New Method Simplifies Preparation of Tumor Genomic DNA Libraries
|
By LabMedica International staff writers Posted on 28 Oct 2019 |
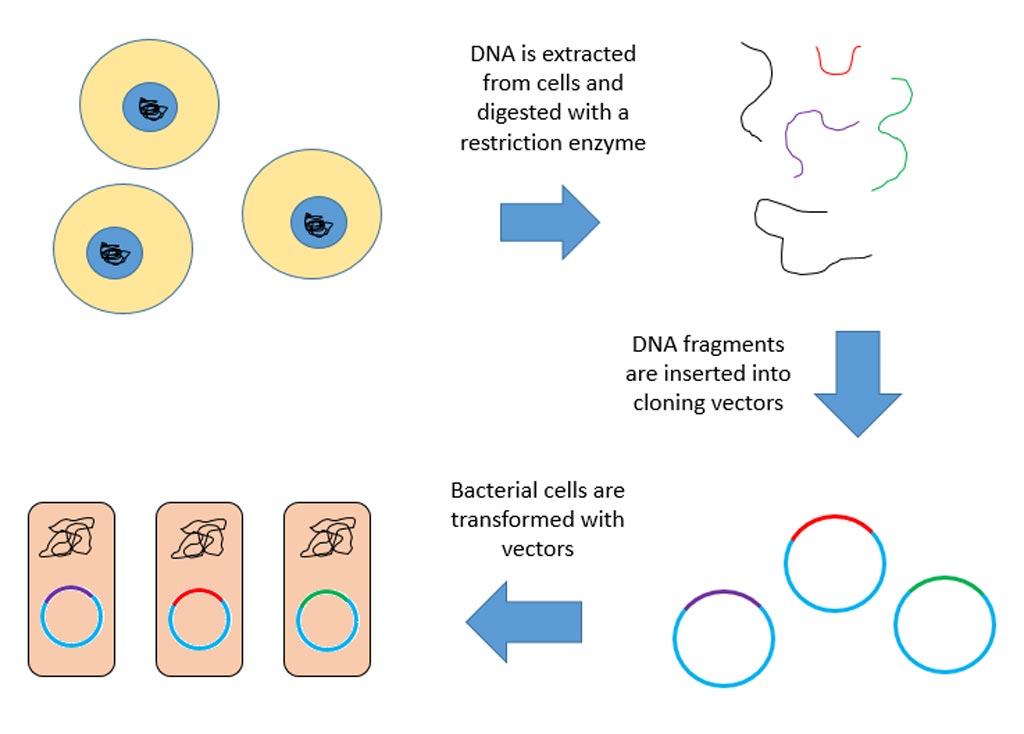
Image: A diagram of construction of a genomic library (Photo courtesy of Wikimedia Commons).
A method has been described that simplifies preparation of tumor genomic DNA libraries by employing restriction enzymes and in vitro transcription to barcode and amplify genomic DNA prior to library construction.
In general, a genomic library is the collection of the total genomic DNA from a single organism. The DNA is stored in a population of identical vectors, each containing a different insert of DNA. In order to construct a genomic library, the organism's DNA is extracted from cells and then digested with a restriction enzyme to cut the DNA into fragments of a specific size. The fragments are then inserted into the vector using DNA ligase.
Copy number alterations or variations are common features of cancer cells. Within the same tumor, cells belonging to different anatomical areas of the tumor may carry different variations. Tumors with many variations are typically very aggressive and tend to be resistant to treatment.
Current strategies for massively parallel sequencing of tumor genomic DNA mainly rely on library indexing in the final steps of library preparation. This procedure is costly and time-consuming, since a library must be generated separately for each sample. Furthermore, whole-genome amplification requires intact DNA and thus is problematic in fixed tissue samples, in particular formalin-fixed, paraffin-embedded (FFPE) specimens, which still represent a cornerstone in pathology.
To overcome these limitations, investigators at the Karolinska Institutet (Solna, Sweden) developed a method, which they named CUTseq, that combined restriction endonucleases with in vitro transcription (IVT) to construct highly multiplexed DNA libraries for reduced representation genome sequencing of multiple samples in parallel.
The investigators showed that CUTseq could be used to barcode gDNA extracted from both non-fixed and fixed samples, including old archival FFPE tissue sections. They benchmarked CUTseq by comparing it with a widely used method of DNA library preparation and demonstrated that CUTseq could be used for reduced representation genome and exome sequencing, enabling reproducible DNA copy number profiling and single-nucleotide variant (SNV) calling in both cell and low-input FFPE tissue samples.
The investigators demonstrated an application of CUTseq for assessing genetic differences within tumors by profiling DNA copy number levels in multiple small regions of individual FFPE tumor sections. Finally, they described a workflow for rapid and cost-effective preparation of highly multiplexed CUTseq libraries, which could be applied in the context of high-throughput genetic screens and for cell line authentication.
"I expect that CUTseq will find many useful applications in cancer diagnostics," said senior author Dr. Nicola Crosetto, senior researcher in medical biochemistry and biophysics at the Karolinska Institutet. "Multi-region tumor sequencing is going to be increasingly used in the diagnostic setting, in order to identify patients with highly heterogeneous tumors that need to be treated more aggressively. I believe that our method can play a leading role here."
The study was published in the October 18, 2019, online edition of the journal Nature Communications.
Related Links:
Karolinska Institutet
In general, a genomic library is the collection of the total genomic DNA from a single organism. The DNA is stored in a population of identical vectors, each containing a different insert of DNA. In order to construct a genomic library, the organism's DNA is extracted from cells and then digested with a restriction enzyme to cut the DNA into fragments of a specific size. The fragments are then inserted into the vector using DNA ligase.
Copy number alterations or variations are common features of cancer cells. Within the same tumor, cells belonging to different anatomical areas of the tumor may carry different variations. Tumors with many variations are typically very aggressive and tend to be resistant to treatment.
Current strategies for massively parallel sequencing of tumor genomic DNA mainly rely on library indexing in the final steps of library preparation. This procedure is costly and time-consuming, since a library must be generated separately for each sample. Furthermore, whole-genome amplification requires intact DNA and thus is problematic in fixed tissue samples, in particular formalin-fixed, paraffin-embedded (FFPE) specimens, which still represent a cornerstone in pathology.
To overcome these limitations, investigators at the Karolinska Institutet (Solna, Sweden) developed a method, which they named CUTseq, that combined restriction endonucleases with in vitro transcription (IVT) to construct highly multiplexed DNA libraries for reduced representation genome sequencing of multiple samples in parallel.
The investigators showed that CUTseq could be used to barcode gDNA extracted from both non-fixed and fixed samples, including old archival FFPE tissue sections. They benchmarked CUTseq by comparing it with a widely used method of DNA library preparation and demonstrated that CUTseq could be used for reduced representation genome and exome sequencing, enabling reproducible DNA copy number profiling and single-nucleotide variant (SNV) calling in both cell and low-input FFPE tissue samples.
The investigators demonstrated an application of CUTseq for assessing genetic differences within tumors by profiling DNA copy number levels in multiple small regions of individual FFPE tumor sections. Finally, they described a workflow for rapid and cost-effective preparation of highly multiplexed CUTseq libraries, which could be applied in the context of high-throughput genetic screens and for cell line authentication.
"I expect that CUTseq will find many useful applications in cancer diagnostics," said senior author Dr. Nicola Crosetto, senior researcher in medical biochemistry and biophysics at the Karolinska Institutet. "Multi-region tumor sequencing is going to be increasingly used in the diagnostic setting, in order to identify patients with highly heterogeneous tumors that need to be treated more aggressively. I believe that our method can play a leading role here."
The study was published in the October 18, 2019, online edition of the journal Nature Communications.
Related Links:
Karolinska Institutet
Latest BioResearch News
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
- Study Links Abnormal Gene Splicing to Treatment Response in Metastatic Kidney Cancer
Channels
Clinical Chemistry
view channel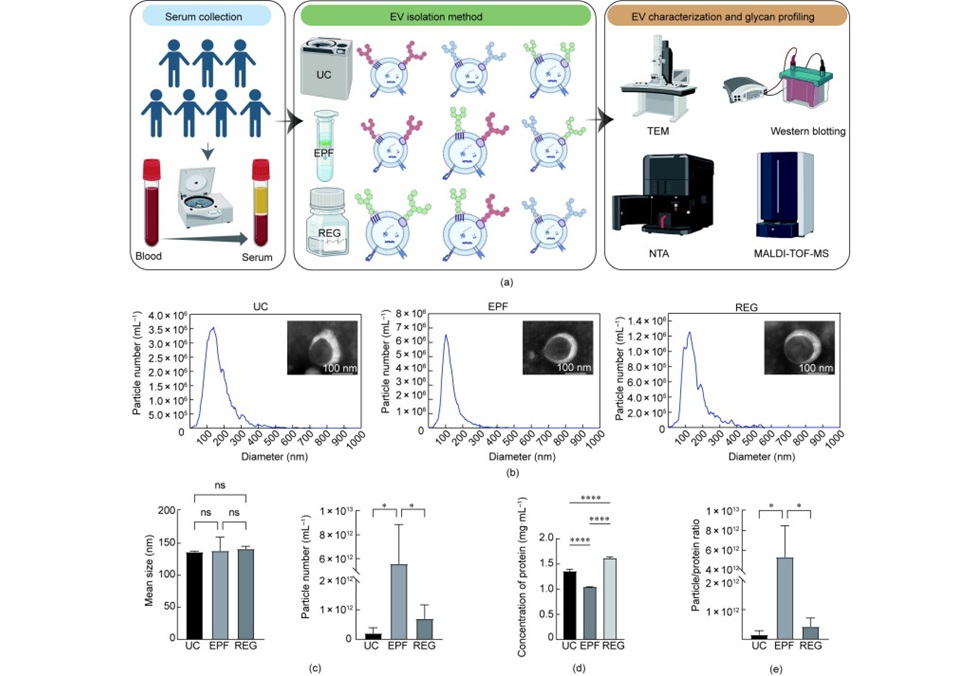
Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
Childhood epilepsy remains a major neurological disorder with unmet needs for accurate, non-invasive biomarkers, as conventional tests such as electroencephalography and neuroimaging can have limited sensitivity... Read more
Blood-Based Sensor Detects Early Signs of Alzheimer’s and Parkinson’s
Alzheimer’s disease and Parkinson’s disease are increasing as populations age, yet diagnosis remains largely symptom-driven and often occurs after irreversible brain damage has begun. Earlier detection,... Read moreMolecular Diagnostics
view channel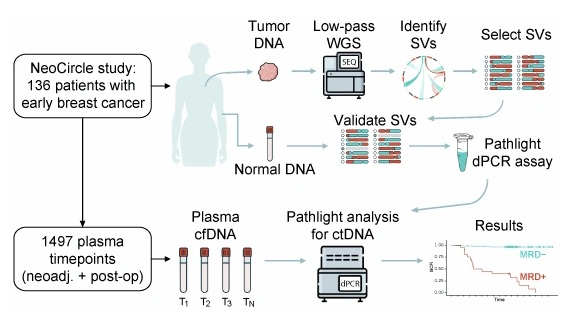
Ultrasensitive MRD Blood Test Detects Early Breast Cancer Recurrence
SAGA Diagnostics (Morrisville, NC, USA), a company specializing in tumor-informed, blood-based cancer detection and precision medicine, announced the publication of a new study evaluating its Pathlight... Read more
Position Statements Outline Evidence Standards for Multi-Cancer Detection Tests
Cancer screening is intended to reduce mortality, but policy decisions often depend on early indicators that may not fully reflect true survival benefit. The emergence of blood-based tests capable of detecting... Read moreHematology
view channel
Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
Iron deficiency in school-age children can affect brain development, learning, growth, and physical performance, yet early deficiency may be missed when screening focuses mainly on anemia.... Read more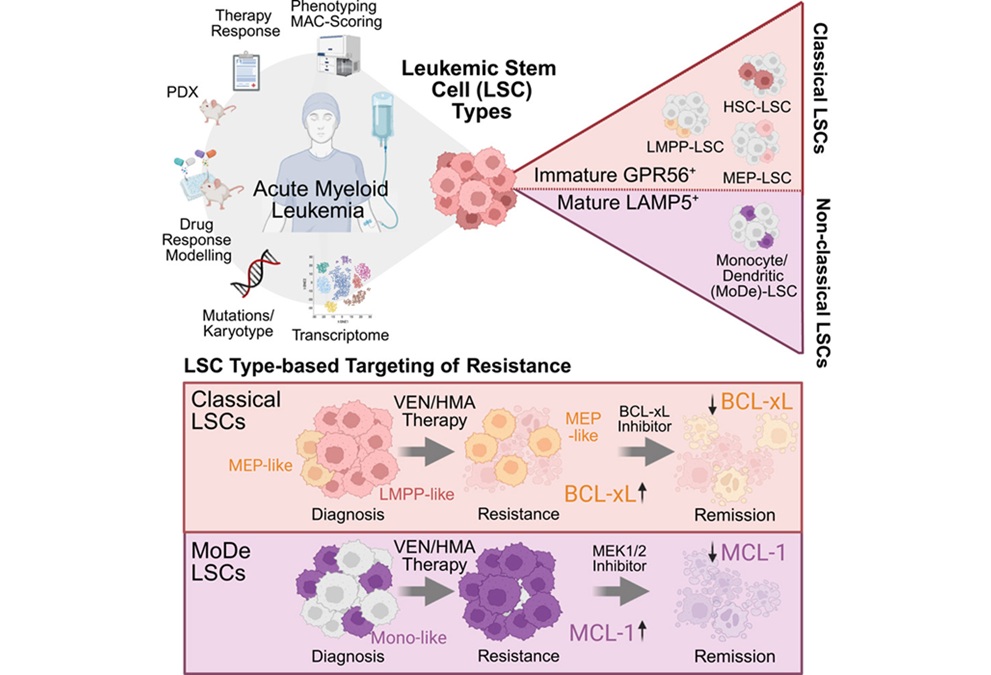
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read moreImmunology
view channel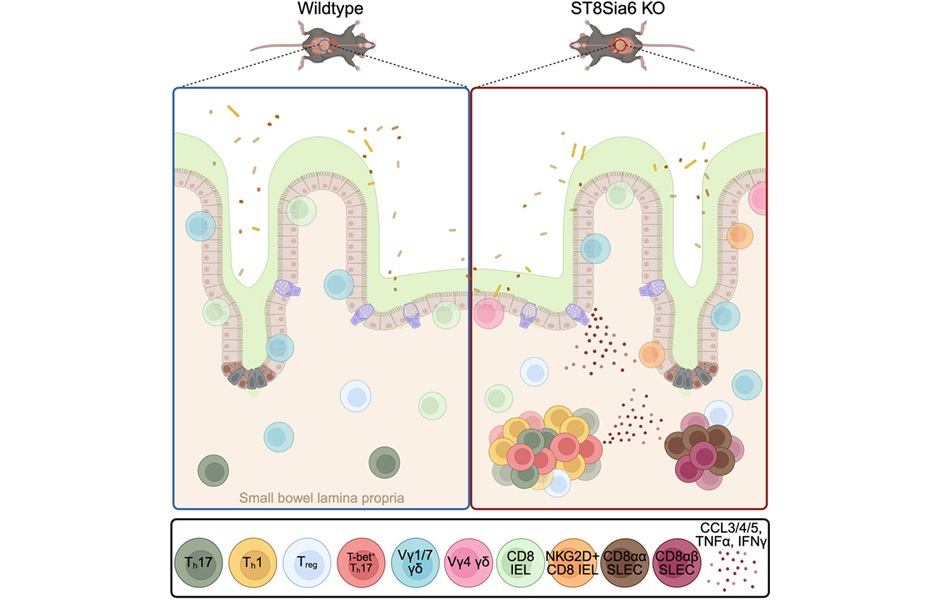
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read more
Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
Lung transplant recipients face some of the highest rates of acute cellular rejection, and routine surveillance often relies on repeated surgical biopsies. These procedures can cause complications such... Read moreMicrobiology
view channel
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read more
New AMR Assay Supports Rapid Infection Control Screening in Hospitals
As antimicrobial resistance spreads worldwide, healthcare-associated infections are placing a growing burden on hospitals, increasing the need for faster and broader diagnostic solutions.... Read morePathology
view channel
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read more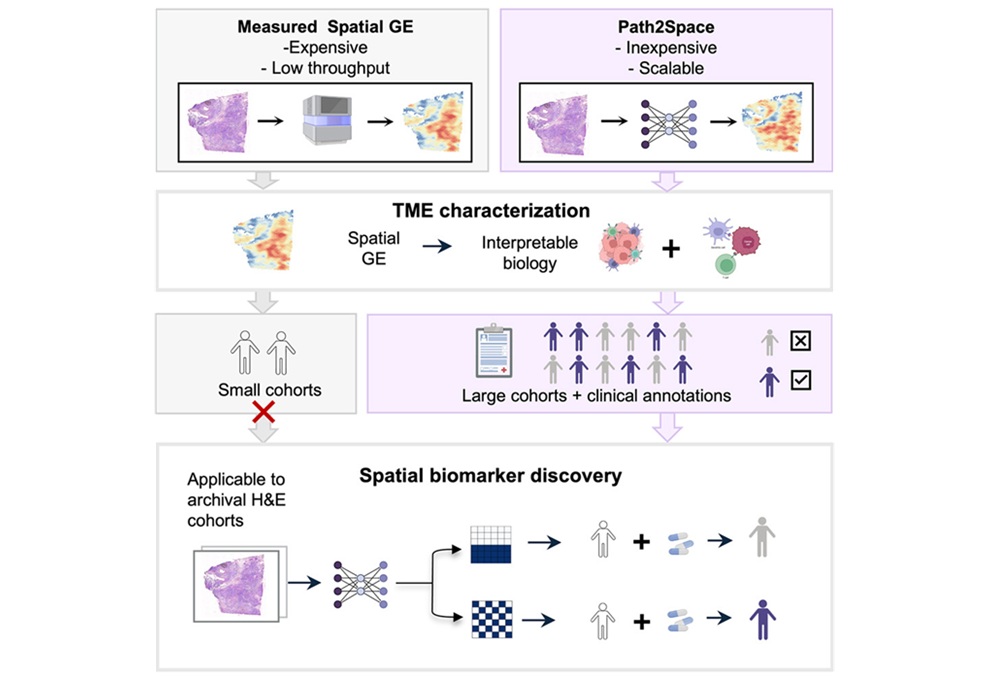
Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
Gene expression profiling can inform tumor biology and treatment selection, but spatial assays remain costly and time-consuming. Results can take weeks and cost thousands of dollars, limiting large-scale... Read moreTechnology
view channel
AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
Clinical laboratories and research groups face increasingly complex molecular workflows and expanding technical documentation spread across multiple systems. Fragmented digital tools can slow experiment... Read more
AI Tool Automates Validation of Laboratory Software Configuration Changes
Regulated laboratories face heavy documentation and requalification demands when software configurations change, slowing improvements and discouraging beneficial updates. A new capability now automates... Read moreIndustry
view channel
New Distribution Agreement Expands Access to CE-Marked Precision Oncology Assays
Eurobio Scientific (Les Ulis, France) has signed a distribution agreement with Canhelp Genomics (Hangzhou, China) to broaden availability of the Canhelp‑UCa and Canhelp‑Origin assays. The agreement extends... Read more