Epigenetic Markers Predict Type 2 Diabetes Patients Response to Metformin
|
By LabMedica International staff writers Posted on 29 Sep 2020 |
Image: Various types of data analysis using BeadStudio Methylation Module with Illumina`s MethylationEpic array (Photo courtesy of Phoebe Lu).
Generally, metformin is the first medication prescribed for type 2 diabetes (T2D). It works by lowering glucose production in the liver and improving the body's sensitivity to insulin so that the body uses insulin more effectively. Currently, there are no phenotypes that successfully predict glycemic response to, or tolerance of, metformin.
About 30% of patients do not respond to metformin and between 20% and 30% experience side effects that can be intolerable Gastrointestinal side effects of metformin are observed in 10% to 15% of patients, depending on the dose, and include abdominal discomfort, anorexia, bloating, and diarrhea. Because insulin secretion is unaltered, hypoglycemia is not a side effect of metformin used as monotherapy.
An international team of clinical scientists led by those at Skåne University Hospital (Malmo, Sweden) conducted multiple epigenome-wide association studies by analyzing in the blood of drug-naïve patients who were recently diagnosed with T2D. Blood samples were collected from the All New Diabetics In Scandia (ANDIS) cohort and analyzed using Illumina's MethylationEpic array (Illumina, San Diego, CA, USA). The team sought to gauge whether differences in DNA methylation prior to treatment could predict whether individuals had changes in glycated hemoglobin (HbA1c), responded to the drug treatment, or experienced intolerance to the drug following about a year and a half of metformin treatment.
The investigators identified more than 2,500 methylation sites that were significantly associated with changes in HbA1c, a marker of blood glucose levels. In the replication cohort, 132 CpGs of these sites were validated. They additionally uncovered 7,916 methylation sites that differed between individuals with T2D who responded to metformin and individuals who did not. Of those, 601 were then validated in the ANDIS replication cohort and 329 in an additional cohorts.
In all, 33 CpG sites were associated with future metformin response in all cohorts, and in a combined meta-analysis 11 sites reached genome-wide significance. At the same time, the team found 9,676 methylation sites that differed between individuals with T2D who could tolerate metformin treatment and those who could not. In the ANDIS replication cohort, 235 CpGs were validated, and in the replication cohort, 352 CpGs were. Seven CpGs were associated with metformin in all cohorts, and in a combined meta-analysis four sites reached genome-wide significance.
The scientists generated two methylation risk scores, one of metformin response and one of metformin intolerance. For the metformin response, they bundled together the 11 sites to form a weighted methylation risk score that could differentiate between responders and non-responders with an area under the curve of between 0.80 and 0.89. Meanwhile, for metformin intolerance, they combined the four sites that reached genome-wide significance into a separate risk score that could differentiate between tolerant and intolerant individuals with an area under the curve of between 0.85 and 0.94.
The authors concludes that they could discriminate between glycemic responders/non-responders and participants tolerant/intolerant to metformin at diagnosis by measuring blood-based epigenetic markers in drug-naïve patients with T2D. This epigenetics-based tool may be further developed to help patients with T2D receive optimal therapy. The study was published on September 16, 2020 in the journal Science Translational Medicine.
About 30% of patients do not respond to metformin and between 20% and 30% experience side effects that can be intolerable Gastrointestinal side effects of metformin are observed in 10% to 15% of patients, depending on the dose, and include abdominal discomfort, anorexia, bloating, and diarrhea. Because insulin secretion is unaltered, hypoglycemia is not a side effect of metformin used as monotherapy.
An international team of clinical scientists led by those at Skåne University Hospital (Malmo, Sweden) conducted multiple epigenome-wide association studies by analyzing in the blood of drug-naïve patients who were recently diagnosed with T2D. Blood samples were collected from the All New Diabetics In Scandia (ANDIS) cohort and analyzed using Illumina's MethylationEpic array (Illumina, San Diego, CA, USA). The team sought to gauge whether differences in DNA methylation prior to treatment could predict whether individuals had changes in glycated hemoglobin (HbA1c), responded to the drug treatment, or experienced intolerance to the drug following about a year and a half of metformin treatment.
The investigators identified more than 2,500 methylation sites that were significantly associated with changes in HbA1c, a marker of blood glucose levels. In the replication cohort, 132 CpGs of these sites were validated. They additionally uncovered 7,916 methylation sites that differed between individuals with T2D who responded to metformin and individuals who did not. Of those, 601 were then validated in the ANDIS replication cohort and 329 in an additional cohorts.
In all, 33 CpG sites were associated with future metformin response in all cohorts, and in a combined meta-analysis 11 sites reached genome-wide significance. At the same time, the team found 9,676 methylation sites that differed between individuals with T2D who could tolerate metformin treatment and those who could not. In the ANDIS replication cohort, 235 CpGs were validated, and in the replication cohort, 352 CpGs were. Seven CpGs were associated with metformin in all cohorts, and in a combined meta-analysis four sites reached genome-wide significance.
The scientists generated two methylation risk scores, one of metformin response and one of metformin intolerance. For the metformin response, they bundled together the 11 sites to form a weighted methylation risk score that could differentiate between responders and non-responders with an area under the curve of between 0.80 and 0.89. Meanwhile, for metformin intolerance, they combined the four sites that reached genome-wide significance into a separate risk score that could differentiate between tolerant and intolerant individuals with an area under the curve of between 0.85 and 0.94.
The authors concludes that they could discriminate between glycemic responders/non-responders and participants tolerant/intolerant to metformin at diagnosis by measuring blood-based epigenetic markers in drug-naïve patients with T2D. This epigenetics-based tool may be further developed to help patients with T2D receive optimal therapy. The study was published on September 16, 2020 in the journal Science Translational Medicine.
Latest Pathology News
- Engineered Yeast Cells Enable Rapid Testing of Cancer Immunotherapy
- First-Of-Its-Kind Test Identifies Autism Risk at Birth
- AI Algorithms Improve Genetic Mutation Detection in Cancer Diagnostics
- Skin Biopsy Offers New Diagnostic Method for Neurodegenerative Diseases
- Fast Label-Free Method Identifies Aggressive Cancer Cells
- New X-Ray Method Promises Advances in Histology
- Single-Cell Profiling Technique Could Guide Early Cancer Detection
- Intraoperative Tumor Histology to Improve Cancer Surgeries
- Rapid Stool Test Could Help Pinpoint IBD Diagnosis
- AI-Powered Label-Free Optical Imaging Accurately Identifies Thyroid Cancer During Surgery
- Deep Learning–Based Method Improves Cancer Diagnosis
- ADLM Updates Expert Guidance on Urine Drug Testing for Patients in Emergency Departments
- New Age-Based Blood Test Thresholds to Catch Ovarian Cancer Earlier
- Genetics and AI Improve Diagnosis of Aortic Stenosis
- AI Tool Simultaneously Identifies Genetic Mutations and Disease Type
- Rapid Low-Cost Tests Can Prevent Child Deaths from Contaminated Medicinal Syrups
Channels
Clinical Chemistry
view channel
New PSA-Based Prognostic Model Improves Prostate Cancer Risk Assessment
Prostate cancer is the second-leading cause of cancer death among American men, and about one in eight will be diagnosed in their lifetime. Screening relies on blood levels of prostate-specific antigen... Read more
Extracellular Vesicles Linked to Heart Failure Risk in CKD Patients
Chronic kidney disease (CKD) affects more than 1 in 7 Americans and is strongly associated with cardiovascular complications, which account for more than half of deaths among people with CKD.... Read moreHematology
view channel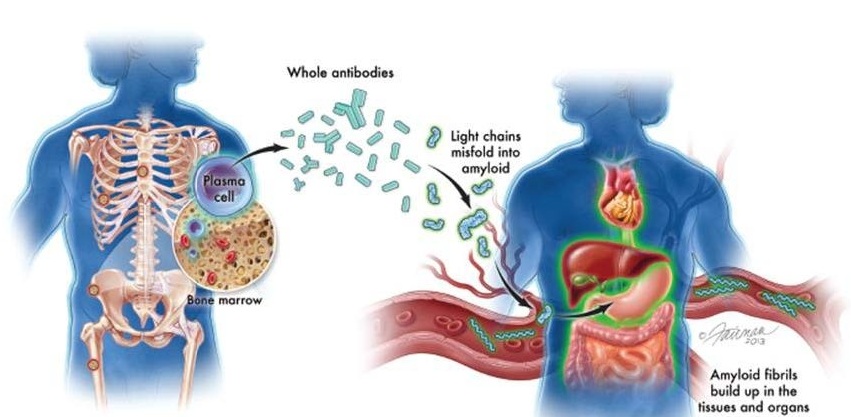
New Guidelines Aim to Improve AL Amyloidosis Diagnosis
Light chain (AL) amyloidosis is a rare, life-threatening bone marrow disorder in which abnormal amyloid proteins accumulate in organs. Approximately 3,260 people in the United States are diagnosed... Read more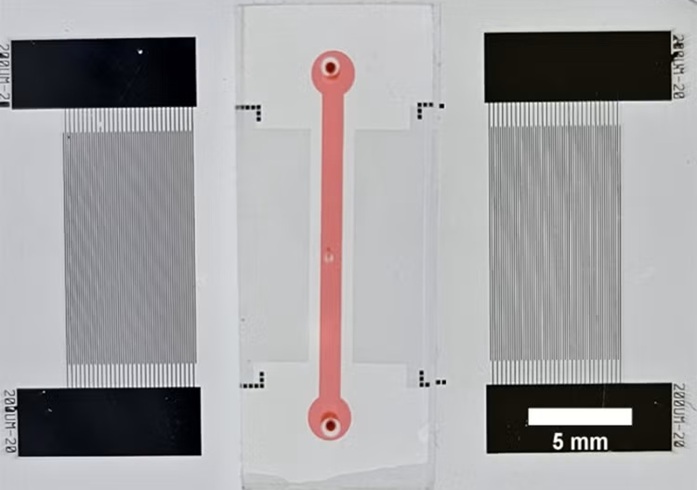
Fast and Easy Test Could Revolutionize Blood Transfusions
Blood transfusions are a cornerstone of modern medicine, yet red blood cells can deteriorate quietly while sitting in cold storage for weeks. Although blood units have a fixed expiration date, cells from... Read more
Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
High-volume hemostasis sections must sustain rapid turnaround while managing reruns and reflex testing. Manual tube handling and preanalytical checks can strain staff time and increase opportunities for error.... Read more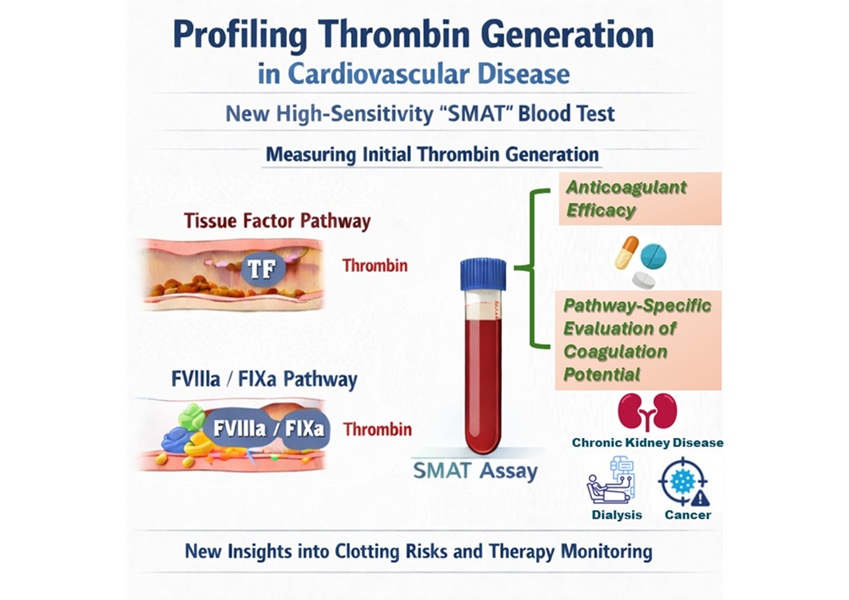
High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
Blood clotting is essential for preventing bleeding, but even small imbalances can lead to serious conditions such as thrombosis or dangerous hemorrhage. In cardiovascular disease, clinicians often struggle... Read moreImmunology
view channelBlood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
Small cell lung cancer (SCLC) is an aggressive disease with limited treatment options, and even newly approved immunotherapies do not benefit all patients. While immunotherapy can extend survival for some,... Read more
Whole-Genome Sequencing Approach Identifies Cancer Patients Benefitting From PARP-Inhibitor Treatment
Targeted cancer therapies such as PARP inhibitors can be highly effective, but only for patients whose tumors carry specific DNA repair defects. Identifying these patients accurately remains challenging,... Read more
Ultrasensitive Liquid Biopsy Demonstrates Efficacy in Predicting Immunotherapy Response
Immunotherapy has transformed cancer treatment, but only a small proportion of patients experience lasting benefit, with response rates often remaining between 10% and 20%. Clinicians currently lack reliable... Read moreMicrobiology
view channel
Comprehensive Review Identifies Gut Microbiome Signatures Associated With Alzheimer’s Disease
Alzheimer’s disease affects approximately 6.7 million people in the United States and nearly 50 million worldwide, yet early cognitive decline remains difficult to characterize. Increasing evidence suggests... Read moreAI-Powered Platform Enables Rapid Detection of Drug-Resistant C. Auris Pathogens
Infections caused by the pathogenic yeast Candida auris pose a significant threat to hospitalized patients, particularly those with weakened immune systems or those who have invasive medical devices.... Read morePathology
view channel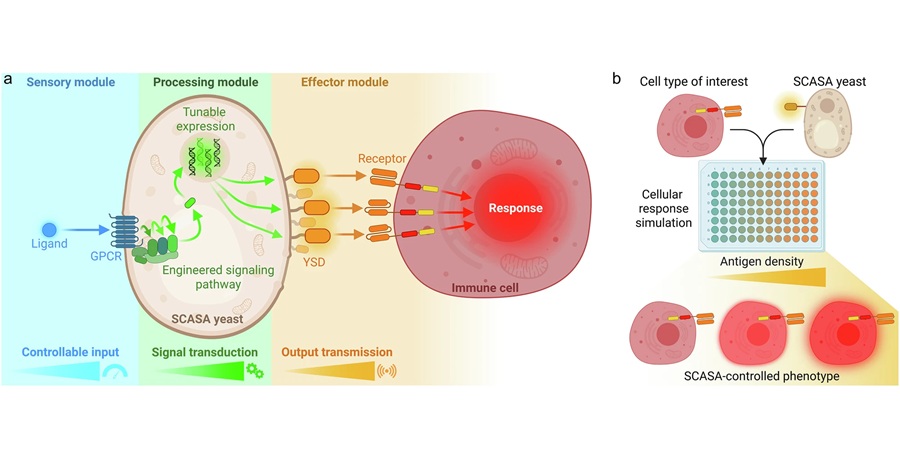
Engineered Yeast Cells Enable Rapid Testing of Cancer Immunotherapy
Developing new cancer immunotherapies is a slow, costly, and high-risk process, particularly for CAR T cell treatments that must precisely recognize cancer-specific antigens. Small differences in tumor... Read more
First-Of-Its-Kind Test Identifies Autism Risk at Birth
Autism spectrum disorder is treatable, and extensive research shows that early intervention can significantly improve cognitive, social, and behavioral outcomes. Yet in the United States, the average age... Read moreTechnology
view channel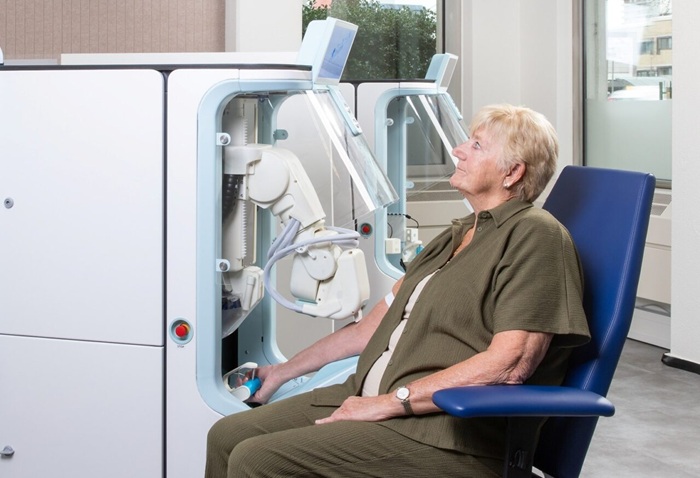
Robotic Technology Unveiled for Automated Diagnostic Blood Draws
Routine diagnostic blood collection is a high‑volume task that can strain staffing and introduce human‑dependent variability, with downstream implications for sample quality and patient experience.... Read more
ADLM Launches First-of-Its-Kind Data Science Program for Laboratory Medicine Professionals
Clinical laboratories generate billions of test results each year, creating a treasure trove of data with the potential to support more personalized testing, improve operational efficiency, and enhance patient care.... Read moreAptamer Biosensor Technology to Transform Virus Detection
Rapid and reliable virus detection is essential for controlling outbreaks, from seasonal influenza to global pandemics such as COVID-19. Conventional diagnostic methods, including cell culture, antigen... Read more
AI Models Could Predict Pre-Eclampsia and Anemia Earlier Using Routine Blood Tests
Pre-eclampsia and anemia are major contributors to maternal and child mortality worldwide, together accounting for more than half a million deaths each year and leaving millions with long-term health complications.... Read moreIndustry
view channelNew Collaboration Brings Automated Mass Spectrometry to Routine Laboratory Testing
Mass spectrometry is a powerful analytical technique that identifies and quantifies molecules based on their mass and electrical charge. Its high selectivity, sensitivity, and accuracy make it indispensable... Read more
AI-Powered Cervical Cancer Test Set for Major Rollout in Latin America
Noul Co., a Korean company specializing in AI-based blood and cancer diagnostics, announced it will supply its intelligence (AI)-based miLab CER cervical cancer diagnostic solution to Mexico under a multi‑year... Read more
Diasorin and Fisher Scientific Enter into US Distribution Agreement for Molecular POC Platform
Diasorin (Saluggia, Italy) has entered into an exclusive distribution agreement with Fisher Scientific, part of Thermo Fisher Scientific (Waltham, MA, USA), for the LIAISON NES molecular point-of-care... Read more