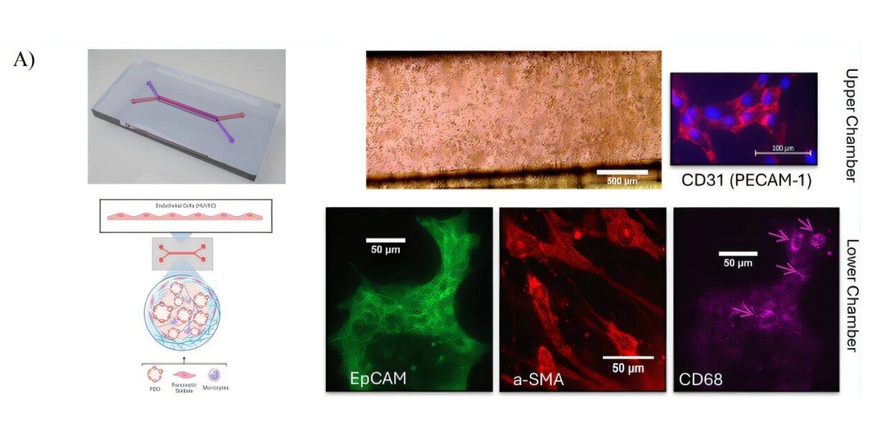
Computer-Designed Proteins Programmed to Deactivate Flu Viruses
|
By LabMedica International staff writers Posted on 20 Jun 2012 |
Computer-designed proteins are now being constructed to fight the flu. Researchers are demonstrating that proteins found in nature, but that do not normally bind the flu, can be engineered to act as broad-spectrum antiviral agents against a range of flu virus strains, including H1N1 pandemic influenza.
“One of these engineered proteins has a flu-fighting potency that rivals that of several human monoclonal antibodies,” said Dr. David Baker, a professor of biochemistry at the University of Washington (Seattle, USA), in a report June 7, 2012, published in the journal Nature Biotechnology.
Dr. Baker’s research team is making major inroads in optimizing the function of computer-designed influenza inhibitors. These proteins are constructed via computer modeling to fit exquisitely into a specific nano-sized target on flu viruses. By binding the target areas similar to a key into a lock, they keep the virus from changing shape, a tactic that the virus uses to infect living cells. The research efforts, analogous to docking a space station but on a molecular level, are made possible by computers that can describe the panoramas of forces involved on the submicroscopic scale.
Dr. Baker is head of the new Institute for Protein Design Center at the University of Washington. Biochemists, engineers, computer scientists, and medical specialists at the center are engineering innovative proteins with new functions for specific purposes in medicine, environmental protection, and other areas. Proteins underlie all typical activities and structures of living cells, and also control disease actions of pathogens such as viruses. Abnormal protein formation and interactions are also implicated in many inherited and later-life chronic disorders.
Because influenza is a serious worldwide public health problem due to its genetic shifts and drifts that sporadically become more virulent, the flu is one of the key interests of the Institutes for Protein Design and its collaborators in the United States and worldwide. Researchers are trying to meet the vital need for better therapeutic agents to protect against this very adaptable and extremely infective virus. Vaccines for new strains of influenza take months to develop, evaluate, and manufacture, and are not helpful for those already sick. The long response time for vaccine creation and distribution is unsettling when a more lethal strain abruptly emerges and spreads rapidly. The speed of transmission is accelerated by the lack of widespread immunity in the general population to the latest form of the virus.
Flu trackers refer to strains by their H and N subtypes. H stands for hemagglutinins, which are the molecules on the flu virus that enable it to invade the cells of respiratory passages. The virus’s hemagglutinin molecules attach to the surface of cells lining the respiratory tract. When the cell tries to engulf the virus, it makes the error of pulling it into a more acidic location. The drop in pH changes the shape of the viral hemagglutinin, thereby allowing the virus to fuse to the cell and open an entry for the virus’ RNA to come in and start making fresh viruses. It is hypothesized that the Baker Lab protein inhibits this shape change by binding the hemagglutinin in a very specific orientation and thus keeps the virus from invading cells.
Dr. Baker and his team wanted to create antivirals that could react against a wide variety of H subtypes, as this versatility could lead to a comprehensive therapy for influenza. Specifically, viruses that have hemagglutinins of the H2 subtype are responsible for the deadly pandemic of 1957 and continued to circulate until 1968. People born after that date have not been exposed to H2 viruses. The recent avian flu has a new version of H1 hemagglutinin. Data suggest that Dr. Baker’s proteins bind to all types of the group I hemagglutinin, a group that includes not only H1 but also the pandemic H2 and avian H5 strains.
The methods developed for the influenza inhibitor protein design, according to Dr. Baker, could be “a powerful route to inhibitors or binders for any surface patch on any desired target of interest.” For example, if a new disease pathogen arises, scientists could figure out how it interacts with human cells or other hosts on a molecular level. Scientists could then employ protein interface design to create a diversity of small proteins that they predict would block the pathogen’s interaction surface.
Genes for large numbers of the most promising, computer-designed proteins could be tested using yeast cells. After additional molecular chemistry research to search for the best binding among those proteins, those could be reprogrammed in the laboratory to undergo mutations, and all the mutated forms could be stored in a “library” for an in-depth examination of their amino acids, molecular architecture, and energy bonds.
Sophisticated technologies would allow the scientists to rapidly browse through the library to find those tiny proteins that clung to the pathogen surface target with pinpoint accuracy. The finalists would be selected from this pool for excelling at blocking the pathogen from attaching to, entering, and infecting human or animal cells.
The utilization of deep sequencing, the same technology now used to sequence human genomes cheaply, was particularly central in creating detailed maps relating sequencing to function. These maps were used to reprogram the design to achieve a more exact interaction between the inhibitor protein and the virus molecule. It also enabled the scientists, they said, “to leapfrog over bottlenecks” to improve the activity of the binder. They were able to see how small contributions from many small alterations in the protein, too difficult to see individually, could together create a binder with better attachment strength.
“We anticipate that our approach combining computational design followed by comprehensive energy landscape mapping,” Dr. Baker said, “will be widely useful in generating high-affinity and high-specificity binders to a broad range of targets for use in therapeutics and diagnostics.”
Related Links:
University of Washington
“One of these engineered proteins has a flu-fighting potency that rivals that of several human monoclonal antibodies,” said Dr. David Baker, a professor of biochemistry at the University of Washington (Seattle, USA), in a report June 7, 2012, published in the journal Nature Biotechnology.
Dr. Baker’s research team is making major inroads in optimizing the function of computer-designed influenza inhibitors. These proteins are constructed via computer modeling to fit exquisitely into a specific nano-sized target on flu viruses. By binding the target areas similar to a key into a lock, they keep the virus from changing shape, a tactic that the virus uses to infect living cells. The research efforts, analogous to docking a space station but on a molecular level, are made possible by computers that can describe the panoramas of forces involved on the submicroscopic scale.
Dr. Baker is head of the new Institute for Protein Design Center at the University of Washington. Biochemists, engineers, computer scientists, and medical specialists at the center are engineering innovative proteins with new functions for specific purposes in medicine, environmental protection, and other areas. Proteins underlie all typical activities and structures of living cells, and also control disease actions of pathogens such as viruses. Abnormal protein formation and interactions are also implicated in many inherited and later-life chronic disorders.
Because influenza is a serious worldwide public health problem due to its genetic shifts and drifts that sporadically become more virulent, the flu is one of the key interests of the Institutes for Protein Design and its collaborators in the United States and worldwide. Researchers are trying to meet the vital need for better therapeutic agents to protect against this very adaptable and extremely infective virus. Vaccines for new strains of influenza take months to develop, evaluate, and manufacture, and are not helpful for those already sick. The long response time for vaccine creation and distribution is unsettling when a more lethal strain abruptly emerges and spreads rapidly. The speed of transmission is accelerated by the lack of widespread immunity in the general population to the latest form of the virus.
Flu trackers refer to strains by their H and N subtypes. H stands for hemagglutinins, which are the molecules on the flu virus that enable it to invade the cells of respiratory passages. The virus’s hemagglutinin molecules attach to the surface of cells lining the respiratory tract. When the cell tries to engulf the virus, it makes the error of pulling it into a more acidic location. The drop in pH changes the shape of the viral hemagglutinin, thereby allowing the virus to fuse to the cell and open an entry for the virus’ RNA to come in and start making fresh viruses. It is hypothesized that the Baker Lab protein inhibits this shape change by binding the hemagglutinin in a very specific orientation and thus keeps the virus from invading cells.
Dr. Baker and his team wanted to create antivirals that could react against a wide variety of H subtypes, as this versatility could lead to a comprehensive therapy for influenza. Specifically, viruses that have hemagglutinins of the H2 subtype are responsible for the deadly pandemic of 1957 and continued to circulate until 1968. People born after that date have not been exposed to H2 viruses. The recent avian flu has a new version of H1 hemagglutinin. Data suggest that Dr. Baker’s proteins bind to all types of the group I hemagglutinin, a group that includes not only H1 but also the pandemic H2 and avian H5 strains.
The methods developed for the influenza inhibitor protein design, according to Dr. Baker, could be “a powerful route to inhibitors or binders for any surface patch on any desired target of interest.” For example, if a new disease pathogen arises, scientists could figure out how it interacts with human cells or other hosts on a molecular level. Scientists could then employ protein interface design to create a diversity of small proteins that they predict would block the pathogen’s interaction surface.
Genes for large numbers of the most promising, computer-designed proteins could be tested using yeast cells. After additional molecular chemistry research to search for the best binding among those proteins, those could be reprogrammed in the laboratory to undergo mutations, and all the mutated forms could be stored in a “library” for an in-depth examination of their amino acids, molecular architecture, and energy bonds.
Sophisticated technologies would allow the scientists to rapidly browse through the library to find those tiny proteins that clung to the pathogen surface target with pinpoint accuracy. The finalists would be selected from this pool for excelling at blocking the pathogen from attaching to, entering, and infecting human or animal cells.
The utilization of deep sequencing, the same technology now used to sequence human genomes cheaply, was particularly central in creating detailed maps relating sequencing to function. These maps were used to reprogram the design to achieve a more exact interaction between the inhibitor protein and the virus molecule. It also enabled the scientists, they said, “to leapfrog over bottlenecks” to improve the activity of the binder. They were able to see how small contributions from many small alterations in the protein, too difficult to see individually, could together create a binder with better attachment strength.
“We anticipate that our approach combining computational design followed by comprehensive energy landscape mapping,” Dr. Baker said, “will be widely useful in generating high-affinity and high-specificity binders to a broad range of targets for use in therapeutics and diagnostics.”
Related Links:
University of Washington
Latest BioResearch News
- Nanopore Method Captures RNA Folding at Single-Molecule Resolution
- Tumor Microenvironment Marker Linked to Worse Survival in Solid Tumors
- Hidden Immune Gene Defect May Explain Kaposi Sarcoma Susceptibility
- Genetic Markers May Help Predict Amputation Risk in Peripheral Artery Disease
- Gene Signature Shows Promise for Depression Biomarker Testing
- AI-Driven Tumor Profiling Initiative Targets Precision Therapy Development
- Researchers Map Protein and Glycosylation Across 15 Human Body Fluids
- Telomere Length Abnormalities Linked to Lymphoma Development
- Biomarker Signals Chemotherapy Resistance in Relapsed Small Cell Lung Cancer
- Inflammatory Gene Signature Links Metabolic Disease to Pancreatic Cancer Recurrence
- Study Links Abnormal Gene Splicing to Treatment Response in Metastatic Kidney Cancer
- Research Reveals How Some Aplastic Anemia Patients Recover Bone Marrow Function
- New Molecular Insights Support Diagnosis of Hodgkin Lymphoma
- Epigenetic Signals and Blood Markers Aid Chronic Fatigue Syndrome Diagnosis
- Microenvironment Biomarkers Could Enable Early Lung Cancer Detection
- Study Identifies Protein Changes Driving Immunotherapy Resistance in Multiple Myeloma
Channels
Clinical Chemistry
view channel
Long-Term Data Show PSA Screening Modestly Reduces Prostate Cancer Deaths
Prostate cancer is among the most common cancers in men, and the role of population screening has remained controversial because of overdiagnosis and overtreatment. Health systems have sought clearer,... Read more
Urine-Based Nanosensor Tracks Lung Cancer and Fibrosis Noninvasively
Lung cancer remains difficult to monitor for early progression and treatment resistance, while pulmonary fibrosis continues to pose major challenges for early diagnosis. Clinicians need repeatable, noninvasive... Read moreMolecular Diagnostics
view channel
New Computational Tool Reveals Genetic Driver of Idiopathic Neuropathy
Peripheral neuropathy is a common neurological disorder that causes pain, sensory loss, imbalance, and weakness, affecting an estimated 12%–20% of people in the U.S. and nearly 30% of adults over age 65.... Read more
Breast Cancer-Specific Signatures Link Genome Instability to Outcomes
Genomic instability is a hallmark of cancer, but most genomic analyses have relied on broad signatures shared across multiple malignancies, limiting their precision for individual tumor types.... Read moreHematology
view channel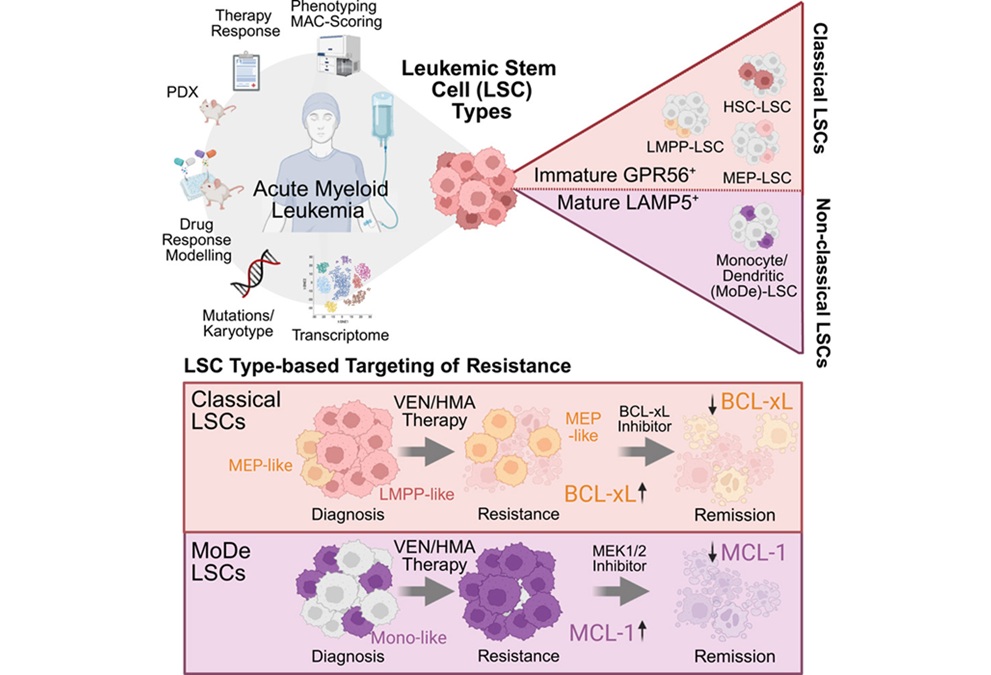
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read more
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read moreImmunology
view channel
Routine TB Screening Test May Reveal Immune Aging and Mortality Risk
Immune aging is associated with weaker responses to vaccination, greater risks of infection, and higher levels of inflammation. Leveraging routinely ordered laboratory tests to quantify that responsiveness... Read more
Biomarkers and Molecular Testing Advance Precision Allergy Care
Allergic diseases often present with similar symptoms but can be driven by distinct biological mechanisms, making standardized care inefficient for many patients. Historically, individuals with pollen... Read moreMicrobiology
view channel
Study Finds Hidden Mpox Infections May Drive Ongoing Spread
Mpox continues to circulate despite vaccination, and many cases show no known link to a symptomatic partner. The role of people without symptoms has remained uncertain, limiting clarity on how transmission persists.... Read more
Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
Antimicrobial resistance (AMR) poses a growing threat to patient safety, with carbapenem-resistant Enterobacterales causing difficult-to-treat infections and leaving clinicians with limited therapeutic options.... Read more
Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
Tuberculosis is the leading cause of death among people living with HIV, and diagnosis in hospital settings remains difficult. Symptoms are often non-specific, disease can be extrapulmonary, and many patients... Read morePathology
view channel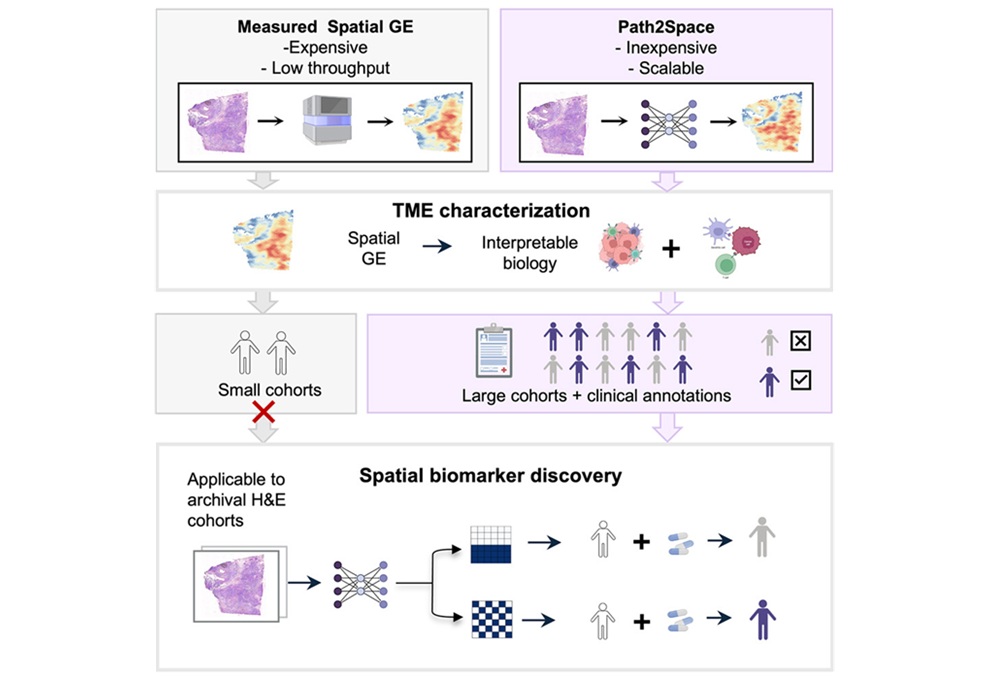
Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
Gene expression profiling can inform tumor biology and treatment selection, but spatial assays remain costly and time-consuming. Results can take weeks and cost thousands of dollars, limiting large-scale... Read more
AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
Non–muscle invasive bladder cancer has highly variable outcomes, complicating surveillance and treatment planning. Risk assessment typically relies on stage, grade, and tumor size, leaving uncertainty... Read moreTechnology
view channel
AI Tool Automates Validation of Laboratory Software Configuration Changes
Regulated laboratories face heavy documentation and requalification demands when software configurations change, slowing improvements and discouraging beneficial updates. A new capability now automates... Read more
Point-of-Care Testing Enhances Health Literacy and Self-Management in Chronic Disease
Limited access to general practitioners and pathology services can delay diagnosis and monitoring for people in regional and remote communities. Rapid, on-the-spot testing can shorten turnaround times... Read moreIndustry
view channel
AI-Powered Multi-Functional Analyzer Wins German Innovation Award
Hematology services are increasingly delivered across distributed care settings, where limited staffing and complex workflows can extend turnaround times. Advanced morphology review still often depends... Read more