One-Tube RNA Ligation and Amplification Method for Rapid Detection of Drug Resistant HIV
|
By LabMedica International staff writers Posted on 28 Oct 2015 |
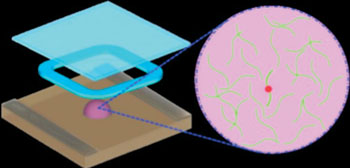
Image: A schematic diagram of a new method that rapidly analyzes the RNA (green strands) of HIV for mutations (red dot) that convey drug resistance. The system does not require transcription of RNA to DNA, as current technologies do, and works within one solution (purple droplet) (Photo courtesy of Dr. Lei Zhang, Brown University).
By not requiring transcription of RNA to DNA, a novel one-tube method allows the rapid detection of drug resistant strains of HIV (human immunodeficiency virus).
In order to detect point mutations in RNA retroviruses, conventional ligase-mediated approaches require the reverse transcription of viral RNA genomes into DNA before separate ligation and amplification steps can be carried out.
To simplify this process, investigators at Brown University (Providence, RI, USA) developed one-step ligation on RNA amplification (LRA) method for the direct detection of RNA point mutations. The system operates directly on viral RNA rather than requiring extra, potentially error-prone steps to examine DNA derived from RNA. In a single tube, the system first combines two engineered probes (ligation). If a mutation is present, it then makes many copies of those combined probes (amplification) for detection.
The investigators used this technique for the detection of a common, clinically relevant HIV-1 reverse transcriptase drug-resistant point mutation, K103N, and compared it with allele-specific PCR and pyrosequencing methodology.
They reported in the November 2015 issue of the Journal of Molecular Diagnostics that the LRA test was sensitive enough to detect the K103N mutation in concentrations as low as one mutant per 10,000 strands of normal viral RNA. The LRA test required about two hours while the alternative technologies took as long as eight hours.
"LRA (ligation on RNA amplification) uniquely optimizes two enzymatic reactions—RNA-based ligation, and quantitative PCR (polymerase chain reaction) amplification—into a single system," said senior author Dr. Anubhav Tripathi, professor of engineering at Brown University. "Each HIV contains about 10,000 nucleotides, or building blocks, in its genetic material, and a drop of blood from a patient with resistant HIV can contain thousands to millions of copies of HIV. To find that one virus, out of thousands to millions, which is mutated at just a single nucleotide is like finding a needle in a haystack."
So far the LRA test has been shown to work on RNA that was derived from laboratory HIV strains, but it has not yet been applied to samples from circulating viruses from AIDS patients.
Related Links:
Brown University
In order to detect point mutations in RNA retroviruses, conventional ligase-mediated approaches require the reverse transcription of viral RNA genomes into DNA before separate ligation and amplification steps can be carried out.
To simplify this process, investigators at Brown University (Providence, RI, USA) developed one-step ligation on RNA amplification (LRA) method for the direct detection of RNA point mutations. The system operates directly on viral RNA rather than requiring extra, potentially error-prone steps to examine DNA derived from RNA. In a single tube, the system first combines two engineered probes (ligation). If a mutation is present, it then makes many copies of those combined probes (amplification) for detection.
The investigators used this technique for the detection of a common, clinically relevant HIV-1 reverse transcriptase drug-resistant point mutation, K103N, and compared it with allele-specific PCR and pyrosequencing methodology.
They reported in the November 2015 issue of the Journal of Molecular Diagnostics that the LRA test was sensitive enough to detect the K103N mutation in concentrations as low as one mutant per 10,000 strands of normal viral RNA. The LRA test required about two hours while the alternative technologies took as long as eight hours.
"LRA (ligation on RNA amplification) uniquely optimizes two enzymatic reactions—RNA-based ligation, and quantitative PCR (polymerase chain reaction) amplification—into a single system," said senior author Dr. Anubhav Tripathi, professor of engineering at Brown University. "Each HIV contains about 10,000 nucleotides, or building blocks, in its genetic material, and a drop of blood from a patient with resistant HIV can contain thousands to millions of copies of HIV. To find that one virus, out of thousands to millions, which is mutated at just a single nucleotide is like finding a needle in a haystack."
So far the LRA test has been shown to work on RNA that was derived from laboratory HIV strains, but it has not yet been applied to samples from circulating viruses from AIDS patients.
Related Links:
Brown University
Latest BioResearch News
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
- Study Links Abnormal Gene Splicing to Treatment Response in Metastatic Kidney Cancer
- Research Reveals How Some Aplastic Anemia Patients Recover Bone Marrow Function
- New Molecular Insights Support Diagnosis of Hodgkin Lymphoma
- Epigenetic Signals and Blood Markers Aid Chronic Fatigue Syndrome Diagnosis
- Microenvironment Biomarkers Could Enable Early Lung Cancer Detection
- Study Identifies Protein Changes Driving Immunotherapy Resistance in Multiple Myeloma
- Genetic Analysis Identifies BRCA-Linked Risks Across Multiple Cancers
- Study Identifies Hidden B-Cell Mutations in Autoimmune Disease
- Single-Cell Method Measures RNA and Proteins to Reveal Immune Responses
- Study Links Midlife Vitamin D to Lower Tau in Alzheimer's
- International Consensus Standardizes Tumor Microbiota Detection and Reporting
- Common Metablolic Enzyme Could Predict Response to Cancer Immunotherapy
- Newly Identfied Genetic Variants in MND Support Prognosis and Family Testing
- Innate Immunity Variants Associated With Earlier Breast Cancer in BRCA1 Carriers
Channels
Clinical Chemistry
view channel
Ultrasensitive Test Detects Key Biomarker of Frontotemporal Dementia Subtype
Dementia affects more than 57 million people worldwide and is projected to nearly double within two decades, straining health systems and families. While biomarkers now enable accurate identification of... Read more
Routine Blood Tests Years Before Pregnancy Could Identify Preeclampsia Risk
High blood pressure during pregnancy is common and can progress to pre-eclampsia, making close monitoring at antenatal visits essential. However, most risk assessment begins only after pregnancy has started.... Read moreMolecular Diagnostics
view channel
Liquid Biopsy Biomarkers Distinguish Inflammatory Breast Cancer and Support Monitoring
Inflammatory breast cancer is among the most aggressive forms of breast malignancy and remains challenging to diagnose and monitor. Obtaining tumor tissue can be difficult, and standard genome and RNA... Read more
Blood Test Maps Tumor Microenvironment to Predict Immunotherapy Response
Immunotherapy has transformed cancer care, yet durable benefit remains limited to a subset of patients, and clinicians still lack reliable tools to predict response before treatment begins.... Read more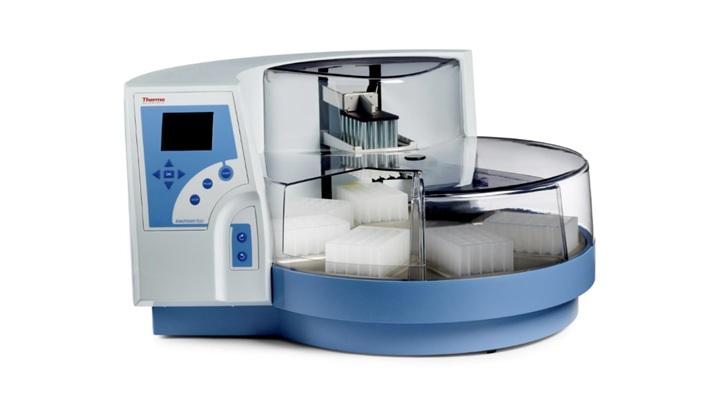
Multiplex Respiratory Panel Integrates Automated Extraction to Streamline High-Volume Testing
Respiratory infections drive heavy testing volumes in clinical laboratories, where accurate, timely results across multiple pathogens are essential. Many labs are seeking to streamline workflows and increase... Read moreHematology
view channel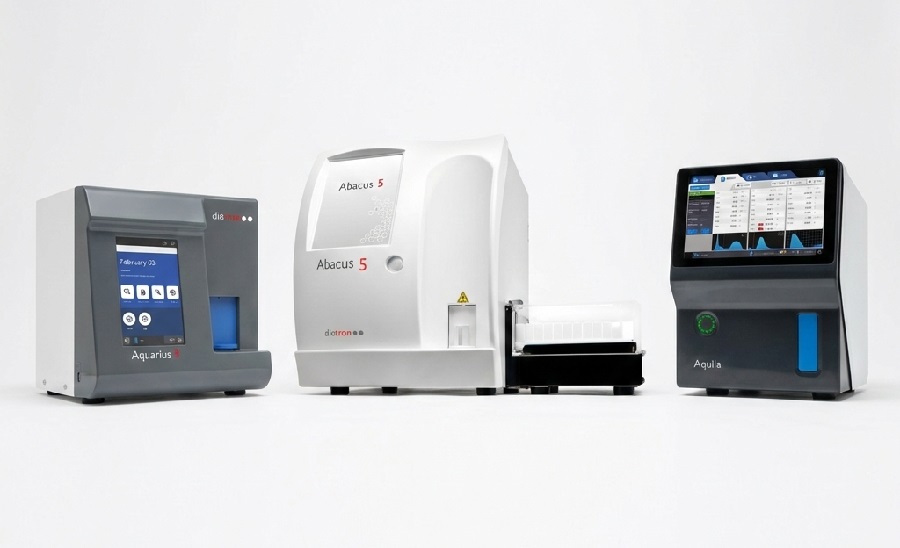
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read more
Blood Test Enables Early Detection of Multiple Myeloma Relapse
Bone marrow biopsies remain central to diagnosing and monitoring multiple myeloma, yet the procedure is painful, invasive, and often repeated over time. Older patients—who represent most new cases—can... Read moreImmunology
view channel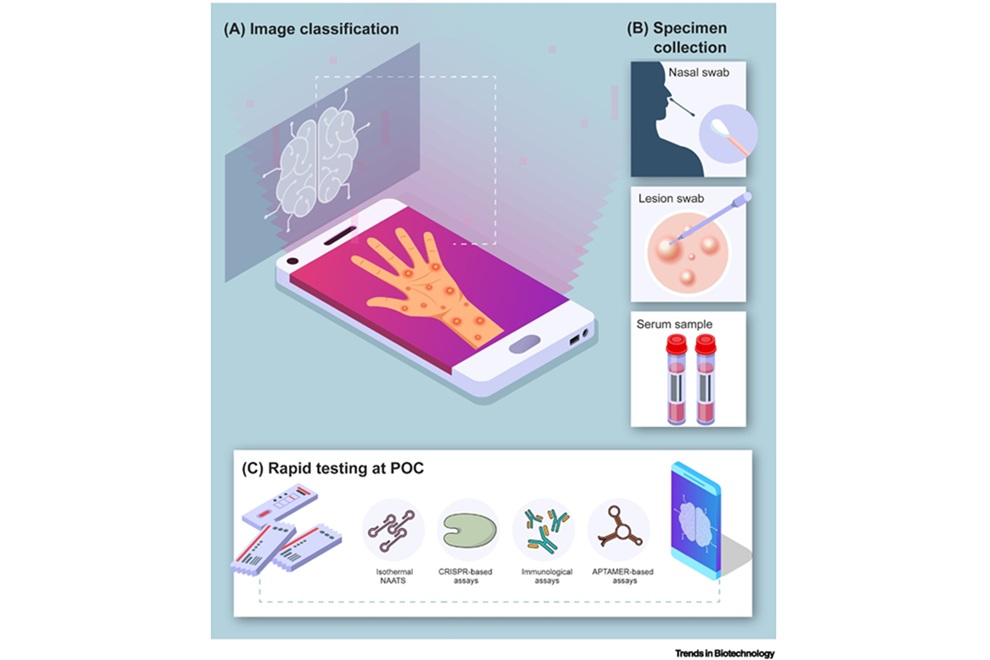
Point-of-Care Tests Could Expand Access to Mpox Diagnosis
Mpox outbreaks in non-endemic regions have underscored the need for rapid, accessible diagnostics to limit transmission. Polymerase chain reaction (PCR) remains the clinical reference, yet it depends on... Read more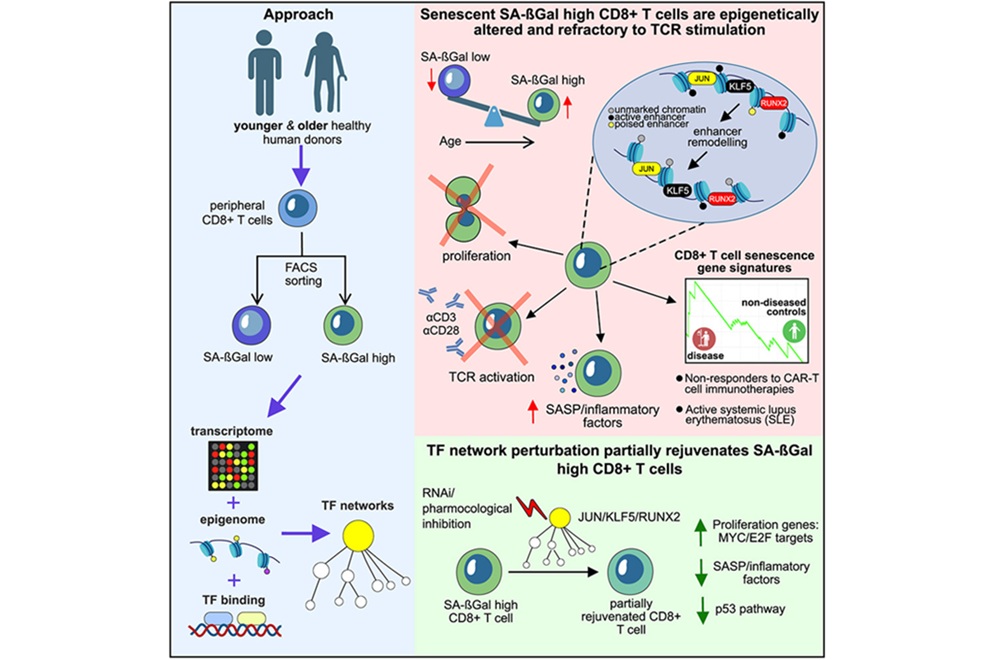
T-Cell Senescence Profiling May Predict CAR T Responses
Chimeric antigen receptor (CAR) T-cell therapy can deliver striking, durable remissions, yet many patients experience minimal or no benefit. The quality of patient-derived cytotoxic T lymphocytes used... Read moreMicrobiology
view channel
Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
Tuberculosis remains a major global health challenge and continues to drive significant morbidity and mortality. The World Health Organization’s 2024 global report cites it as the leading cause of death... Read more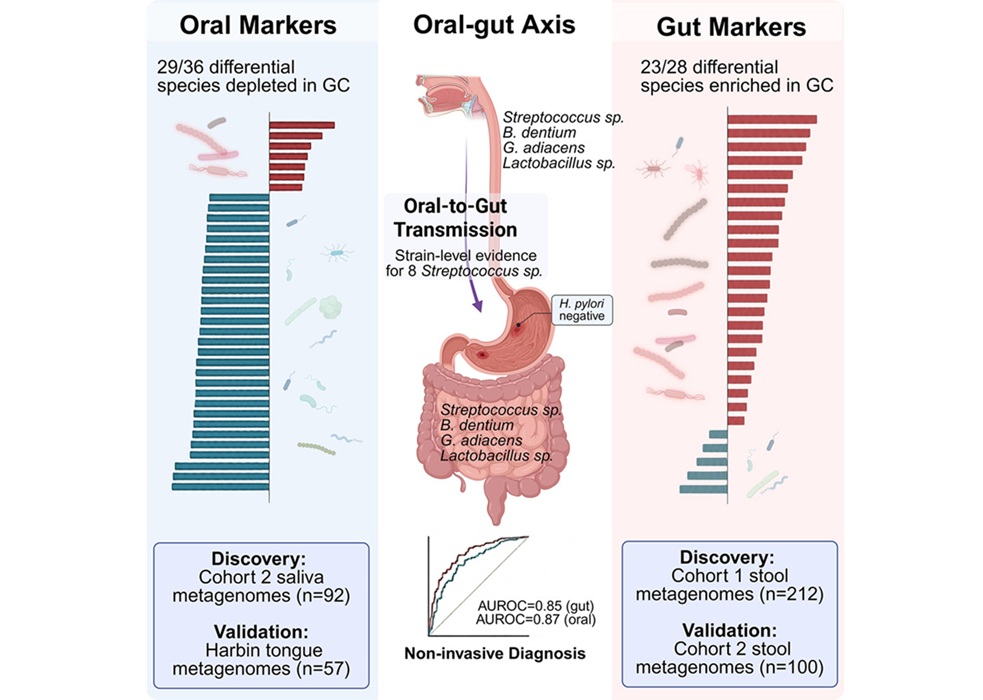
Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
Early detection of gastric cancer could be advanced by scalable screening strategies using minimally invasive sampling. Saliva collection is noninvasive and cost-effective, supporting wider adoption... Read morePathology
view channel
FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
Risk assessment at diagnosis is central to guiding therapy for early-stage, hormone receptor-positive, human epidermal growth factor receptor 2-negative (HR+/HER2-) invasive breast cancer, where overtreatment... Read more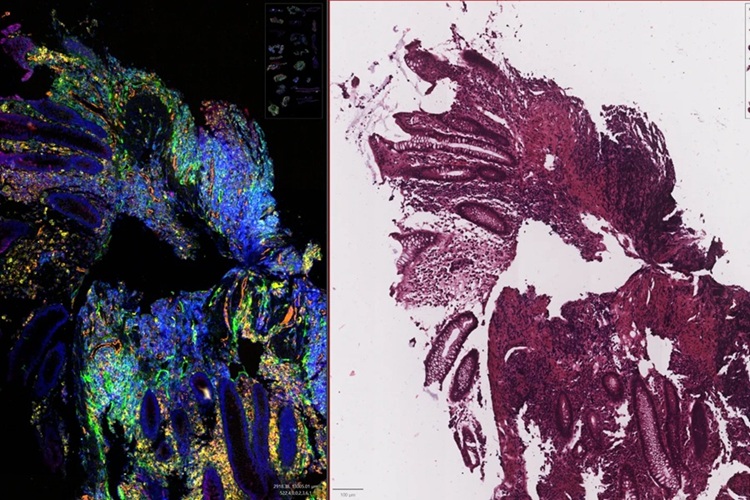
New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
Pathologists worldwide rely on hematoxylin and eosin (H&E) slides to examine tissue architecture, yet these stains do not reveal the underlying molecular activity that often drives disease.... Read moreTechnology
view channel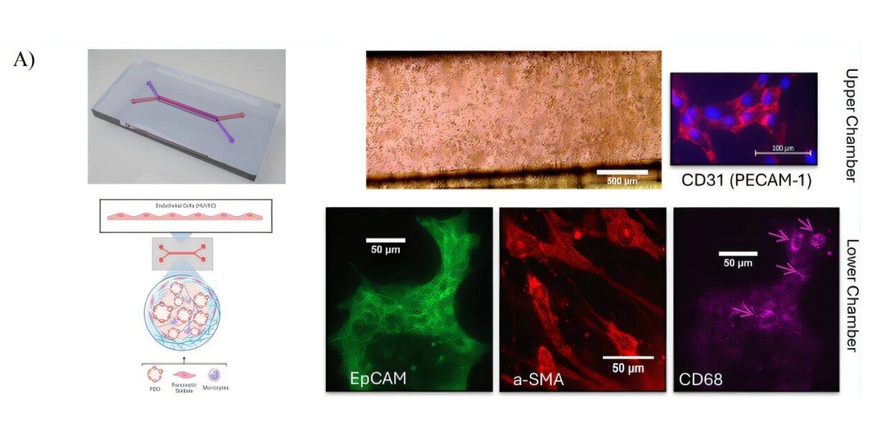
Tumor-on-a-Chip Platform Models Pancreatic Cancer Treatment Response
Pancreatic cancer remains one of the hardest malignancies to treat because tumors are embedded within a dense microenvironment that shapes growth and therapy response. Standard laboratory models often... Read more
New Platform Captures Extracellular Vesicles for Early Cancer Detection
Early diagnosis remains the most effective way to reduce cancer mortality, yet many screening tools miss disease at its earliest stages. Biomarkers shed by tumors into blood and other fluids can be scarce... Read moreIndustry
view channel
Roche to Acquire PathAI for Up to $1.05 Billion to Strengthen AI Diagnostics Portfolio
Roche has entered into a definitive merger agreement to acquire PathAI, a company focused on digital pathology and artificial intelligence for pathology laboratories and the biopharma industry.... Read more