CRISPR Variant Enables More Accurate Genome Editing
|
By LabMedica International staff writers Posted on 13 Aug 2018 |
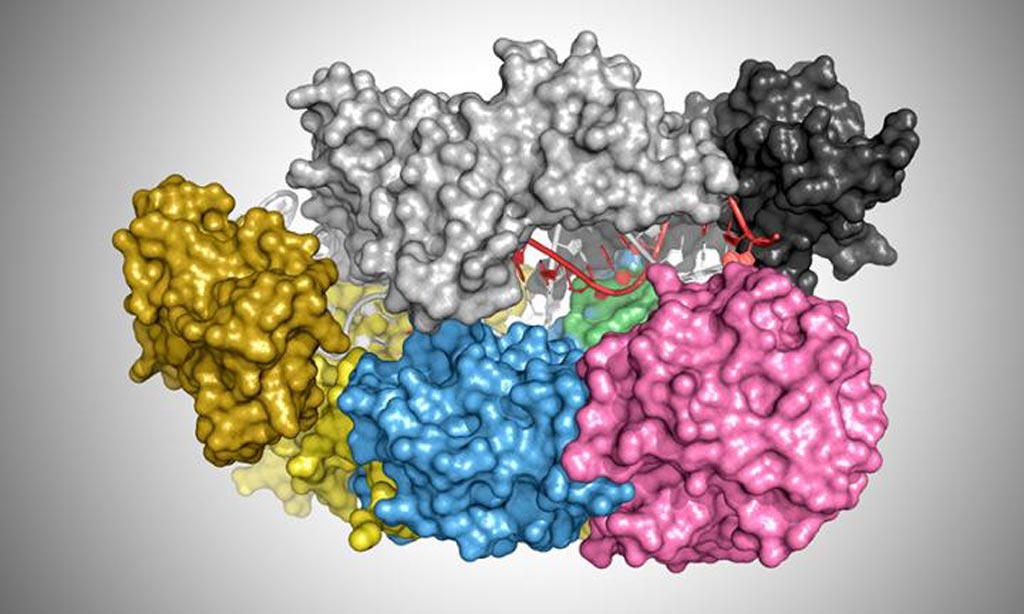
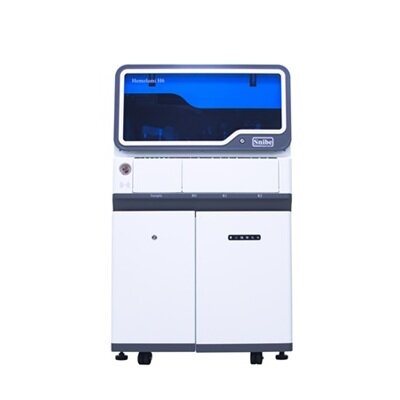
Image: An illustration showing the protein cas12a bound to a DNA helix (red and white) (Photo courtesy of T. Yamano and H. Nishimasu / James Rybarski).
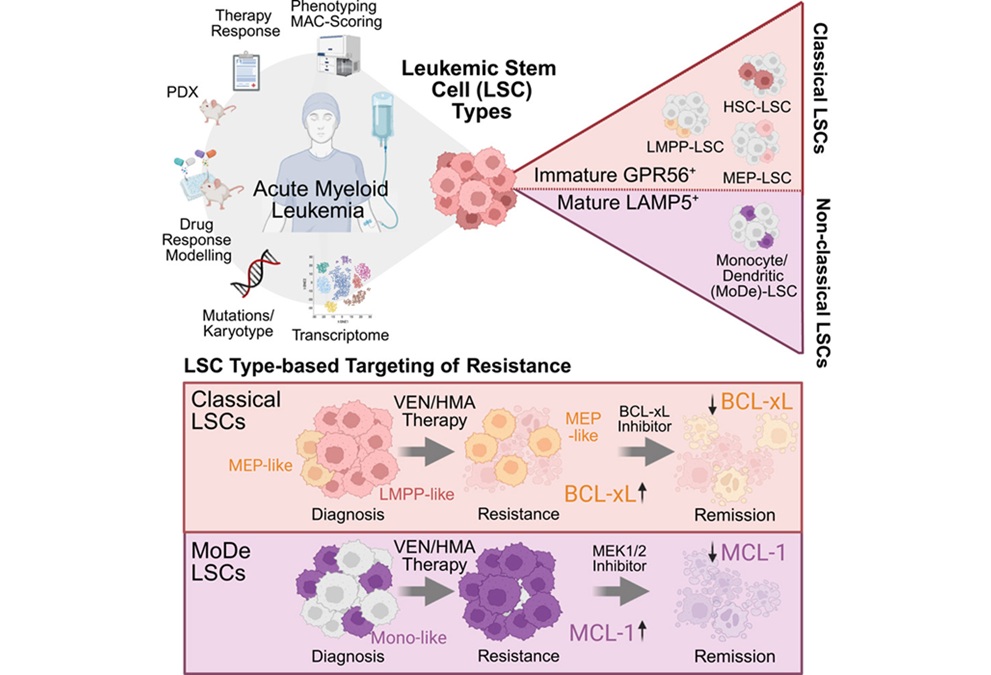
Results of a recent study suggested that inherent binding properties of the Cas12a enzyme make it a better choice for gene editing purposes than the currently used CRISPR/Cas9 combination.
CRISPR/Cas9 is regarded as the cutting edge of molecular biology technology. CRISPRs (clustered regularly interspaced short palindromic repeats) are segments of prokaryotic DNA containing short repetitions of base sequences. Each repetition is followed by short segments of "spacer DNA" from previous exposures to a bacterial virus or plasmid. Since 2013, the CRISPR/Cas9 system has been used in research for gene editing (adding, disrupting, or changing the sequence of specific genes) and gene regulation. By delivering the Cas9 enzyme and appropriate guide RNAs (sgRNAs) into a cell, the organism's genome can be cut at any desired location. The conventional CRISPR/Cas9 system is composed of two parts: the Cas9 enzyme, which cleaves the DNA molecule and specific RNA guides that shepherd the Cas9 protein to the target gene on a DNA strand.
Despite the widespread usage of CRISPR/Cas9, DNA cleavage at off-target sites that resemble the target sequence continues to be a pervasive problem that remains poorly understood mechanistically.
To solve the problem of off-target DNA cleavage, investigators at the University of Texas at Austin (USA) used quantitative kinetics to dissect the reaction steps of DNA targeting by Cas12a (also known as Cpf1 or Centromere and Promoter Factor 1). CRISPR/Cpf1 differs from CRISPR/Cas9 in a number of key ways. Cpf1 is much smaller than the Cas9 enzyme, which makes it easier to package inside a virus and therefore easier to deliver to muscle cells. It also recognizes a different sequence of DNA than Cas9 does, which provides greater flexibility in terms of use.
The investigators reported in the August 2, 2018, online edition of the journal Molecular Cell that Cas12a bound DNA tightly in two kinetically separable steps and discriminated strongly against mismatches along most of the DNA target sequence. In mechanistic terms, the investigators explained that Cas9, initially bound base pairs tightly together, but after the first seven or eight base pairs in the genomic target it paid less attention. This implied that Cas9 could overlook mismatches that could lead it to edit the wrong part of the genome. In contrast, Cas12a formed bonds that were relatively weak. Thus, it required a good match all along the target molecule to hold the complex together long enough to make an edit. Therefore it was much more likely to edit only the intended part of the genome.
"It makes the process of base-pair formation more reversible," said senior author Dr. Rick Russell, professor of molecular biosciences at University of Texas at Austin. "In other words, Cas12a does a better job of checking each base pair before moving on to the next one. After seven or eight letters, Cas9 stops checking, whereas Cas12a keeps on checking out to about 18 letters."
Related Links:
University of Texas at Austin
CRISPR/Cas9 is regarded as the cutting edge of molecular biology technology. CRISPRs (clustered regularly interspaced short palindromic repeats) are segments of prokaryotic DNA containing short repetitions of base sequences. Each repetition is followed by short segments of "spacer DNA" from previous exposures to a bacterial virus or plasmid. Since 2013, the CRISPR/Cas9 system has been used in research for gene editing (adding, disrupting, or changing the sequence of specific genes) and gene regulation. By delivering the Cas9 enzyme and appropriate guide RNAs (sgRNAs) into a cell, the organism's genome can be cut at any desired location. The conventional CRISPR/Cas9 system is composed of two parts: the Cas9 enzyme, which cleaves the DNA molecule and specific RNA guides that shepherd the Cas9 protein to the target gene on a DNA strand.
Despite the widespread usage of CRISPR/Cas9, DNA cleavage at off-target sites that resemble the target sequence continues to be a pervasive problem that remains poorly understood mechanistically.
To solve the problem of off-target DNA cleavage, investigators at the University of Texas at Austin (USA) used quantitative kinetics to dissect the reaction steps of DNA targeting by Cas12a (also known as Cpf1 or Centromere and Promoter Factor 1). CRISPR/Cpf1 differs from CRISPR/Cas9 in a number of key ways. Cpf1 is much smaller than the Cas9 enzyme, which makes it easier to package inside a virus and therefore easier to deliver to muscle cells. It also recognizes a different sequence of DNA than Cas9 does, which provides greater flexibility in terms of use.
The investigators reported in the August 2, 2018, online edition of the journal Molecular Cell that Cas12a bound DNA tightly in two kinetically separable steps and discriminated strongly against mismatches along most of the DNA target sequence. In mechanistic terms, the investigators explained that Cas9, initially bound base pairs tightly together, but after the first seven or eight base pairs in the genomic target it paid less attention. This implied that Cas9 could overlook mismatches that could lead it to edit the wrong part of the genome. In contrast, Cas12a formed bonds that were relatively weak. Thus, it required a good match all along the target molecule to hold the complex together long enough to make an edit. Therefore it was much more likely to edit only the intended part of the genome.
"It makes the process of base-pair formation more reversible," said senior author Dr. Rick Russell, professor of molecular biosciences at University of Texas at Austin. "In other words, Cas12a does a better job of checking each base pair before moving on to the next one. After seven or eight letters, Cas9 stops checking, whereas Cas12a keeps on checking out to about 18 letters."
Related Links:
University of Texas at Austin
Latest BioResearch News
- Lung Cancer Study Reveals Cellular Program Behind Therapy Resistance
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
Channels
Clinical Chemistry
view channel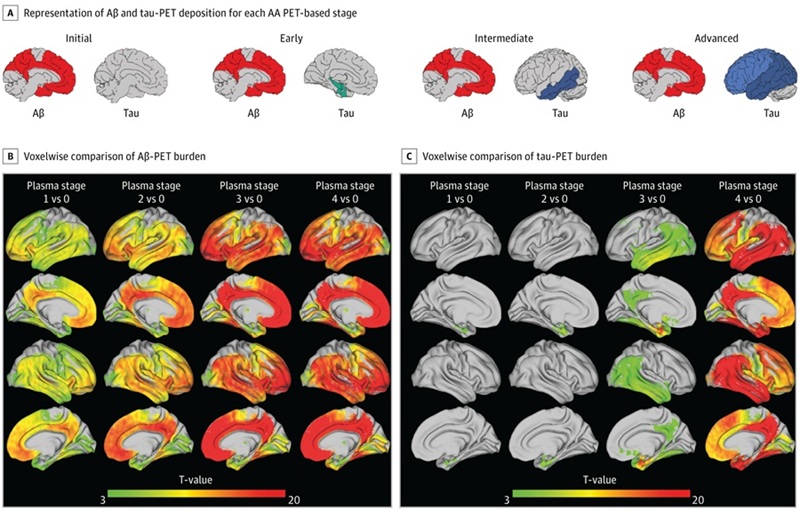
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreMolecular Diagnostics
view channel
Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
Selecting adjuvant therapy for early-stage hormone receptor-positive breast cancer depends on accurately assessing long-term risk of distant recurrence. Clinical features alone can leave uncertainty about... Read more
Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
Many cancers deviate from the normal chromosome number, but the clinical impact of extreme chromosome loss remains unclear. This widespread genomic disruption is associated with aggressive disease and... Read more
Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
Sarcomas remain difficult to monitor during immunotherapy, as low tumor mutation burden can limit traditional circulating tumor DNA approaches and repeat tissue biopsies are often impractical in advanced disease.... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more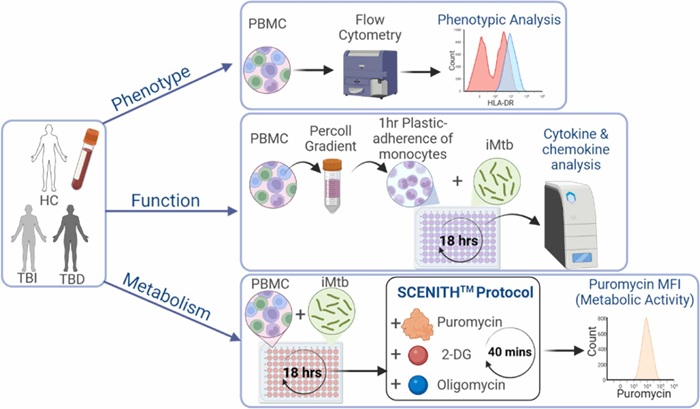
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read moreMicrobiology
view channel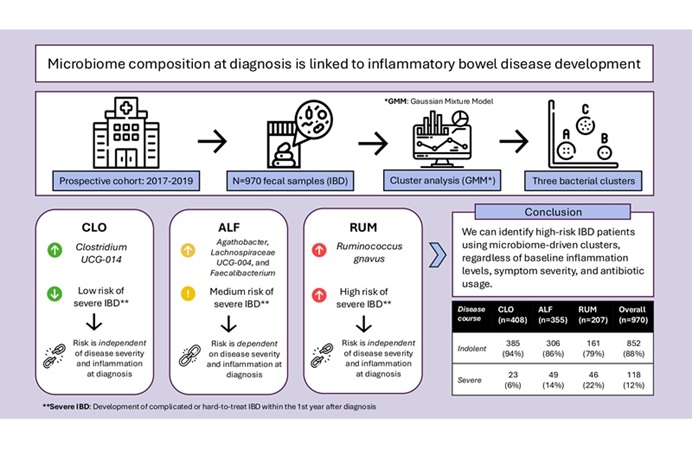
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more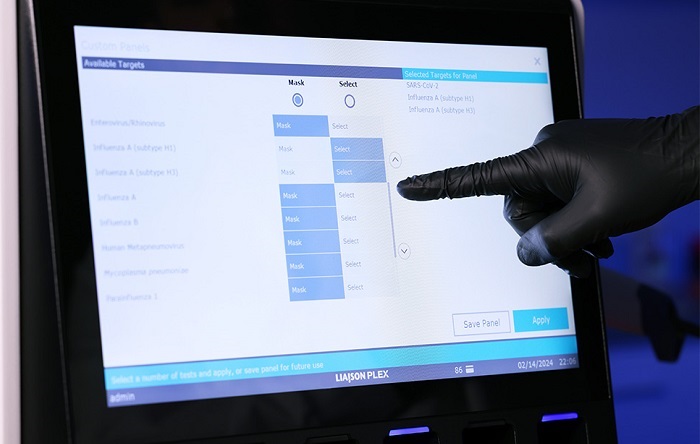
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel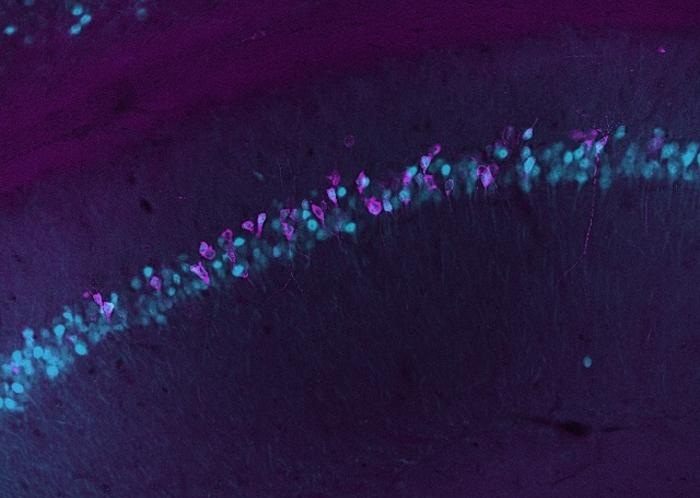
Blood-Based Method Tracks Gene Activity in the Living Brain
Real-time measurement of gene activity in the brain has been limited by assays requiring destructive tissue sampling. Tracking active genes could reveal how the body responds to environmental factors,... Read more
FDA Approval Expands Automated PD-L1 Testing Across Solid Tumors
Clinical laboratories play a central role in guiding immunotherapy by reporting programmed death ligand-1 (PD‑L1) status across multiple solid tumors. Many sites are standardizing this work on fully automated... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel




.jpg)