fluid from patients to estimate the damage to mitochondria, and quickly identify those individuals who may benefit from drugs and other treatments that give a mitochondria boost and improve psychiatric symptoms,” said senior author Dr. Marquis P. Vawter, a researcher in the department of psychiatry and human behavior at the University of California, Irvine. “This technique allows us to use a single test to measure the accumulation of many types of these deletions and to determine an overall burden of these deletions upon mitochondria functions.”
The study was published in the March 14, 2019, online edition of the journal Nucleic Acids Research.
Related Links:
University of California, Irvine
Deletions in the mitochondrial genome have been implicated in numerous human disorders that often display muscular and/or neurological symptoms due to the high-energy demands of these tissues. Among these "mitochondrial myopathies" are Kearns–Sayre syndrome (KSS), Pearson Syndrome (PS), chronic progressive external ophthalmoplegia (CPEO), Leigh syndrome, and diabetes mellitus.
Investigators at the University of California, Irvine (USA) described a catalogue of 4,489 putative mtDNA deletions, including their frequency and relative read rate. To do this, they employed a combinatorial approach of mitochondria-targeted PCR, next-generation sequencing, bioinformatics, post-hoc filtering, annotation, and validation steps. Their bioinformatics pipeline incorporated MapSplice, an RNA-seq splice junction detection algorithm, to detect and quantify mtDNA deletion breakpoints rather than mRNA splices.
The investigators used their technique to analyze 93 samples from postmortem brain and blood. They found that the 4977-base pairs "common deletion" was neither the most frequent deletion nor the most abundant and that brain contained significantly more deletions than blood.
“Taken together, the pipeline will enable us to look in many brain regions for an accumulation of damage to mitochondria DNA for individuals with various psychiatric symptoms such as depression and psychosis. The ultimate use will be to test other more accessible samples such as blood, saliva, or cerebrospinal fluid from patients to estimate the damage to mitochondria, and quickly identify those individuals who may benefit from drugs and other treatments that give a mitochondria boost and improve psychiatric symptoms,” said senior author Dr. Marquis P. Vawter, a researcher in the department of psychiatry and human behavior at the University of California, Irvine. “This technique allows us to use a single test to measure the accumulation of many types of these deletions and to determine an overall burden of these deletions upon mitochondria functions.”
The study was published in the March 14, 2019, online edition of the journal Nucleic Acids Research.
Related Links:
University of California, Irvine
Latest BioResearch News
- Microenvironment Biomarkers Could Enable Early Lung Cancer Detection
- Study Identifies Protein Changes Driving Immunotherapy Resistance in Multiple Myeloma
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- Innate Immunity Variants Associated With Earlier Breast Cancer in BRCA1 Carriers
- Genetic Cause Identified for Severe Infant Epilepsy
- Study Reveals Diagnostic and Therapeutic Target in Rare Pancreatic Tumors
- Researchers Identify Survival Pathway Undermining Targeted Cancer Drugs
- Large-Scale Study Maps DNA Damage Signatures Across Multiple Cancers
- Study Identifies Distinct Immune Signatures to Early Depression and Psychosis
- Genetic Mutation Behind Aggressive Adult Leukemia Offers Treatment Clues
Channels
Clinical Chemistry
view channel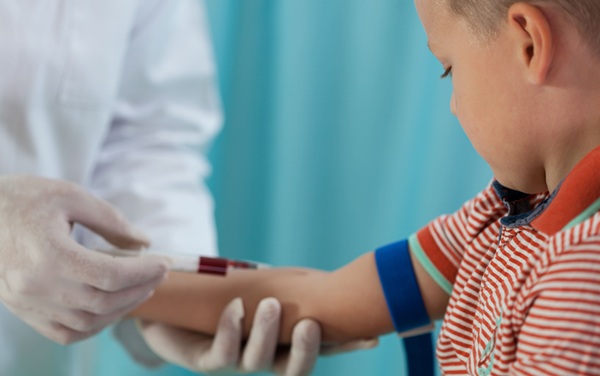
Proteomic Data Underscore Need for Age-Specific Pediatric Reference Ranges
Serum proteins underpin many routine tests used to detect inflammation, hormonal imbalance, cardiovascular disease, and metabolic disorders. Yet pediatric interpretation often relies on adult reference... Read more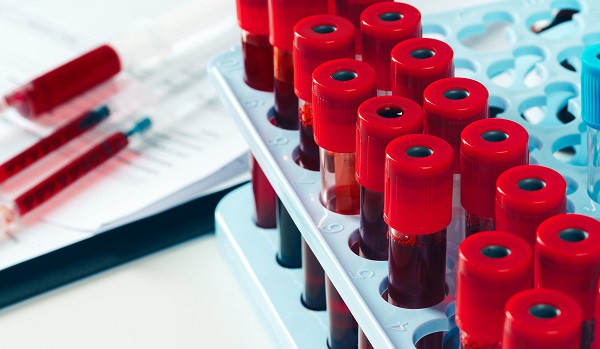
Routine Blood Count Ratio Linked to Future Alzheimer’s and Dementia Risk
Alzheimer’s disease and related dementias develop over years, making it difficult to identify at-risk patients before symptoms appear. Clinicians therefore need widely available laboratory markers that... Read more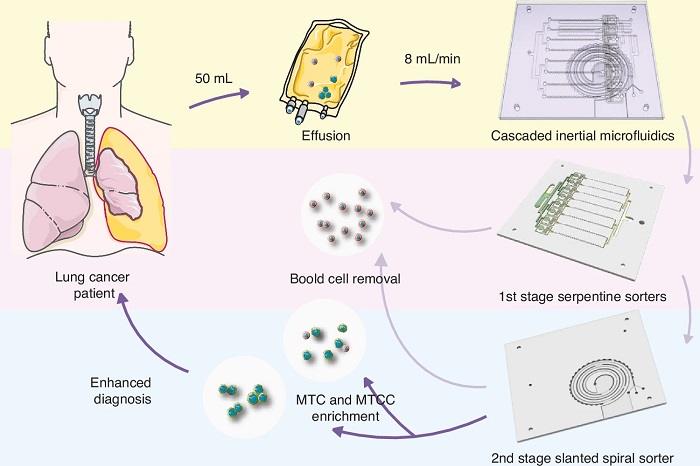
Label-Free Microfluidic Device Enriches Tumor Cells and Clusters from Pleural Effusions
Diagnosing malignancy from pleural effusion remains challenging because tumor cells are rare and clusters are easily disrupted during processing. Conventional cytology can miss malignant tumor cells and... Read moreMolecular Diagnostics
view channel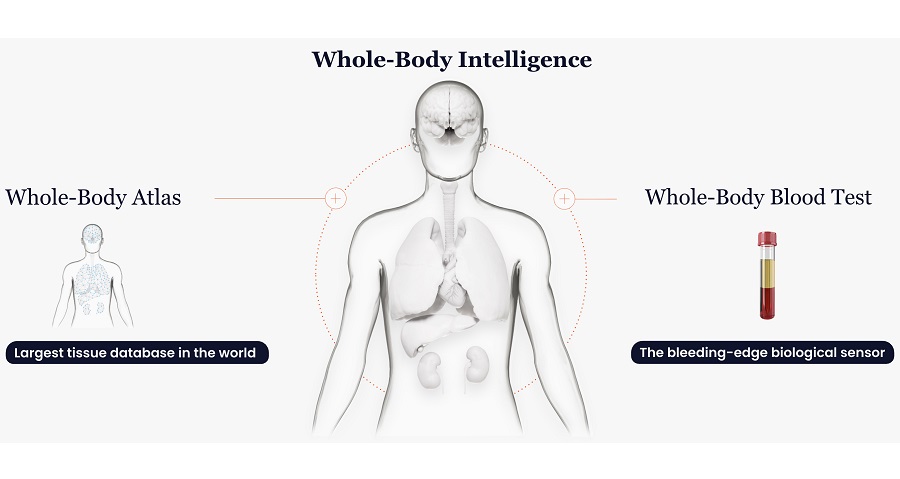
AI Blood Test Enhances Monitoring of Liver Cirrhosis Progression
Monitoring chronic liver disease remains difficult because clinicians rely on tools that can be inconsistent and may miss early progression. Standard approaches often combine ultrasound imaging with blood-based... Read more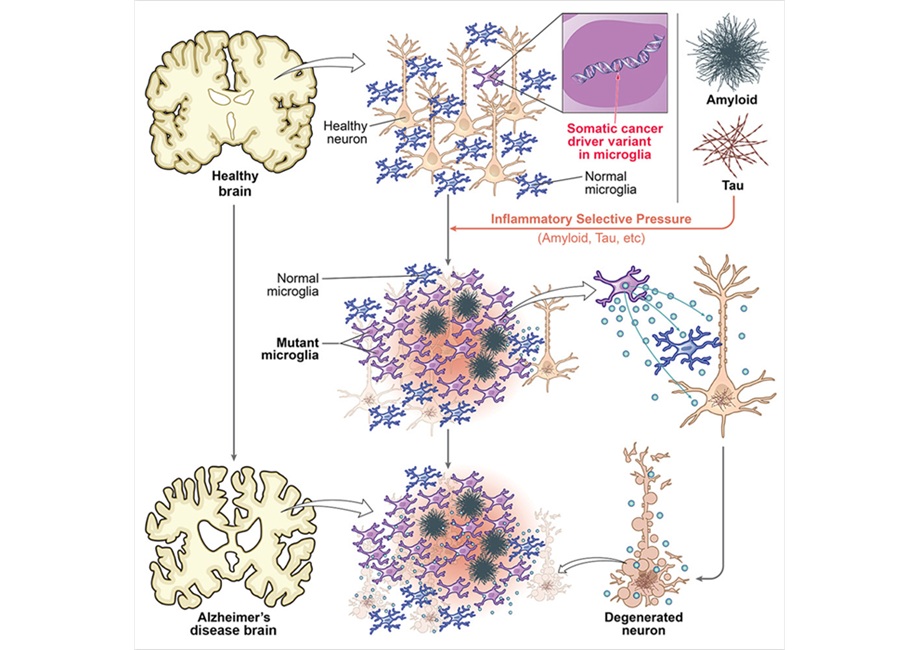
Cancer-Related Mutations in Immune Cells Linked to Alzheimer’s
Alzheimer’s disease is marked by protein aggregation and inflammatory changes in the brain’s immune system, yet its molecular drivers remain incompletely understood. With aging, human cells accumulate... Read more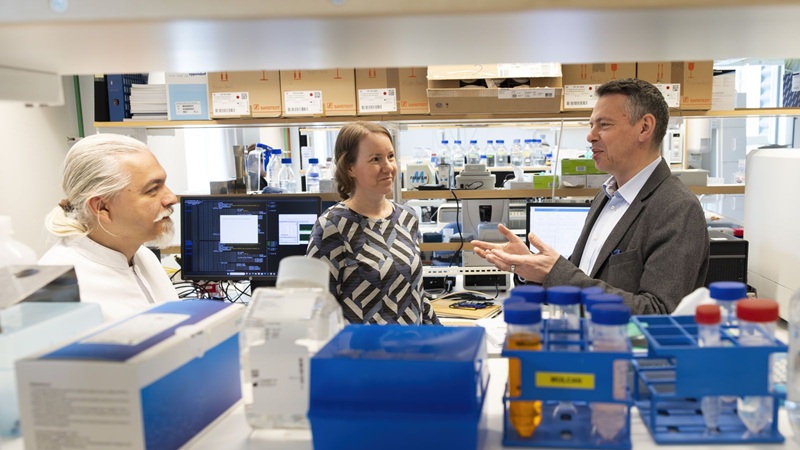
Composite Blood Biomarkers Enable Early Detection of Common Cancers
Early diagnosis of colorectal, lung, and ovarian cancers remains challenging, with many patients identified only after tumors have begun to spread. A scalable blood test could expand access to screening,... Read more
Machine Learning Model Uses DNA Methylation to Predict Tumor Origin in Cancers of Unknown Primary
Cancers of unknown primary (CUP) are metastatic malignancies in which the primary site cannot be identified, complicating treatment selection. Many patients consequently receive broad, nonspecific chemotherapy... Read moreHematology
view channel
Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
CareDx (Brisbane, CA, USA) has introduced AlloSeq Nano, a nanopore‑based HLA (human leukocyte antigen) and ABO genotyping solution unveiled at the European Federation for Immunogenetics (EFI) Conference 2026.... Read more
Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
Diffuse large B-cell lymphoma (DLBCL) is the most common form of non-Hodgkin lymphoma and often presents with aggressive clinical behavior. Although many patients respond to standard chemotherapy with... Read moreImmunology
view channel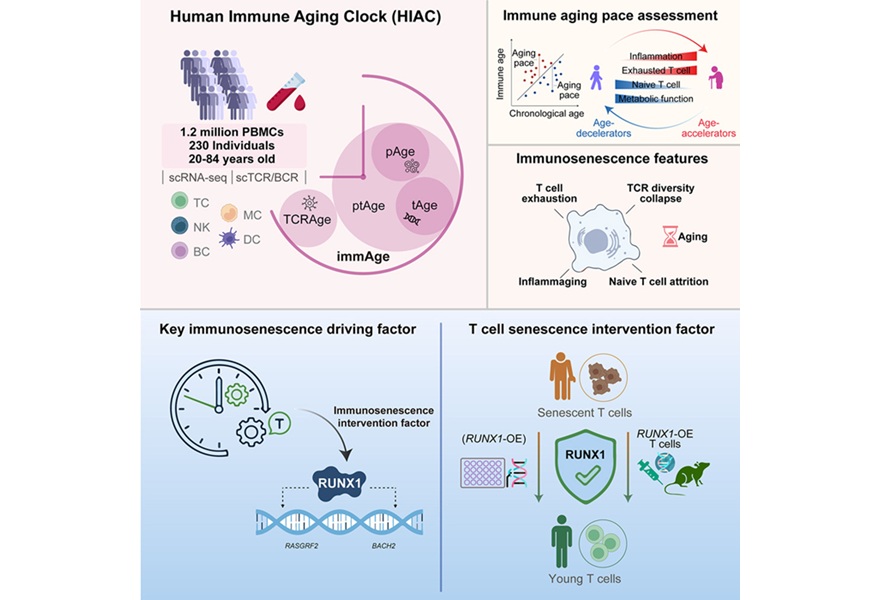
Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
Immune aging undermines host defense and contributes to multiple age-related diseases, yet its heterogeneity complicates measurement and intervention. Clinical laboratories increasingly seek objective... Read more
Study Finds Influenza Often Undiagnosed in Winter Deaths
Seasonal influenza drives substantial excess mortality, yet its contribution is often obscured when infections go undiagnosed near the time of death. Many deaths occur outside hospitals or in older adults... Read moreMicrobiology
view channel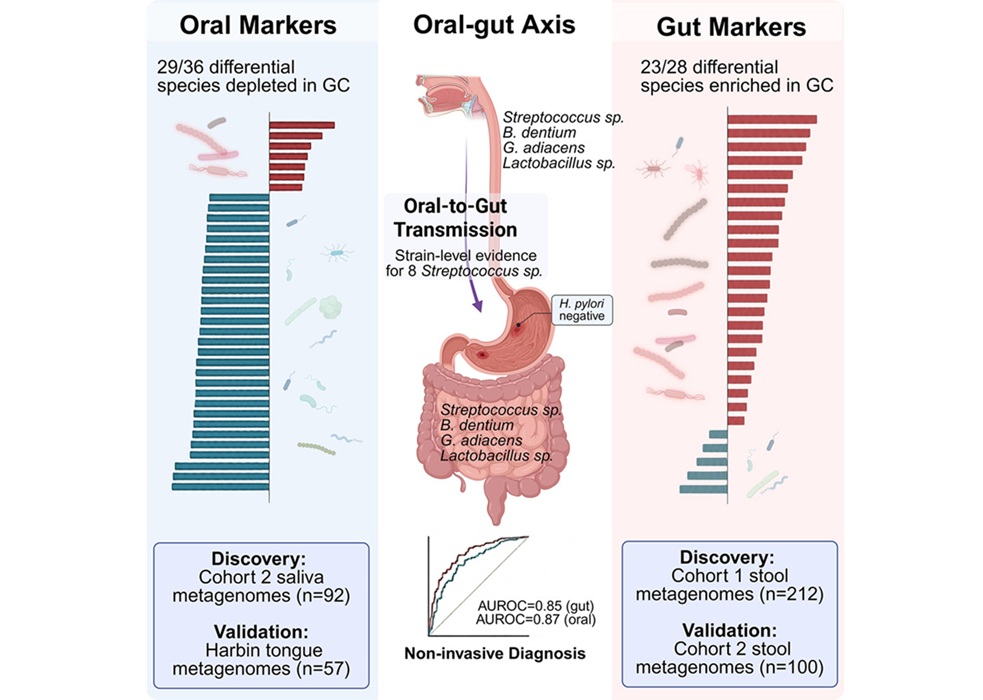
Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
Early detection of gastric cancer could be advanced by scalable screening strategies using minimally invasive sampling. Saliva collection is noninvasive and cost-effective, supporting wider adoption... Read more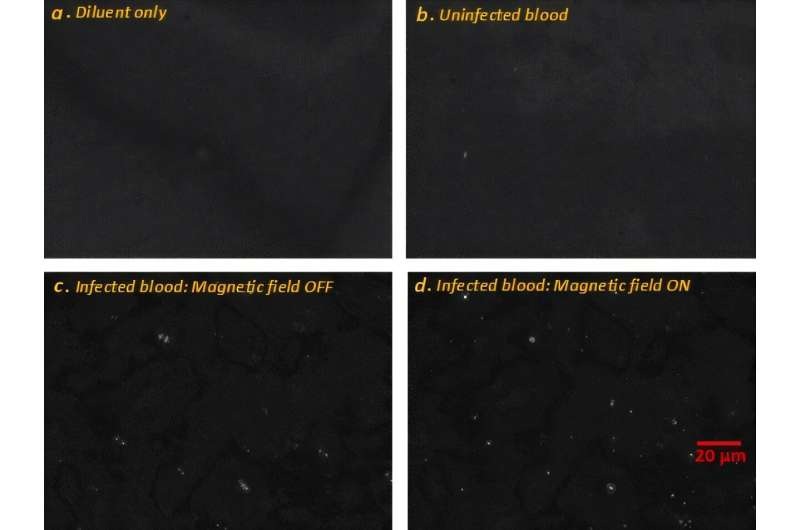
Label-Free Microscopy Methodd Enables Faster, Quantitative Detection of Malaria
Microscopy of blood smears remains a cornerstone for malaria diagnosis but can be slow, stain-dependent, and operator intensive. With more than 200 million infections and over 600,000 deaths annually,... Read more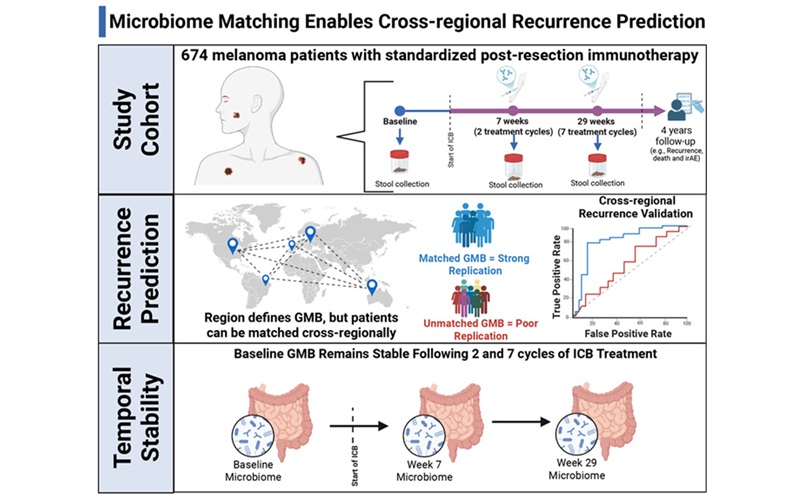
Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
Melanoma remains prone to relapse even after surgery and adjuvant immunotherapy, with 25% to 40% of patients experiencing recurrence. Clinicians lack reliable pre-treatment indicators to identify those... Read more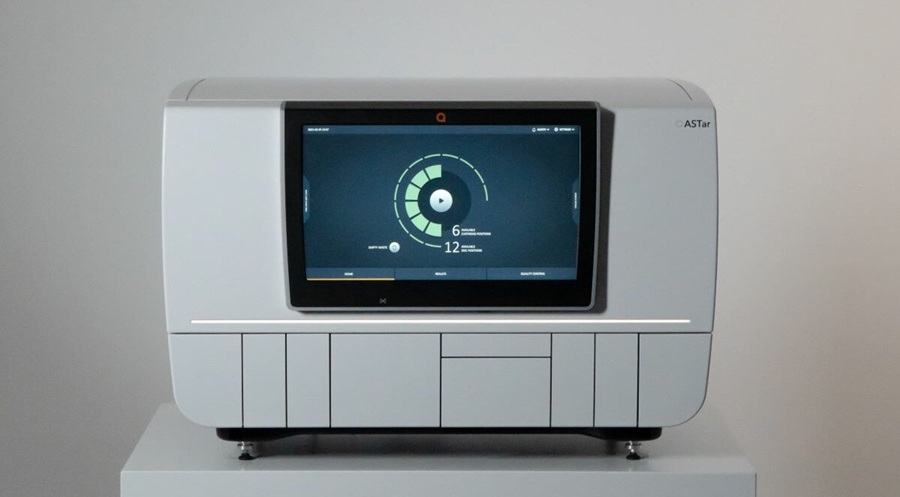
Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
Gram-negative bloodstream infections and sepsis demand fast, precise antimicrobial therapy, yet conventional susceptibility workflows can delay targeted treatment. Clinical laboratories need platforms... Read morePathology
view channel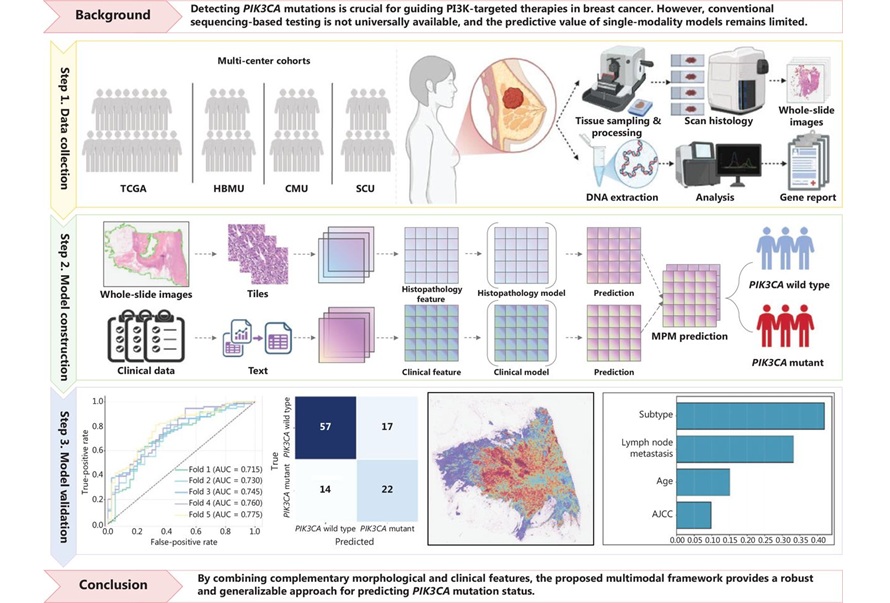
Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
PIK3CA mutations are key biomarkers for selecting phosphoinositide 3-kinase (PI3K)–targeted therapies in breast cancer, yet access to molecular testing can be inconsistent and costly. Conventional polymerase... Read more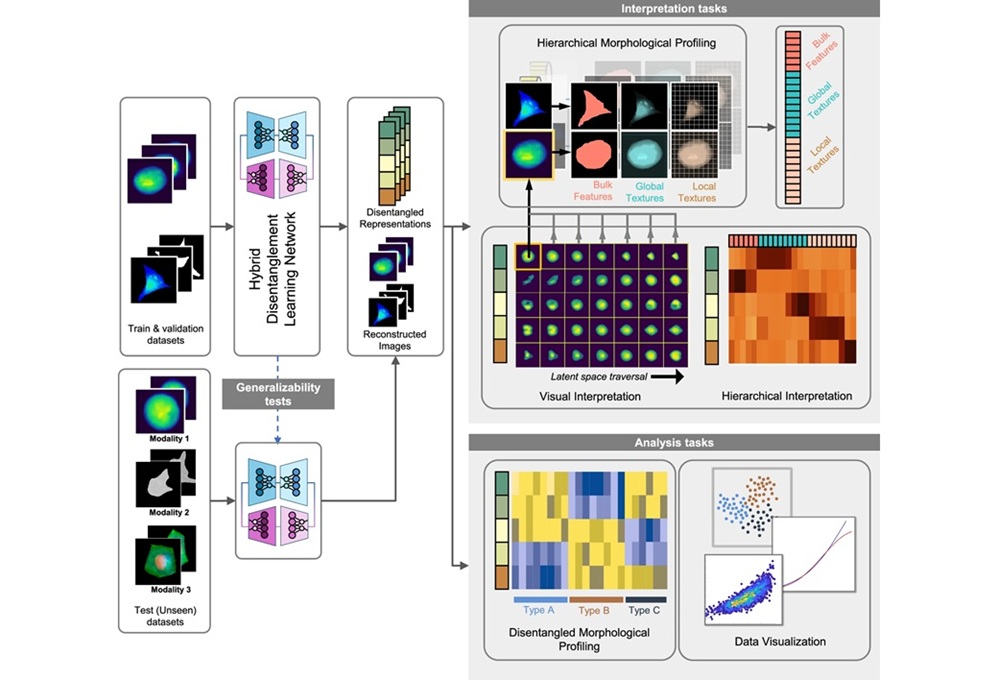
Interpretable AI Reveals Hidden Cellular Features from Microscopy Images
Microscopy images contain rich clues about cell health, but many disease-relevant morphological differences are too subtle to see and difficult to quantify consistently. Artificial intelligence (AI) has... Read moreTechnology
view channel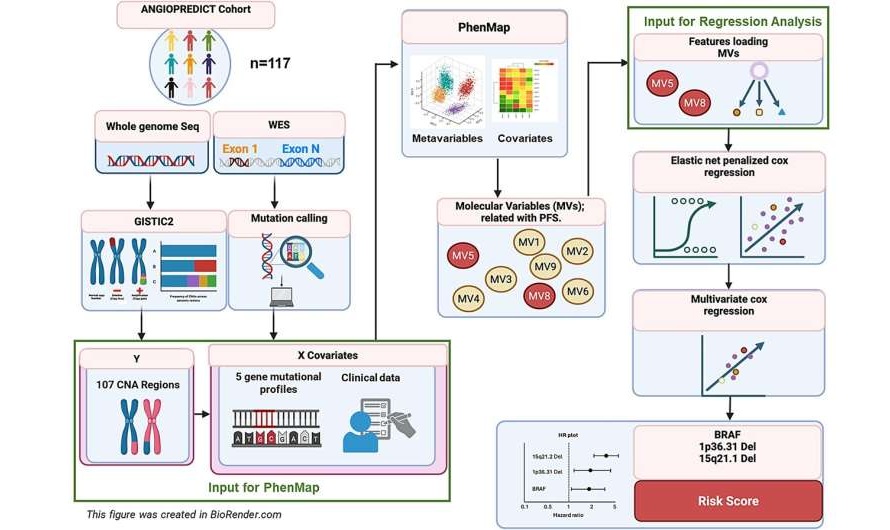
AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
Advanced bowel cancer remains difficult to treat, and many patients receive targeted therapies that do not help them but still cause harm. Clinicians need reliable ways to identify likely responders before... Read more
Integrated System Streamlines Pre-Analytical Workflow for Molecular Testing
Pre-analytical variation remains a leading source of inconsistent molecular test results and added costs, particularly when laboratories rely on multiple instruments and protocols. Standardizing nucleic... Read moreIndustry
view channel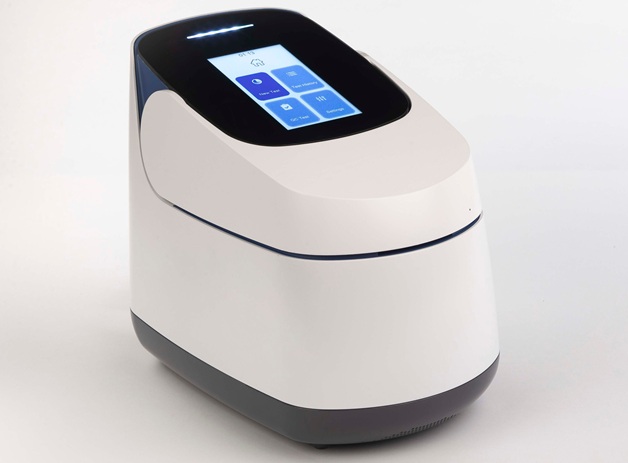
QuidelOrtho Adds Ultra-Fast PCR Platform with LEX Acquisition
QuidelOrtho Corporation has completed the acquisition of LEX Diagnostics for approximately USD 100 million in cash. The transaction adds the LEX VELO System to QuidelOrtho’s portfolio. The platform received U.... Read more
Seegene Showcases Real-Time PCR Data Analytics Platform at ESCMID
Seegene introduced STAgora, a real-time data analytics platform built on aggregated statistical testing data, at ESCMID Global 2026 in Munich, where it also presented an enhanced model of its automated... Read more
Roche Affiliate Expands MRD Portfolio with SAGA Acquisition
Foundation Medicine, Inc., an independent affiliate of Roche, announced plans to expand its monitoring portfolio with SAGA Diagnostics’ Pathlight, a personalized, tumor-informed molecular residual disease... Read more