Novel Class of Enhancer RNAs Linked to Growth of Cancers
|
By LabMedica International staff writers Posted on 27 Aug 2018 |
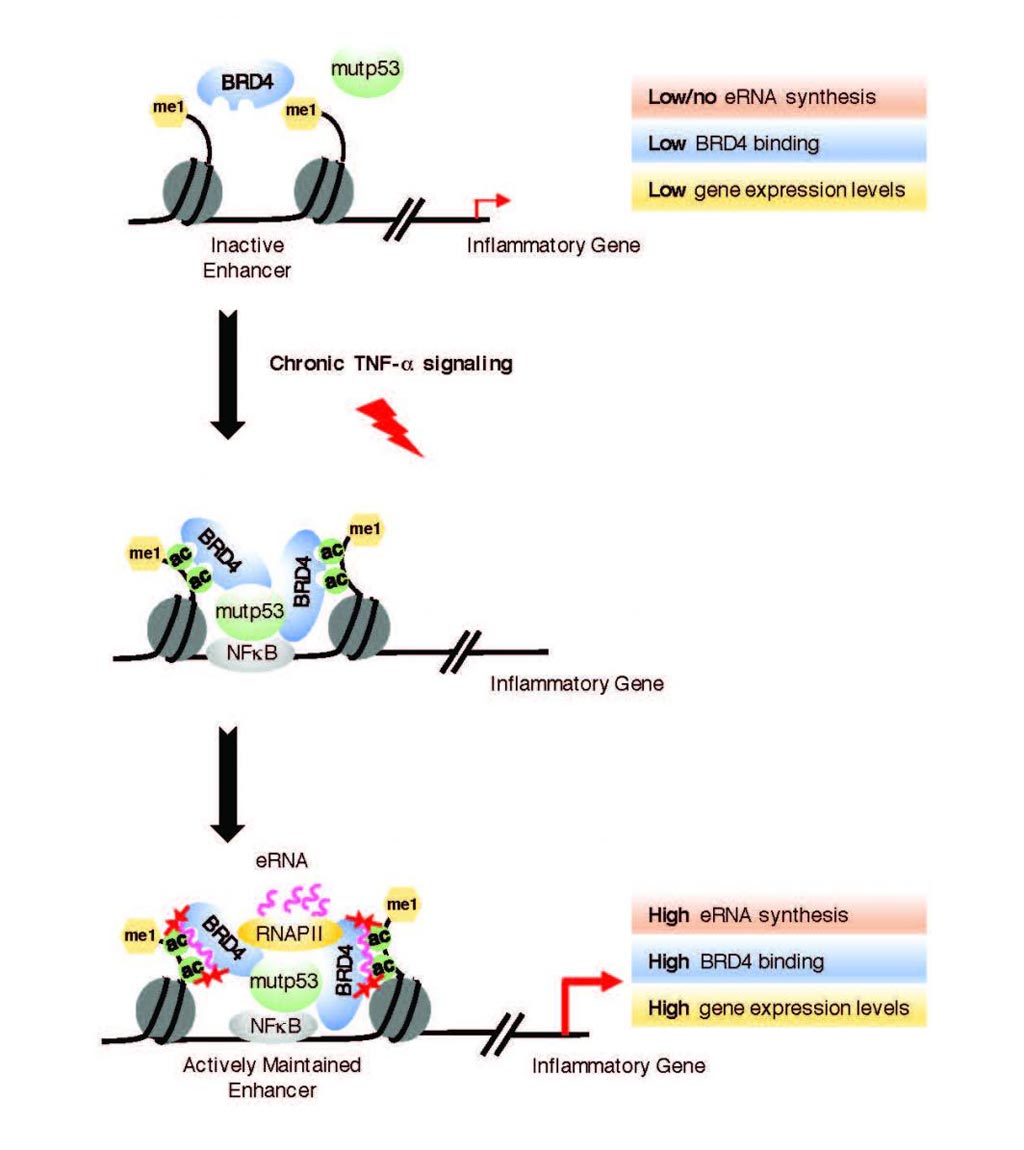
Image: Active enhancer RNAs (eRNAs) interact directly with BRD4, a protein linked to tumor development (Photo courtesy of the Lauberth laboratory, University of California, San Diego).
A recently described class of microRNA has been linked to the cancer-promoting activity of the mutated form of p53 protein.
MicroRNAs (miRNAs) and short interfering RNAs (siRNA) comprise a class of about 20 nucleotides-long RNA fragments that block gene expression by attaching to molecules of messenger RNA in a fashion that prevents them from transmitting the protein synthesizing instructions they had received from the DNA. MiRNAs resemble siRNAs of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. With their capacity to fine-tune protein expression via sequence-specific interactions, miRNAs help regulate cell maintenance and differentiation.
Augmenting the repertoire of "classical" miRNAs, investigators at the University of California, San Diego (USA) identified several thousand enhancer RNAs (eRNAs) that were robustly produced in colon cancer cells in response to chronic immune signaling. Enhancer RNAs are transcribed from DNA sequences upstream and downstream of extragenic enhancer regions. Depending on the directionality of transcription, enhancer regions generate two different types of non-coding transcripts, unidirectional-eRNAs and bidirectional-eRNAs. The nature of the pre-initiation complex and specific transcription factors recruited to the enhancer may control the type of eRNAs generated.
After transcription, the majority of eRNAs remain in the nucleus, and as they are very unstable, they are actively degraded by the nuclear exosome. Not all enhancers are transcribed, with non-transcribed enhancers greatly outnumbering the transcribed ones in the order of magnitude of dozens of thousands in every given cell type. The theory that not all enhancers are transcribed at the same time and that eRNA transcription correlates with enhancer-specific activity support the idea that individual eRNAs carry distinct and relevant biological functions.
The investigators reported in the August 3, 2018, issue of the journal Nature Structural and Molecular Biology that in human colorectal cancer cells, Bromodomain-containing protein 4 (BRD4) was recruited to enhancers that were co-occupied by mutant p53 and supported the synthesis of enhancer-directed transcripts (eRNAs) in response to chronic immune signaling. BRD4 selectively associated with eRNAs that were produced from BRD4-bound enhancers.
BRD4 is a member of the BET (bromodomain and extra terminal domain) protein family, which also includes BRD2, BRD3, and BRDT. BRD4, similar to other BET family members, contains two bromodomains (BDs) that recognize acetylated lysine residues. BRD4 also has an extended C-terminal domain with little sequence homology to other BET family members.
The investigators used biochemical and biophysical methods to show that BRD4 BDs functioned cooperatively as docking sites for eRNAs and that the BDs of BRD2, BRD3, BRDT, BRG1, and BRD7 directly interacted with eRNAs. BRD4-eRNA interactions increased BRD4 binding to acetylated histones in vitro and augmented BRD4 enhancer recruitment and transcriptional cofactor activities.
"Our findings reveal that eRNAs are key regulators of cancer by acting to reinforce BRD4 binding and keep it anchored on DNA, which keeps the tumor-promoting genes turned on at high levels," said senior author Dr. Shannon Lauberth, assistant professor of biology at the University of California, San Diego. "Interestingly, when we deplete several of these eRNAs, we can significantly reduce the expression of the tumor-promoting genes that the eRNAs and BRD4 are co-regulating. Now that we see that eRNAs impact BRD4 function, we have to rethink the way that we therapeutically target BRD4. Taken together, our findings are consistent with the emerging notion that eRNAs are functional molecules, rather than merely reflections of enhancer activation or simply transcriptional noise."
Related Links:
University of California, San Diego
MicroRNAs (miRNAs) and short interfering RNAs (siRNA) comprise a class of about 20 nucleotides-long RNA fragments that block gene expression by attaching to molecules of messenger RNA in a fashion that prevents them from transmitting the protein synthesizing instructions they had received from the DNA. MiRNAs resemble siRNAs of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of double-stranded RNA. With their capacity to fine-tune protein expression via sequence-specific interactions, miRNAs help regulate cell maintenance and differentiation.
Augmenting the repertoire of "classical" miRNAs, investigators at the University of California, San Diego (USA) identified several thousand enhancer RNAs (eRNAs) that were robustly produced in colon cancer cells in response to chronic immune signaling. Enhancer RNAs are transcribed from DNA sequences upstream and downstream of extragenic enhancer regions. Depending on the directionality of transcription, enhancer regions generate two different types of non-coding transcripts, unidirectional-eRNAs and bidirectional-eRNAs. The nature of the pre-initiation complex and specific transcription factors recruited to the enhancer may control the type of eRNAs generated.
After transcription, the majority of eRNAs remain in the nucleus, and as they are very unstable, they are actively degraded by the nuclear exosome. Not all enhancers are transcribed, with non-transcribed enhancers greatly outnumbering the transcribed ones in the order of magnitude of dozens of thousands in every given cell type. The theory that not all enhancers are transcribed at the same time and that eRNA transcription correlates with enhancer-specific activity support the idea that individual eRNAs carry distinct and relevant biological functions.
The investigators reported in the August 3, 2018, issue of the journal Nature Structural and Molecular Biology that in human colorectal cancer cells, Bromodomain-containing protein 4 (BRD4) was recruited to enhancers that were co-occupied by mutant p53 and supported the synthesis of enhancer-directed transcripts (eRNAs) in response to chronic immune signaling. BRD4 selectively associated with eRNAs that were produced from BRD4-bound enhancers.
BRD4 is a member of the BET (bromodomain and extra terminal domain) protein family, which also includes BRD2, BRD3, and BRDT. BRD4, similar to other BET family members, contains two bromodomains (BDs) that recognize acetylated lysine residues. BRD4 also has an extended C-terminal domain with little sequence homology to other BET family members.
The investigators used biochemical and biophysical methods to show that BRD4 BDs functioned cooperatively as docking sites for eRNAs and that the BDs of BRD2, BRD3, BRDT, BRG1, and BRD7 directly interacted with eRNAs. BRD4-eRNA interactions increased BRD4 binding to acetylated histones in vitro and augmented BRD4 enhancer recruitment and transcriptional cofactor activities.
"Our findings reveal that eRNAs are key regulators of cancer by acting to reinforce BRD4 binding and keep it anchored on DNA, which keeps the tumor-promoting genes turned on at high levels," said senior author Dr. Shannon Lauberth, assistant professor of biology at the University of California, San Diego. "Interestingly, when we deplete several of these eRNAs, we can significantly reduce the expression of the tumor-promoting genes that the eRNAs and BRD4 are co-regulating. Now that we see that eRNAs impact BRD4 function, we have to rethink the way that we therapeutically target BRD4. Taken together, our findings are consistent with the emerging notion that eRNAs are functional molecules, rather than merely reflections of enhancer activation or simply transcriptional noise."
Related Links:
University of California, San Diego
Latest BioResearch News
- Genetic Testing Identifies Greater Inherited Sudden Cardiac Arrest Risk in Younger Individuals
- Hidden 'Jumping Gene' Variant Linked to Higher Pancreatic Cancer Risk
- Common White Blood Cells Produce Schizophrenia-Linked Protein
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
- Study Links Abnormal Gene Splicing to Treatment Response in Metastatic Kidney Cancer
- Research Reveals How Some Aplastic Anemia Patients Recover Bone Marrow Function
- New Molecular Insights Support Diagnosis of Hodgkin Lymphoma
Channels
Clinical Chemistry
view channel
New CA19-9 Cutoff Value Helps Identify High-Risk Pancreatic Cancer Patients
Pancreatic ductal adenocarcinoma (PDAC) is frequently diagnosed at an advanced stage and remains one of the most lethal solid tumors. Clinicians commonly use serum carbohydrate antigen 19-9 (CA19-9) to... Read more
Blood-Based Biomarkers Show Promise for Psychosis Risk Prediction
Psychosis commonly emerges in adolescence or early adulthood and can severely disrupt social and occupational functioning. Hallucinations, delusions, and disorganized thinking often evolve gradually, hindering... Read moreMolecular Diagnostics
view channel
New RNA Origami Method Supports Faster Targeted Testing for Repeat Expansion Disorders
Repeat expansion disorders drive conditions such as myotonic dystrophy, Huntington’s disease, and amyotrophic lateral sclerosis (ALS), yet accurately sizing the mutated sequences remains difficult.... Read more
FDA Approves Expanded Liquid Biopsy Panel for Advanced Cancer Profiling
Timely, comprehensive tumor profiling helps clinicians make treatment selection decisions for patients with advanced cancer. Blood-based approaches can provide actionable insights from a simple draw and... Read moreHematology
view channel
Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
Iron deficiency in school-age children can affect brain development, learning, growth, and physical performance, yet early deficiency may be missed when screening focuses mainly on anemia.... Read more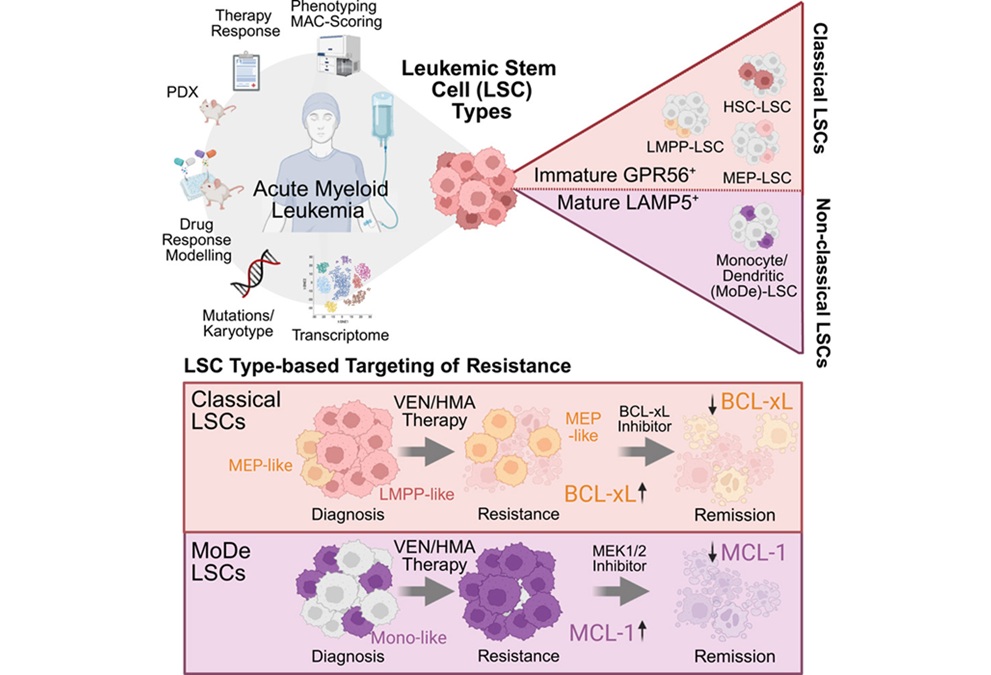
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read moreImmunology
view channel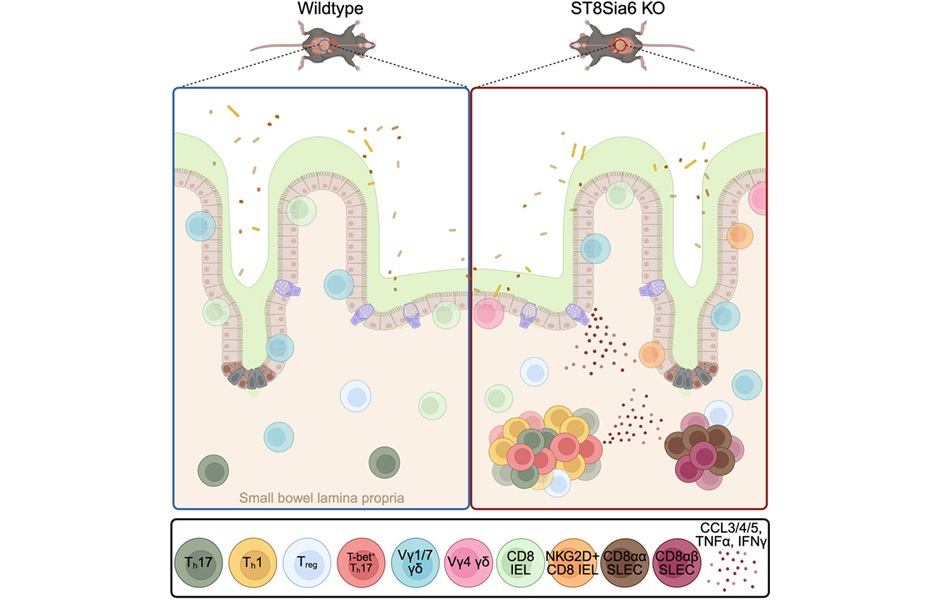
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read more
Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
Lung transplant recipients face some of the highest rates of acute cellular rejection, and routine surveillance often relies on repeated surgical biopsies. These procedures can cause complications such... Read moreMicrobiology
view channel
New AMR Assay Supports Rapid Infection Control Screening in Hospitals
As antimicrobial resistance spreads worldwide, healthcare-associated infections are placing a growing burden on hospitals, increasing the need for faster and broader diagnostic solutions.... Read more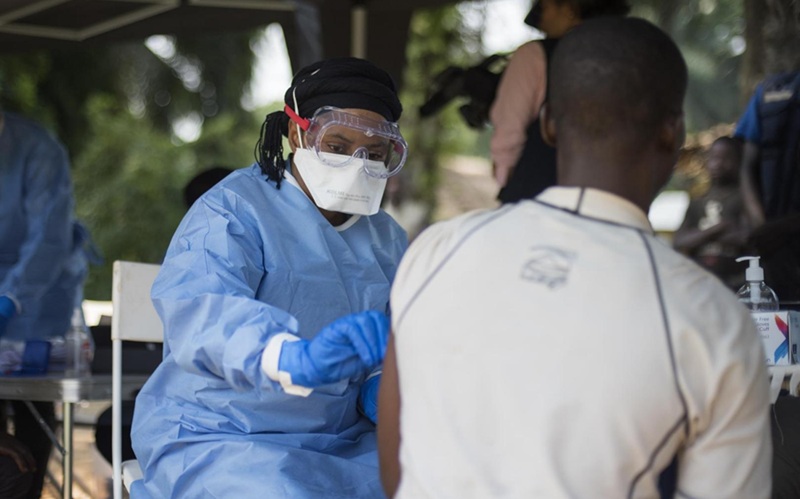
Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
In eastern Democratic Republic of the Congo, communities are confronting a resurgence of Bundibugyo ebolavirus, a rarer species for which no vaccines or treatments have been approved. Ebola is a highly... Read more
Study Finds Hidden Mpox Infections May Drive Ongoing Spread
Mpox continues to circulate despite vaccination, and many cases show no known link to a symptomatic partner. The role of people without symptoms has remained uncertain, limiting clarity on how transmission persists.... Read more
Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
Antimicrobial resistance (AMR) poses a growing threat to patient safety, with carbapenem-resistant Enterobacterales causing difficult-to-treat infections and leaving clinicians with limited therapeutic options.... Read morePathology
view channel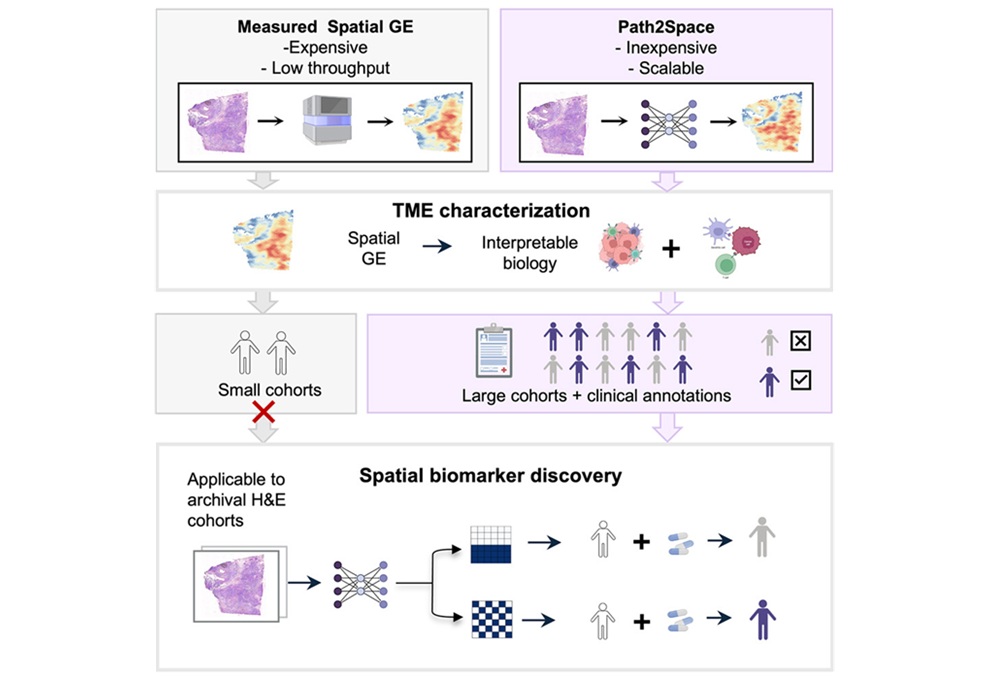
Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
Gene expression profiling can inform tumor biology and treatment selection, but spatial assays remain costly and time-consuming. Results can take weeks and cost thousands of dollars, limiting large-scale... Read more
AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
Non–muscle invasive bladder cancer has highly variable outcomes, complicating surveillance and treatment planning. Risk assessment typically relies on stage, grade, and tumor size, leaving uncertainty... Read moreTechnology
view channel
AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
Clinical laboratories and research groups face increasingly complex molecular workflows and expanding technical documentation spread across multiple systems. Fragmented digital tools can slow experiment... Read more
AI Tool Automates Validation of Laboratory Software Configuration Changes
Regulated laboratories face heavy documentation and requalification demands when software configurations change, slowing improvements and discouraging beneficial updates. A new capability now automates... Read moreIndustry
view channel
Strategic Collaboration Advances RNA Foundation Models for Precision Oncology
Bulk RNA sequencing is increasingly used to study tumor biology, but standard analyses often reduce results to gene-level summaries that miss important transcript variants and mutation patterns.... Read more