Largest Data Set of Cancer-Related Genetic Variations Generated for Researchers
|
By LabMedica International staff writers Posted on 29 Jul 2013 |
US scientists have generated a data set of cancer-specific genetic variations and are making these data available to the research community.
The investigators, from the US National Cancer Institute (NCI; Bethesda, MD, USA), published their study’s findings July 15, 2013, online in Cancer Research, a journal of the American Association for Cancer Research.
This new technology will help cancer researchers better illuminate drug response and resistance to cancer treatments. “To date, this is the largest database worldwide, containing six billion data points that connect drugs with genomic variants for the whole human genome across cell lines from nine tissues of origin, including breast, ovary, prostate, colon, lung, kidney, brain, blood, and skin,” said Yves Pommier, MD, PhD, chief of the laboratory of molecular pharmacology at the NCI in an interview. “We are making this data set public for the greater community to use and analyze. Opening this extensive data set to researchers will expand our knowledge and understanding of tumorigenesis, as more and more cancer-related gene aberrations are discovered. This comes at a great time, because genomic medicine is becoming a reality, and I am very hopeful this valuable information will change the way we use drugs for precision medicine.”
Dr. Pommier and colleagues conducted whole-exome sequencing of the NCI-60 human cancer cell-line panel, which is an assortment of 60 human cancer cell lines, and generated a comprehensive list of cancer-specific genetic variations. Early research conducted by the researchers show that the extensive data set has the potential to greatly enhance understanding of the links between specific cancer-related genetic variations and drug response, which will hasten the drug development process.
The NCI-60 human cancer cell-line panel is used extensively by cancer researchers to discover novel anticancer drugs. To conduct whole-exome sequencing, Dr. Pommier and his NCI team extracted DNA from the 60 different cell lines from tumors of the lung, colon, brain, ovary, prostate, breast, and kidney, as well as melanoma and leukemia, and cataloged the genetic coding variants for the complete human genome. The genetic variations identified were of two types: type I variants corresponding to variants found in the normal population, and type II variants, which are cancer-specific.
The scientists then employed the Super Learner algorithm to predict the sensitivity of cells harboring type II variants to 103 anticancer drugs approved by the US Food and Drug Administration (FDA) and an additional 207 investigational new pharmaceutical agents. They were able to assess the correlations between key cancer-related genes and clinically pertinent anticancer drugs, and predict the outcome.
The data generated in this project provide a way to identify new determinants of response and processes of drug resistance, and offer opportunities to target genomic defects and overcome acquired resistance, according to Dr. Pommier. To accomplish this, the researchers are making these data available to all researchers by way of two database portals, called the CellMiner database and the Ingenuity systems database.
Related Links:
US National Cancer Institute
The investigators, from the US National Cancer Institute (NCI; Bethesda, MD, USA), published their study’s findings July 15, 2013, online in Cancer Research, a journal of the American Association for Cancer Research.
This new technology will help cancer researchers better illuminate drug response and resistance to cancer treatments. “To date, this is the largest database worldwide, containing six billion data points that connect drugs with genomic variants for the whole human genome across cell lines from nine tissues of origin, including breast, ovary, prostate, colon, lung, kidney, brain, blood, and skin,” said Yves Pommier, MD, PhD, chief of the laboratory of molecular pharmacology at the NCI in an interview. “We are making this data set public for the greater community to use and analyze. Opening this extensive data set to researchers will expand our knowledge and understanding of tumorigenesis, as more and more cancer-related gene aberrations are discovered. This comes at a great time, because genomic medicine is becoming a reality, and I am very hopeful this valuable information will change the way we use drugs for precision medicine.”
Dr. Pommier and colleagues conducted whole-exome sequencing of the NCI-60 human cancer cell-line panel, which is an assortment of 60 human cancer cell lines, and generated a comprehensive list of cancer-specific genetic variations. Early research conducted by the researchers show that the extensive data set has the potential to greatly enhance understanding of the links between specific cancer-related genetic variations and drug response, which will hasten the drug development process.
The NCI-60 human cancer cell-line panel is used extensively by cancer researchers to discover novel anticancer drugs. To conduct whole-exome sequencing, Dr. Pommier and his NCI team extracted DNA from the 60 different cell lines from tumors of the lung, colon, brain, ovary, prostate, breast, and kidney, as well as melanoma and leukemia, and cataloged the genetic coding variants for the complete human genome. The genetic variations identified were of two types: type I variants corresponding to variants found in the normal population, and type II variants, which are cancer-specific.
The scientists then employed the Super Learner algorithm to predict the sensitivity of cells harboring type II variants to 103 anticancer drugs approved by the US Food and Drug Administration (FDA) and an additional 207 investigational new pharmaceutical agents. They were able to assess the correlations between key cancer-related genes and clinically pertinent anticancer drugs, and predict the outcome.
The data generated in this project provide a way to identify new determinants of response and processes of drug resistance, and offer opportunities to target genomic defects and overcome acquired resistance, according to Dr. Pommier. To accomplish this, the researchers are making these data available to all researchers by way of two database portals, called the CellMiner database and the Ingenuity systems database.
Related Links:
US National Cancer Institute
Latest BioResearch News
- Lung Cancer Study Reveals Cellular Program Behind Therapy Resistance
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
Channels
Clinical Chemistry
view channel
Urine-Based Test Shows Promise for Autism Screening in Children
Autism spectrum disorder (ASD) is commonly diagnosed through behavioral assessments, which can involve long waits that delay intervention. Earlier identification is linked to better developmental outcomes,... Read more
Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
Childhood epilepsy remains a major neurological disorder with unmet needs for accurate, non-invasive biomarkers, as conventional tests such as electroencephalography and neuroimaging can have limited sensitivity... Read moreMolecular Diagnostics
view channel
Updated Guidance Prioritizes Stool-Based Colorectal Cancer Screening Tests
Colorectal cancer is the second-leading cause of cancer death in the United States and claimed an estimated 55,000 lives in 2026. Incidence is rising among adults younger than 50, even as overall mortality... Read more
Digital PCR Assays Support Surveillance of Bundibugyo Ebolavirus Outbreak
QIAGEN (Venlo, Netherlands) has introduced two custom-designed research-use-only (RUO) QIAcuity dPCR assays to support infectious disease research and surveillance connected to the Bundibugyo ebolavirus outbreak.... Read more
Blood-Based Proteomic Test May Predict Treatment Response in Non-Small Cell Lung Cancer
Lung cancer remains the leading cause of cancer death, with non-small cell lung cancer (NSCLC) accounting for most cases. Treatment decisions are often made without a clear indication of how a patient... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel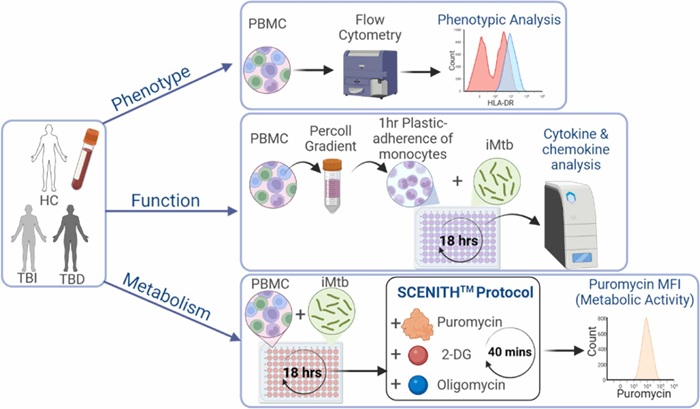
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read more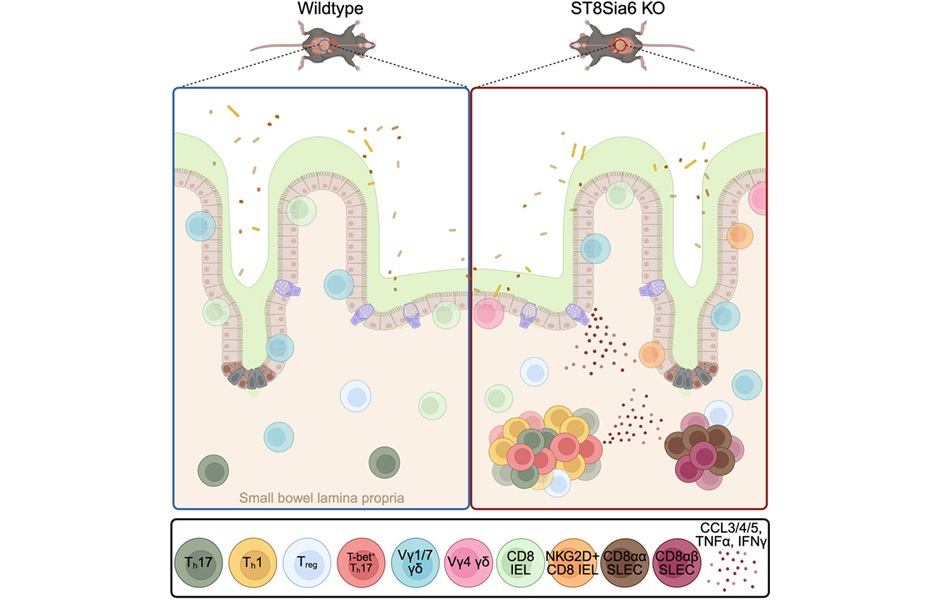
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read moreMicrobiology
view channel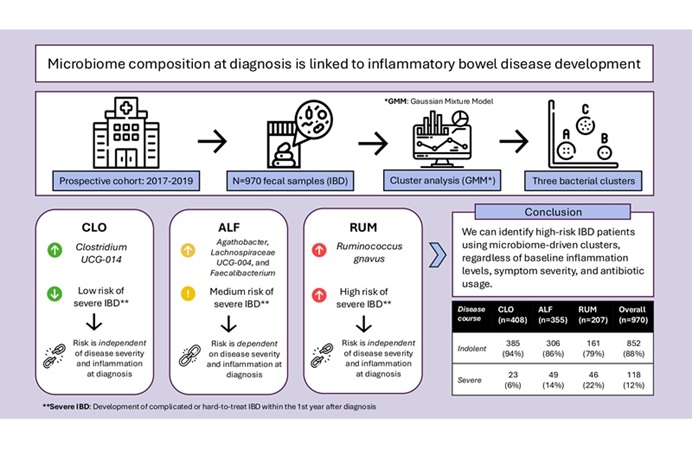
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more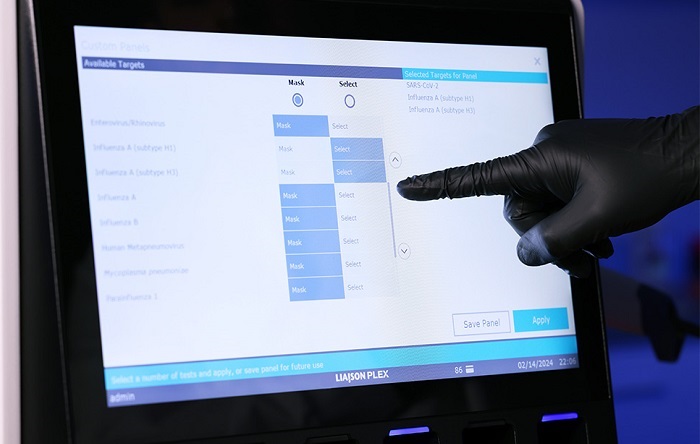
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel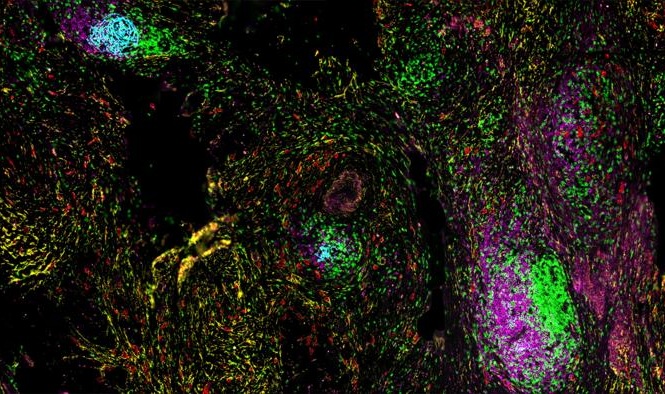
AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
Tertiary lymphoid structures are emerging as important indicators of antitumor immunity, but their heterogeneity and spatial context within tumors remain difficult to capture through routine diagnostics.... Read more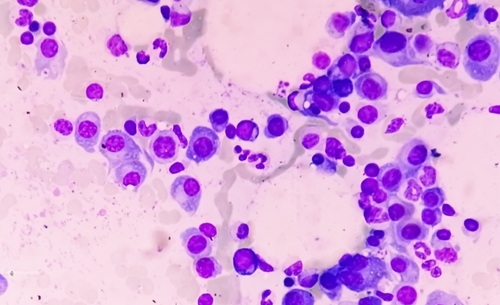
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read moreTechnology
view channel
Mailed Screening Kits Help Reduce Colorectal Cancer Screening Gaps
Colorectal cancer screening is a longstanding preventive priority, yet participation and follow-up remain uneven across patient groups. Safety‑net primary care settings often face barriers that limit screening... Read more
Algorithm Panel Aids Liver Fibrosis Assessment and Liver Cancer Surveillance
Chronic liver disease is common and often progresses silently, increasing the risk of cirrhosis and hepatocellular carcinoma when not detected early. With an estimated 1.5 billion people affected worldwide... Read moreIndustry
view channelWerfen and Oxford Nanopore Collaborate on Transplant Assay Development
Werfen (Barcelona, Spain), a global specialized diagnostics company, has announced a strategic collaboration with Oxford Nanopore Technologies (Oxford, UK), which develops nanopore-based sequencing technology,... Read more