Methodology Devised to Improve Stem Cell Reprogramming
|
By LabMedica International staff writers Posted on 27 Jan 2015 |
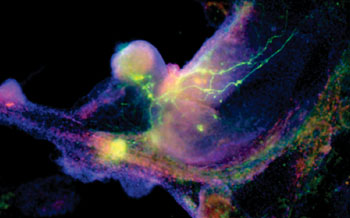
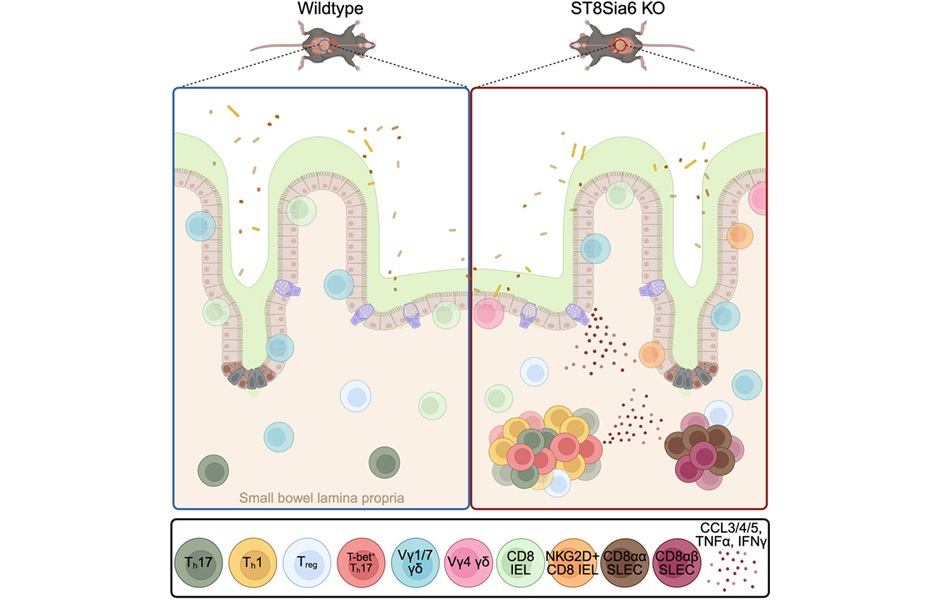
Image: Induced pluripotent stem (iPS) cells, which act very much like embryonic stem cells, are shown growing into heart cells (blue) and nerve cells (green) (Photo courtesy of Gladstone Institutes/Chris Goodfellow).
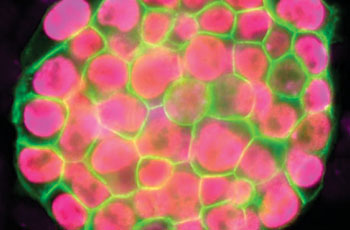
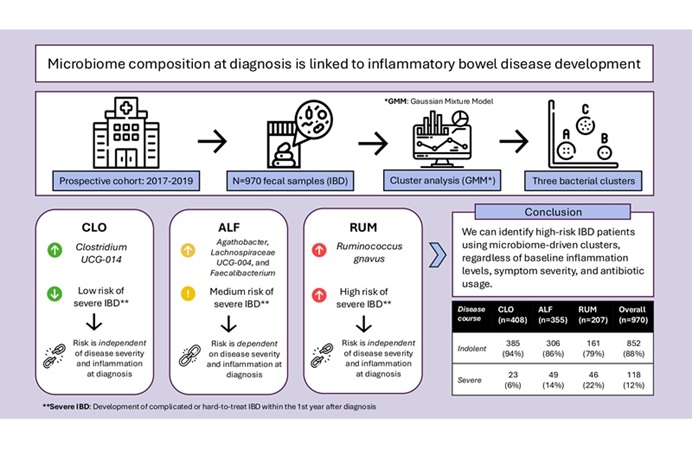
Image: Microscopic view of a colony of induced pluripotent stem cells obtained by reprogramming a specialized cell for two weeks (Photo courtesy of UCLA Broad Stem Cell Research Center/Plath Lab).
In a study that provides scientists with a critical new determination of stem cell development and its role in disease, researchers have established a first-of-its-kind approach that outlines the stages by which specialized cells are reprogrammed into stem cells resembling those found in embryos. The research could have wide ranging, long-term implications in enhancing disease modeling and devising new therapies for patients.
The study, conducted by researchers from the University of California, Los Angeles (UCLA; USA) Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research and led by Dr. Kathrin Plath, a professor of biological chemistry, was published January 2015 in the journal Cell. Induced pluripotent stem cells (iPSCs) are cells that can be generated from adult cells and then, like embryonic stem cells, be directed to become any cell in the human body. Adult cells can also be reprogrammed in the lab to change from a specialized cell back to an iPSC (and thereby becoming a cell similar to that of an embryonic stem cell).
Reprogramming takes one to two weeks and is a mostly inefficient process, with typically less than one percent of the beginning cells effectively becoming an iPSC. The exact stages a cell goes through during the reprogramming process are not well understood. This knowledge is vital, because iPSCs have great potential in the field of regenerative medicine, as they can constantly reproduce and provide a single source of patient-specific cells to replace those lost to injury or disease. They can also be used to create innovative disease models from which new drugs and therapies can be developed.
Vincent Pasque and Jason Tchieu, postdoctoral fellows in Plath’s lab and co-first authors of the study, developed a roadmap of the reprogramming process using detailed time-course analyses. They induced the reprogramming of specialized cells (that could only make more of themselves, and no other cell types), then observed and analyzed on a daily basis or every other day the process of transformation at the single-cell level. The data were gathered and recorded during a time period of up to two weeks.
Dr. Plath’s team found that the changes that happen in cells during reprogramming occur in sequentially, and that notably, the stages of the sequence were the same across the diverse reprogramming systems and different cell types analyzed. “The exact stage of reprogramming of any cell can now be determined,” Dr. Pasque said. “This study signals a big change in thinking, because it provides simple and efficient tools for scientists to study stem cell creation in a stage-by-stage manner. Most studies to date ignore the stages of reprogramming, but we can now seek to better understand the entire process on both a macro and micro level.”
Dr. Plath’s group additionally discovered that the stages of reprogramming to iPSC are different from what was expected. They found that it is not simply the reversed sequence of stages of embryo development. Some steps are reversed in the expected order; others do not actually happen in the exact reverse order and resist a change until late during reprogramming to iPSCs. “This reflects how cells do not like to change from one specialized cell type to another and resist a change in cell identity,” Dr. Pasque said. “Resistance to reprogramming also helps to explain why reprogramming takes place only in a very small proportion of the starting cells.”
With these findings, Dr. Plath’s lab plans future studies to actively isolate specific cell types during specific stages of reprogramming. They also hope the research will encourage further investigation into the characteristics of iPSC development. “This research has broad impact, because by understanding cell reprogramming better we have the potential to improve disease modeling and the generation of better sources of patient-specific specialized cells suitable for replacement therapy,” concluded Dr. Plath. “This can ultimately benefit patients with new and better treatments for a wide range of diseases.”
Related Links:
University of California, Los Angeles’ Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research
The study, conducted by researchers from the University of California, Los Angeles (UCLA; USA) Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research and led by Dr. Kathrin Plath, a professor of biological chemistry, was published January 2015 in the journal Cell. Induced pluripotent stem cells (iPSCs) are cells that can be generated from adult cells and then, like embryonic stem cells, be directed to become any cell in the human body. Adult cells can also be reprogrammed in the lab to change from a specialized cell back to an iPSC (and thereby becoming a cell similar to that of an embryonic stem cell).
Reprogramming takes one to two weeks and is a mostly inefficient process, with typically less than one percent of the beginning cells effectively becoming an iPSC. The exact stages a cell goes through during the reprogramming process are not well understood. This knowledge is vital, because iPSCs have great potential in the field of regenerative medicine, as they can constantly reproduce and provide a single source of patient-specific cells to replace those lost to injury or disease. They can also be used to create innovative disease models from which new drugs and therapies can be developed.
Vincent Pasque and Jason Tchieu, postdoctoral fellows in Plath’s lab and co-first authors of the study, developed a roadmap of the reprogramming process using detailed time-course analyses. They induced the reprogramming of specialized cells (that could only make more of themselves, and no other cell types), then observed and analyzed on a daily basis or every other day the process of transformation at the single-cell level. The data were gathered and recorded during a time period of up to two weeks.
Dr. Plath’s team found that the changes that happen in cells during reprogramming occur in sequentially, and that notably, the stages of the sequence were the same across the diverse reprogramming systems and different cell types analyzed. “The exact stage of reprogramming of any cell can now be determined,” Dr. Pasque said. “This study signals a big change in thinking, because it provides simple and efficient tools for scientists to study stem cell creation in a stage-by-stage manner. Most studies to date ignore the stages of reprogramming, but we can now seek to better understand the entire process on both a macro and micro level.”
Dr. Plath’s group additionally discovered that the stages of reprogramming to iPSC are different from what was expected. They found that it is not simply the reversed sequence of stages of embryo development. Some steps are reversed in the expected order; others do not actually happen in the exact reverse order and resist a change until late during reprogramming to iPSCs. “This reflects how cells do not like to change from one specialized cell type to another and resist a change in cell identity,” Dr. Pasque said. “Resistance to reprogramming also helps to explain why reprogramming takes place only in a very small proportion of the starting cells.”
With these findings, Dr. Plath’s lab plans future studies to actively isolate specific cell types during specific stages of reprogramming. They also hope the research will encourage further investigation into the characteristics of iPSC development. “This research has broad impact, because by understanding cell reprogramming better we have the potential to improve disease modeling and the generation of better sources of patient-specific specialized cells suitable for replacement therapy,” concluded Dr. Plath. “This can ultimately benefit patients with new and better treatments for a wide range of diseases.”
Related Links:
University of California, Los Angeles’ Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research
Latest BioResearch News
- Lung Cancer Study Reveals Cellular Program Behind Therapy Resistance
- Tumor Genome Marker May Predict Treatment Benefit in Pediatric Cancers
- Lysosomal Gene Defect Linked to Severe Childhood Brain Disorders
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
Channels
Clinical Chemistry
view channel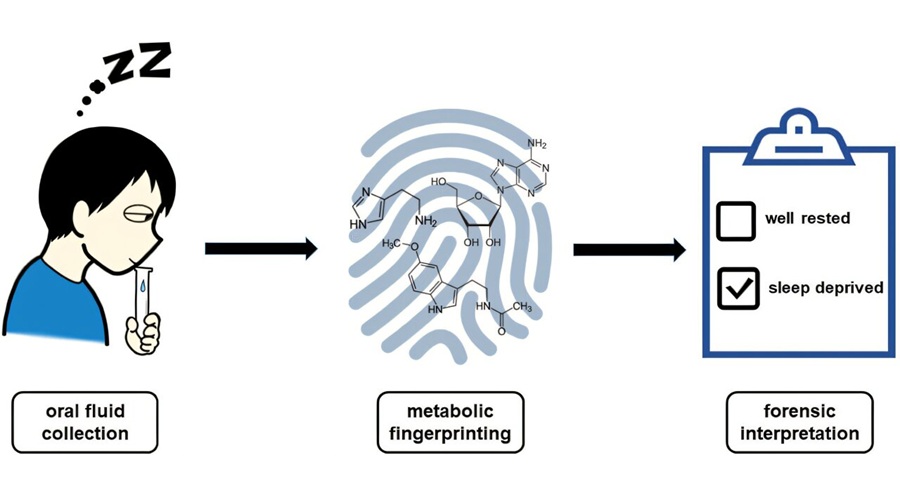
Saliva-Based Test Detects Biochemical Signs of Sleep Loss
Acute sleep loss impairs cognition and motor skills, raising safety risks that resemble alcohol intoxication. Clinicians currently lack an objective biochemical test to determine when someone is dangerously... Read more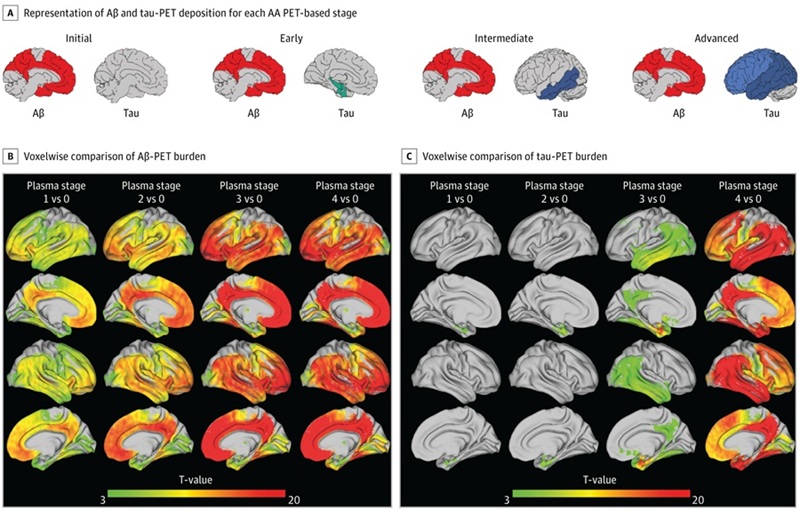
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreMolecular Diagnostics
view channel
New PCR Assay Supports Bundibugyo Ebola Outbreak Surveillance
Rapid identification of Ebola infections is essential to limit transmission and guide public health response, yet detection can be difficult when outbreaks involve rare variants. The current outbreaks... Read more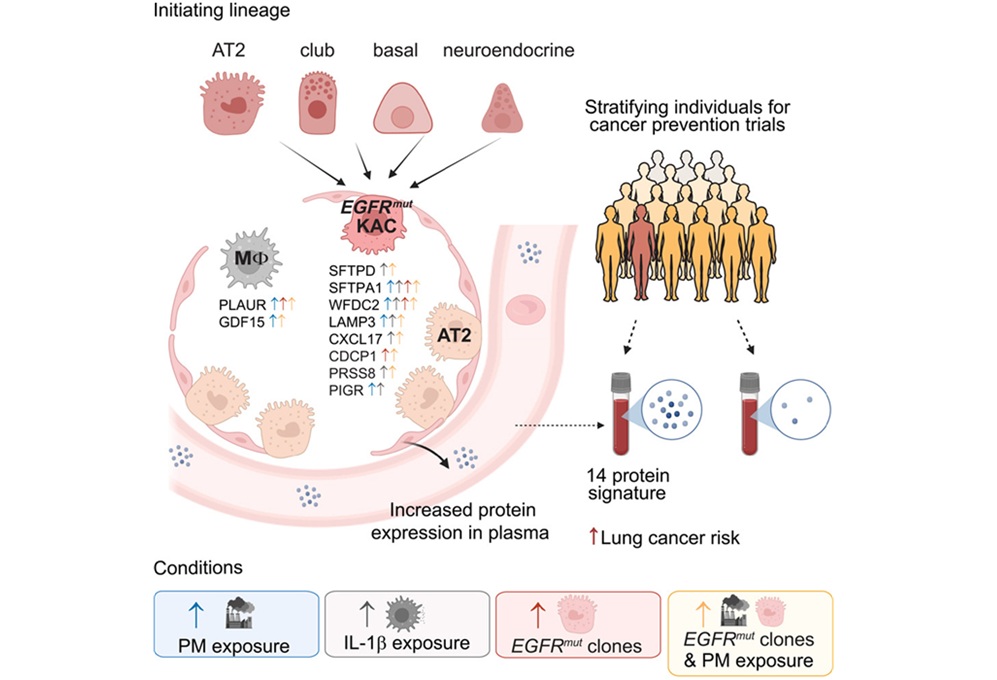
Plasma Protein Signature Predicts Lung Cancer Risk Up to Five Years Ahead
Lung cancer remains a leading cause of cancer death, and many cases are detected only after symptoms appear. Current screening programs largely target people with a history of smoking, leaving other at-risk... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more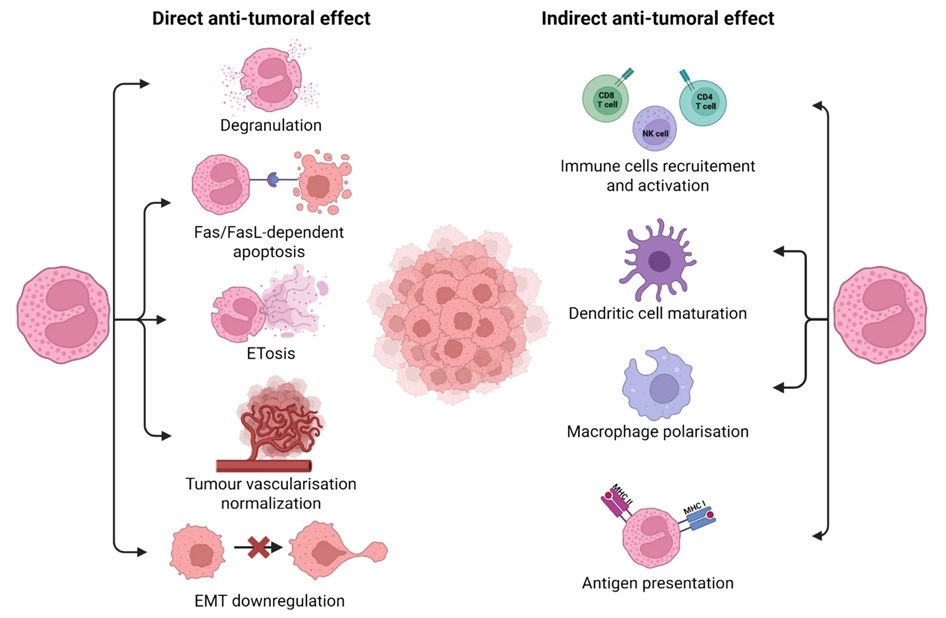
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channelAptamer-Based Biosensor Enables Mutation-Resilient SARS-CoV-2 Detection
Rapid evolution of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) can undermine existing molecular diagnostics, especially when assays target small viral components. Double-antibody sandwich... Read more
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more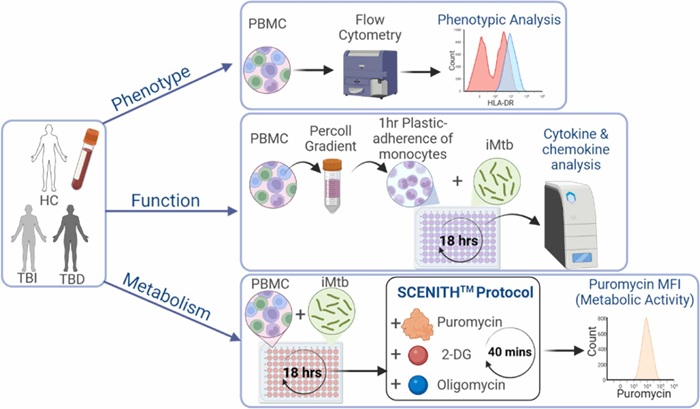
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read moreMicrobiology
view channel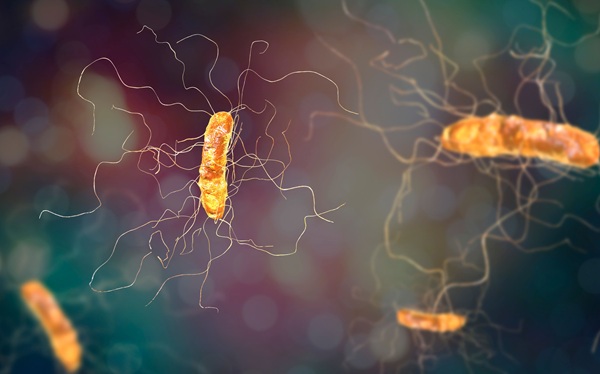
New Culture Medium Speeds C. difficile Resistance Detection and Reduces Costs
Clostridioides difficile infections remain a persistent threat in hospitals and communities, affecting about 500,000 people in the United States each year. Severe cases can be fatal within 30 days of diagnosis,... Read more
Automated Blood Culture System Speeds Detection of Bloodstream Infections
Bloodstream infections and sepsis require rapid laboratory detection to guide targeted antimicrobial therapy and reduce mortality. Conventional blood culture workflows can delay actionable results by critical... Read morePathology
view channel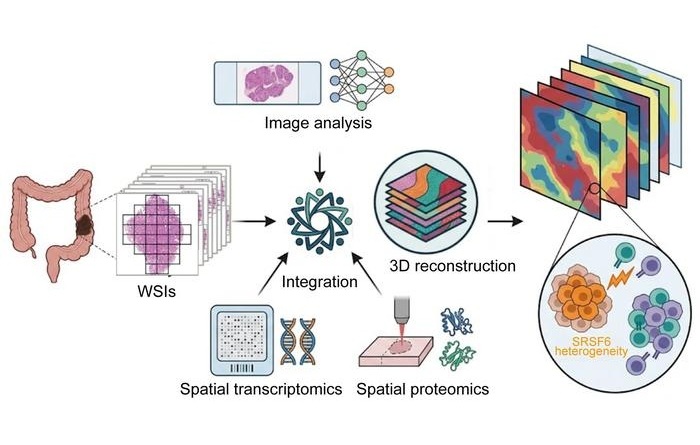
3D Spatial Multi-Omics Maps Intra-Tumor Diversity in Colorectal Cancer
Colorectal cancer remains a leading cause of cancer death, and clinical decision-making is complicated by marked intra-tumor heterogeneity. Conventional bulk sequencing averages molecular signals across... Read more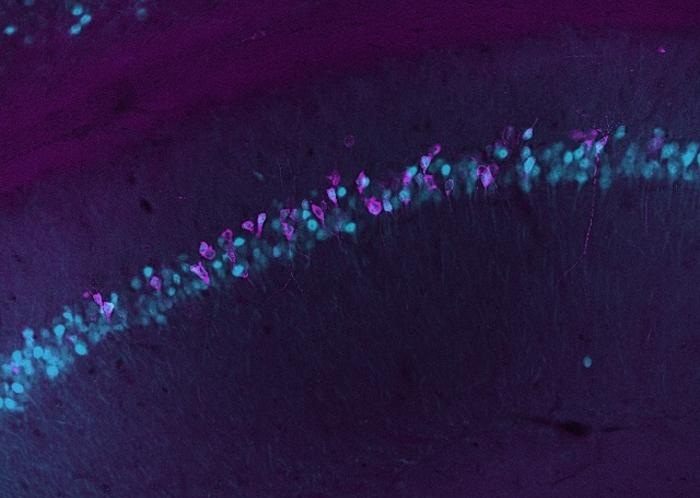
Blood-Based Method Tracks Gene Activity in the Living Brain
Real-time measurement of gene activity in the brain has been limited by assays requiring destructive tissue sampling. Tracking active genes could reveal how the body responds to environmental factors,... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel
Collaboration Advances ctDNA-Guided Development in Metastatic Colorectal Cancer
Natera, Inc. (Austin, TX, USA) and CytoDyn Inc. (Vancouver, WA, USA) announced a strategic collaboration focused on metastatic colorectal cancer (mCRC). Under the agreement, Natera will evaluate circulating... Read more