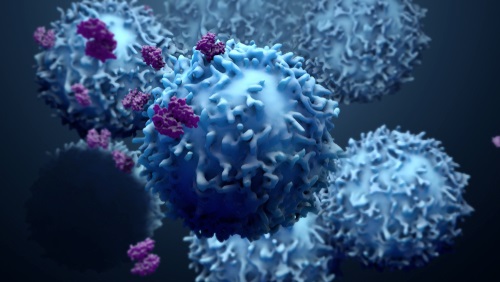
T Cells Recognize Recent SARS-CoV-2 Variants
|
By LabMedica International staff writers Posted on 15 Apr 2021 |
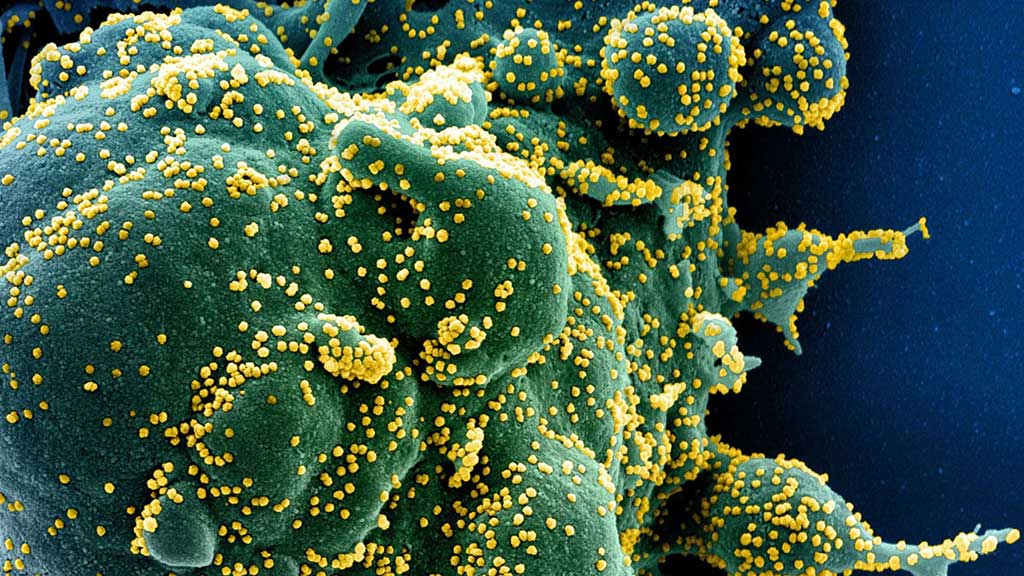
Image: Immune T cells lymphocytes can seek and destroy a cell (green) infected with and making copies of SARS-CoV-2 (yellow) (Photo courtesy of US National Institute of Allergy and Infectious Diseases)
Due to the proofreading ability of the coronavirus (CoV) RNA-dependent RNA polymerase, the evolution of the global SARS-CoV-2 viral population during the current pandemic has been relatively constrained as compared to other endemic RNA viruses that do not possess this ability.
During late 2020, three distinct variants that each possessed a significantly increased amount of amino acid polymorphisms were identified in association with spikes in cases of COVID-19 in the United Kingdom (variant B.1.1.7), South Africa (variant B.1.351), and Brazil (variant B.1.1.248). These variants all possess the N501Y mutation in the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein, a primary target for neutralizing antibody (NAb) binding.
A collaborating team of scientists led by the Johns Hopkins University School of Medicine (Baltimore, MD, USA) analyzed blood cell samples from 30 people who had contracted and recovered from coronavirus disease 2019 (COVID-19) prior to the emergence of virus variants. Most (60%) individuals included in the analysis were male and samples were collected a median of 42.5 days (interquartile range, 37.5-48 days) from initial diagnosis. The team aimed to determine whether CD8+ T cells in the blood could still recognize the three main SARS-CoV-2 variants. Peripheral blood mononuclear cell (PBMC) samples from the patients were collected and examined across six different human leukocyte antigen (HLA) haplotypes (HLAA*01:01, HLA-A*02:01, HLA-A03:01, HLA-A*11:01, HLA-A*24:02 and HLA-B*07:02).
A multiplexed peptide-MHC tetramer staining approach permitted the screening of 408 potential SARS-CoV-2 candidate epitopes for CD8+ T cell recognition. T cells were also evaluated using a 28-marker phenotypic panel (Immunoscape Pte. Ltd, Singapore). A total of 52 unique epitope responses were found and were directed against several structural and non-structural viral proteins. For controls, CD8+ T cells were probed for reactivity for up to 20 different SARS-CoV-2-unrelated control peptides per HLA (Adenovirus-, CMV-, EBV-, Influenza-, and MART-1-derived epitopes).
The team reported that only one mutation found in the B.1.351-Spike overlapped with a previously identified epitope (1/52), suggesting that virtually all anti-SARS-CoV-2 CD8+ T-cell responses should recognize these newly described variants. This mutation is the D80A mutation in the Spike protein, and occurs in the third residue of the RFDN VLPF epitope. This is a HLA*A24:02-restricted epitope for which a CD8+ T cell response was detected in 1/5 HLA*A24:02+ individuals, and at a low frequency (0.005 of total CD8+ T-cells), indicating this is not a high-prevalence epitope.
The authors concluded that their data highlighted the potential significant role of a multi-epitope T cell response in limiting viral escape, and partly mediate protection from disease caused by the SARS-CoV-2 variants. It is important that vaccines used for widespread campaigns generate strong multivalent T-cell responses in addition to neutralizing antibody (Nab) and other humoral responses in order to optimize efficacy against the current SARS-CoV-2 and emerging strains. It will be important to continue to monitor the breadth, magnitude, and durability of the anti-SARS-CoV-2 T cell responses in recovered and vaccinated individuals as part of any assessment to determine if booster vaccinations are needed. The study was published on March 30, 2021 in the journal Open Forum Infectious Diseases.
Related Links:
Johns Hopkins University School of Medicine
Immunoscape Pte. Ltd
During late 2020, three distinct variants that each possessed a significantly increased amount of amino acid polymorphisms were identified in association with spikes in cases of COVID-19 in the United Kingdom (variant B.1.1.7), South Africa (variant B.1.351), and Brazil (variant B.1.1.248). These variants all possess the N501Y mutation in the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein, a primary target for neutralizing antibody (NAb) binding.
A collaborating team of scientists led by the Johns Hopkins University School of Medicine (Baltimore, MD, USA) analyzed blood cell samples from 30 people who had contracted and recovered from coronavirus disease 2019 (COVID-19) prior to the emergence of virus variants. Most (60%) individuals included in the analysis were male and samples were collected a median of 42.5 days (interquartile range, 37.5-48 days) from initial diagnosis. The team aimed to determine whether CD8+ T cells in the blood could still recognize the three main SARS-CoV-2 variants. Peripheral blood mononuclear cell (PBMC) samples from the patients were collected and examined across six different human leukocyte antigen (HLA) haplotypes (HLAA*01:01, HLA-A*02:01, HLA-A03:01, HLA-A*11:01, HLA-A*24:02 and HLA-B*07:02).
A multiplexed peptide-MHC tetramer staining approach permitted the screening of 408 potential SARS-CoV-2 candidate epitopes for CD8+ T cell recognition. T cells were also evaluated using a 28-marker phenotypic panel (Immunoscape Pte. Ltd, Singapore). A total of 52 unique epitope responses were found and were directed against several structural and non-structural viral proteins. For controls, CD8+ T cells were probed for reactivity for up to 20 different SARS-CoV-2-unrelated control peptides per HLA (Adenovirus-, CMV-, EBV-, Influenza-, and MART-1-derived epitopes).
The team reported that only one mutation found in the B.1.351-Spike overlapped with a previously identified epitope (1/52), suggesting that virtually all anti-SARS-CoV-2 CD8+ T-cell responses should recognize these newly described variants. This mutation is the D80A mutation in the Spike protein, and occurs in the third residue of the RFDN VLPF epitope. This is a HLA*A24:02-restricted epitope for which a CD8+ T cell response was detected in 1/5 HLA*A24:02+ individuals, and at a low frequency (0.005 of total CD8+ T-cells), indicating this is not a high-prevalence epitope.
The authors concluded that their data highlighted the potential significant role of a multi-epitope T cell response in limiting viral escape, and partly mediate protection from disease caused by the SARS-CoV-2 variants. It is important that vaccines used for widespread campaigns generate strong multivalent T-cell responses in addition to neutralizing antibody (Nab) and other humoral responses in order to optimize efficacy against the current SARS-CoV-2 and emerging strains. It will be important to continue to monitor the breadth, magnitude, and durability of the anti-SARS-CoV-2 T cell responses in recovered and vaccinated individuals as part of any assessment to determine if booster vaccinations are needed. The study was published on March 30, 2021 in the journal Open Forum Infectious Diseases.
Related Links:
Johns Hopkins University School of Medicine
Immunoscape Pte. Ltd
Latest Immunology News
- Finger-Prick Lateral Flow Test Detects Sepsis Biomarkers at Point of Care
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
- Immune Signature Identified in Treatment-Resistant Myasthenia Gravis
- New Biomarker Predicts Chemotherapy Response in Triple-Negative Breast Cancer
- Blood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
- Whole-Genome Sequencing Approach Identifies Cancer Patients Benefitting From PARP-Inhibitor Treatment
- Ultrasensitive Liquid Biopsy Demonstrates Efficacy in Predicting Immunotherapy Response
- Blood Test Could Identify Colon Cancer Patients to Benefit from NSAIDs
Channels
Clinical Chemistry
view channel
Blood Test Detects Testicular Cancer Missed by Standard Markers
Testicular cancer most often affects adolescents and young adults and is highly treatable when found early. Diagnosis can be difficult when tumors do not produce sufficient levels of standard blood-based... Read more
Routine Blood Tests Identify Biomarkers Linked to PTSD
Post-traumatic stress disorder (PTSD) is associated with a range of chronic physical health conditions and affects multiple organ systems. Clinical laboratories routinely measure blood analytes that reflect... Read moreMolecular Diagnostics
view channel
CRISPR-Based Test Identifies Multiple Respiratory Viruses Simultaneously
Respiratory virus co-circulation complicates differential diagnosis, as overlapping symptoms can obscure etiology. Multiplex testing typically depends on multiple enzymes or fluorophores and multistep... Read more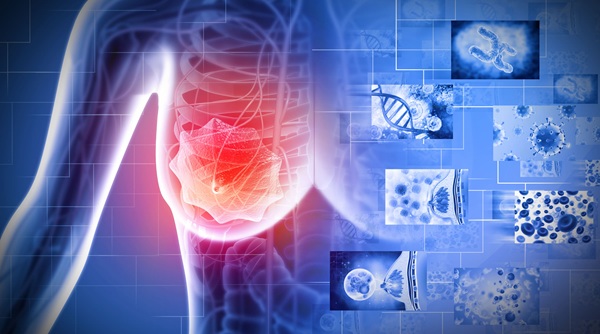
Multi-Omic Assay Predicts Recurrence and Radiation Benefit in Early Breast Cancer
Early-stage invasive breast cancer is frequently managed with breast-conserving surgery followed by adjuvant radiation therapy, but the magnitude of benefit from radiation varies among patients.... Read more
Portable Test Detects Tuberculosis from Tongue Swabs in 30 Minutes
Despite decades of effective drug regimens, tuberculosis remains the leading cause of death from an infectious disease, driven in part by limited access to accurate, rapid testing. Conventional diagnostics... Read more
Blood Test Receives FDA Breakthrough Status to Differentiate Schizophrenia and Bipolar Disorder
Differentiating schizophrenia from bipolar I disorder in symptomatic patients is challenging because early presentations often overlap. Current diagnostic pathways rely heavily on subjective assessments... Read moreHematology
view channel
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read more
Blood Test Enables Early Detection of Multiple Myeloma Relapse
Bone marrow biopsies remain central to diagnosing and monitoring multiple myeloma, yet the procedure is painful, invasive, and often repeated over time. Older patients—who represent most new cases—can... Read moreMicrobiology
view channel
Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
Tuberculosis remains a major global health challenge and continues to drive significant morbidity and mortality. The World Health Organization’s 2024 global report cites it as the leading cause of death... Read more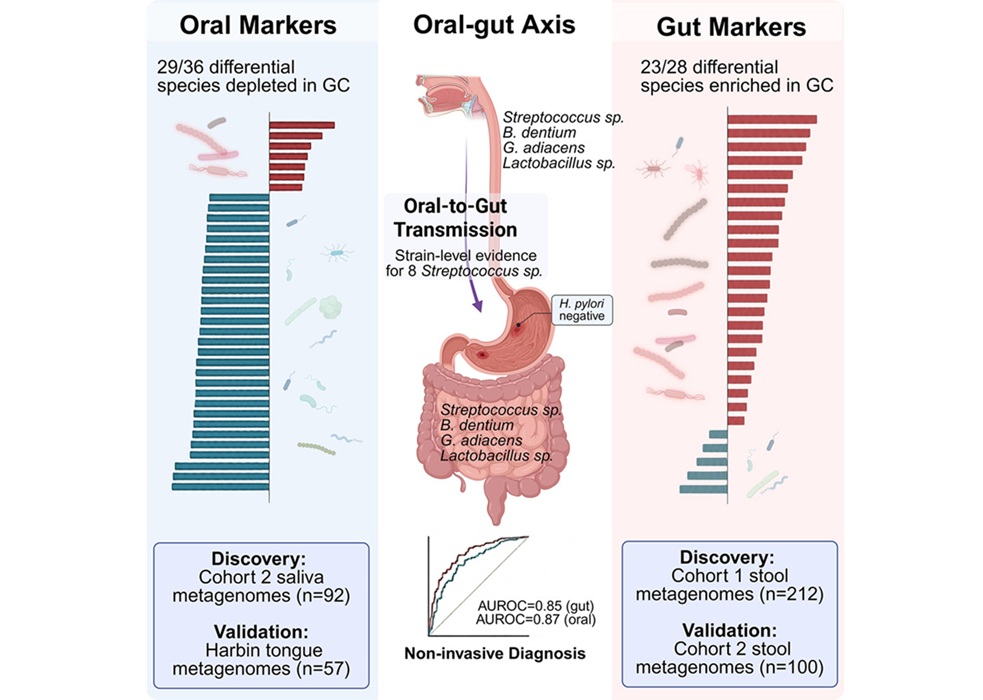
Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
Early detection of gastric cancer could be advanced by scalable screening strategies using minimally invasive sampling. Saliva collection is noninvasive and cost-effective, supporting wider adoption... Read morePathology
view channel
New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease
Diabetic kidney disease is a leading cause of chronic kidney disease and end-stage kidney disease, affecting 20%–40% of people with diabetes and more than 107 million individuals worldwide as of 2021.... Read more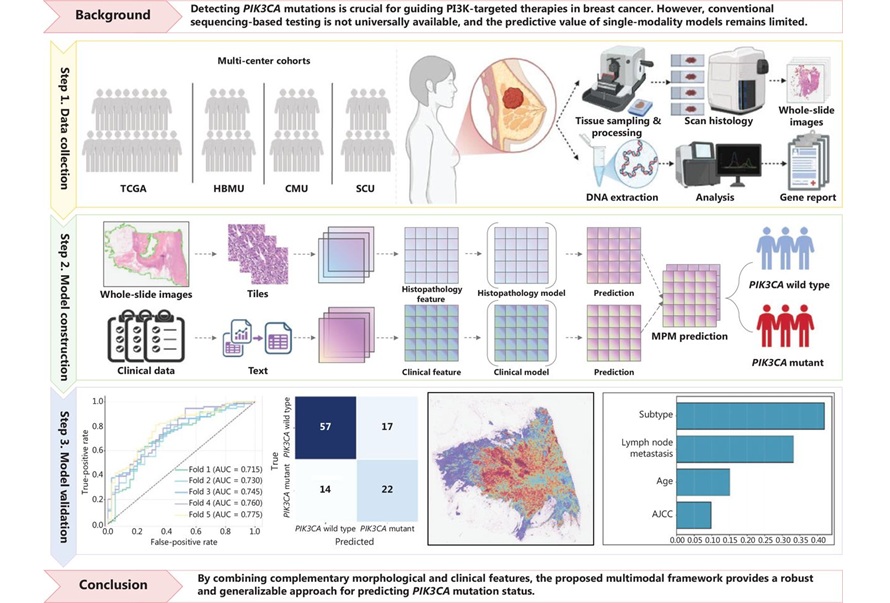
Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
PIK3CA mutations are key biomarkers for selecting phosphoinositide 3-kinase (PI3K)–targeted therapies in breast cancer, yet access to molecular testing can be inconsistent and costly. Conventional polymerase... Read moreTechnology
view channel
Microfluidic Single-Cell Assay Predicts Breast Cancer Risk
Risk stratification for breast cancer remains imprecise, as population-based models and breast density can over- or underestimate individual risk, potentially leading to over- or under-screening.... Read more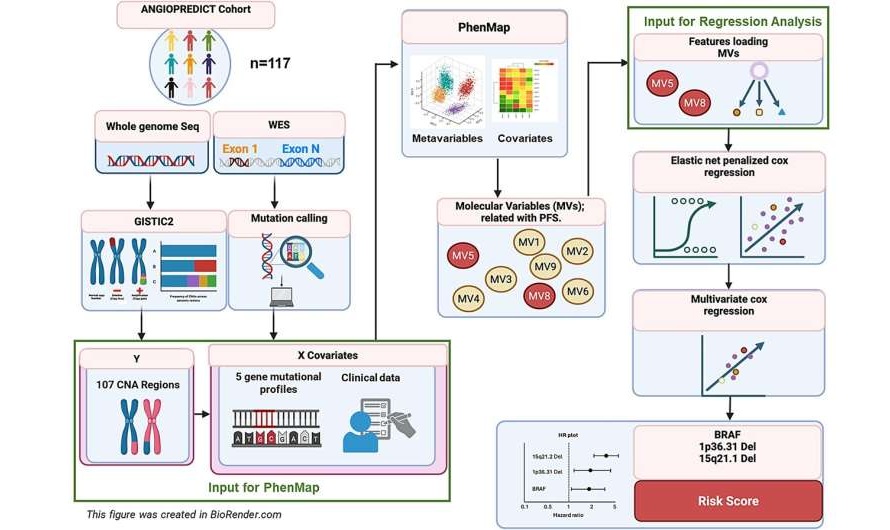




.jpg)