DNA Repair Map Expected to Enhance Human Genome Studies
|
By LabMedica International staff writers Posted on 10 May 2015 |
Researchers have created a map of the entire human genome that shows where repairs to DNA that has been damaged by radiation or chemical treatment are made.
Investigators at the University of North Carolina (Chapel Hill, USA) developed a technique called XR-Seq (excision repair sequencing) in order to prepare the map, which was presented in a paper published in the May 1, 2015, issue of the journal Genes & Development.
Human nucleotide excision repair generates two incisions surrounding the site of damage, creating a polymer of approximately 30 repeating units. In XR-seq, this fragment is isolated and subjected to high-throughput sequencing.
The investigators used XR-seq to produce stranded, nucleotide-resolution maps of repair of two UV-induced DNA damages in human cells: cyclobutane pyrimidine dimers (CPDs) and (6-4) pyrimidine–pyrimidone photoproducts [(6-4)PPs]. In wild-type cells, CPD repair was highly associated with transcription, specifically with the template strand.
Experiments in cells defective in either transcription-coupled excision repair or general excision repair isolated the contribution of each pathway to the overall repair pattern and showed that transcription-coupled repair of both photoproducts occurred exclusively on the template strand. XR-seq maps captured transcription-coupled repair at sites of divergent gene promoters and bidirectional enhancer RNA (eRNA) production at enhancers. XR-seq data also uncovered the repair characteristics and novel sequence preferences of CPDs and (6-4)PPs.
"Now we can say to a fellow scientist, tell us the gene you are interested in or any spot on the genome, and we will tell you how it is repaired," said senior author Dr. Aziz Sancar, professor of biochemistry and biophysics at the University of North Carolina. "Out of six billion base pairs, pick out a spot and we will tell you how it is repaired."
It is expected that XR-seq and the resulting repair maps will facilitate studies of the effects of genomic location, chromatin context, transcription, and replication on DNA repair in human cells.
Related Links:
University of North Carolina
Investigators at the University of North Carolina (Chapel Hill, USA) developed a technique called XR-Seq (excision repair sequencing) in order to prepare the map, which was presented in a paper published in the May 1, 2015, issue of the journal Genes & Development.
Human nucleotide excision repair generates two incisions surrounding the site of damage, creating a polymer of approximately 30 repeating units. In XR-seq, this fragment is isolated and subjected to high-throughput sequencing.
The investigators used XR-seq to produce stranded, nucleotide-resolution maps of repair of two UV-induced DNA damages in human cells: cyclobutane pyrimidine dimers (CPDs) and (6-4) pyrimidine–pyrimidone photoproducts [(6-4)PPs]. In wild-type cells, CPD repair was highly associated with transcription, specifically with the template strand.
Experiments in cells defective in either transcription-coupled excision repair or general excision repair isolated the contribution of each pathway to the overall repair pattern and showed that transcription-coupled repair of both photoproducts occurred exclusively on the template strand. XR-seq maps captured transcription-coupled repair at sites of divergent gene promoters and bidirectional enhancer RNA (eRNA) production at enhancers. XR-seq data also uncovered the repair characteristics and novel sequence preferences of CPDs and (6-4)PPs.
"Now we can say to a fellow scientist, tell us the gene you are interested in or any spot on the genome, and we will tell you how it is repaired," said senior author Dr. Aziz Sancar, professor of biochemistry and biophysics at the University of North Carolina. "Out of six billion base pairs, pick out a spot and we will tell you how it is repaired."
It is expected that XR-seq and the resulting repair maps will facilitate studies of the effects of genomic location, chromatin context, transcription, and replication on DNA repair in human cells.
Related Links:
University of North Carolina
Latest BioResearch News
- Genome Analysis Predicts Likelihood of Neurodisability in Oxygen-Deprived Newborns
- Gene Panel Predicts Disease Progession for Patients with B-cell Lymphoma
- New Method Simplifies Preparation of Tumor Genomic DNA Libraries
- New Tool Developed for Diagnosis of Chronic HBV Infection
- Panel of Genetic Loci Accurately Predicts Risk of Developing Gout
- Disrupted TGFB Signaling Linked to Increased Cancer-Related Bacteria
- Gene Fusion Protein Proposed as Prostate Cancer Biomarker
- NIV Test to Diagnose and Monitor Vascular Complications in Diabetes
- Semen Exosome MicroRNA Proves Biomarker for Prostate Cancer
- Genetic Loci Link Plasma Lipid Levels to CVD Risk
- Newly Identified Gene Network Aids in Early Diagnosis of Autism Spectrum Disorder
- Link Confirmed between Living in Poverty and Developing Diseases
- Genomic Study Identifies Kidney Disease Loci in Type I Diabetes Patients
- Liquid Biopsy More Effective for Analyzing Tumor Drug Resistance Mutations
- New Liquid Biopsy Assay Reveals Host-Pathogen Interactions
- Method Developed for Enriching Trophoblast Population in Samples
Channels
Clinical Chemistry
view channel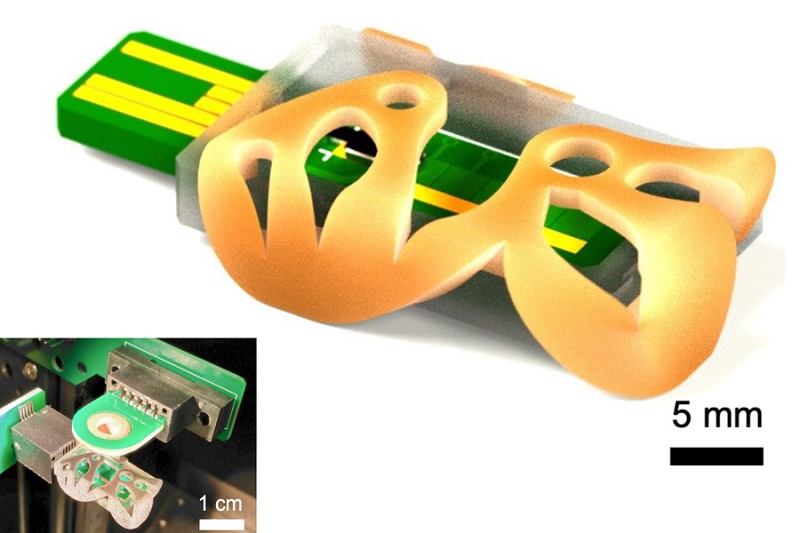
3D Printed Point-Of-Care Mass Spectrometer Outperforms State-Of-The-Art Models
Mass spectrometry is a precise technique for identifying the chemical components of a sample and has significant potential for monitoring chronic illness health states, such as measuring hormone levels... Read more.jpg)
POC Biomedical Test Spins Water Droplet Using Sound Waves for Cancer Detection
Exosomes, tiny cellular bioparticles carrying a specific set of proteins, lipids, and genetic materials, play a crucial role in cell communication and hold promise for non-invasive diagnostics.... Read more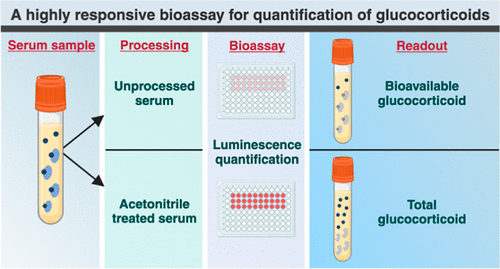
Highly Reliable Cell-Based Assay Enables Accurate Diagnosis of Endocrine Diseases
The conventional methods for measuring free cortisol, the body's stress hormone, from blood or saliva are quite demanding and require sample processing. The most common method, therefore, involves collecting... Read moreMolecular Diagnostics
view channel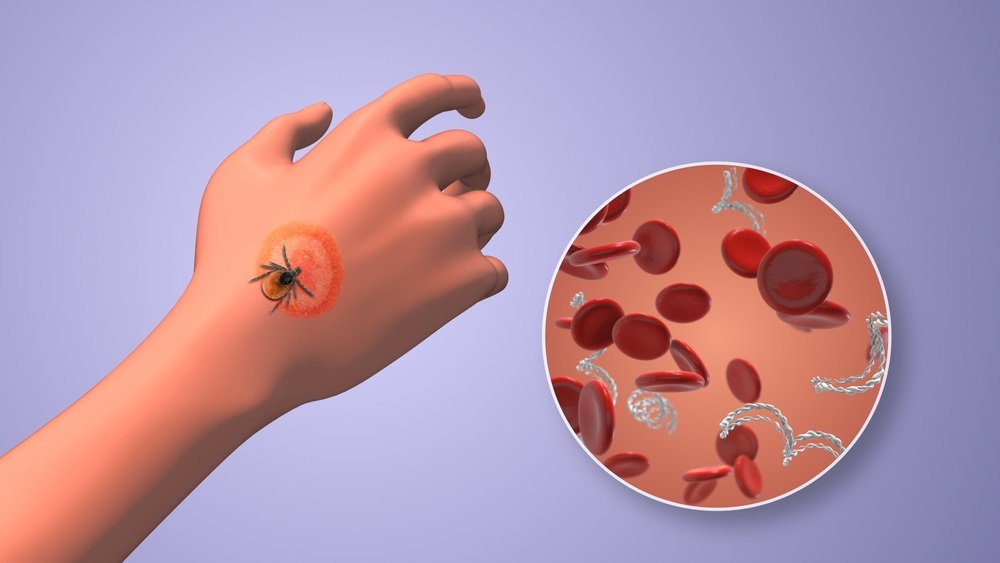
Urine Test to Revolutionize Lyme Disease Testing
Lyme disease is the most common animal-to-human transmitted disease in the United States, with around 476,000 people diagnosed and treated annually, and its incidence has been increasing.... Read more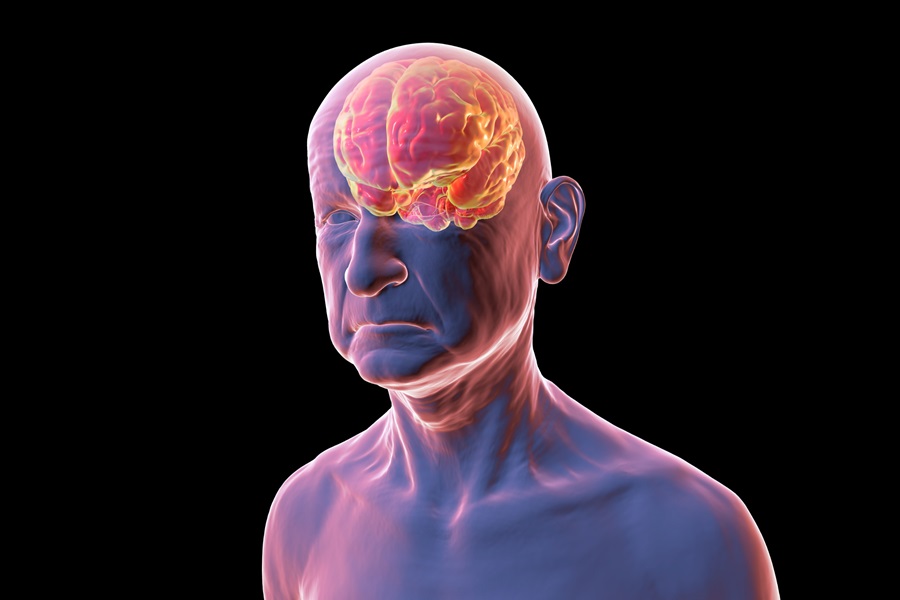
Simple Blood Test Could Enable First Quantitative Assessments for Future Cerebrovascular Disease
Cerebral small vessel disease is a common cause of stroke and cognitive decline, particularly in the elderly. Presently, assessing the risk for cerebral vascular diseases involves using a mix of diagnostic... Read more
New Genetic Testing Procedure Combined With Ultrasound Detects High Cardiovascular Risk
A key interest area in cardiovascular research today is the impact of clonal hematopoiesis on cardiovascular diseases. Clonal hematopoiesis results from mutations in hematopoietic stem cells and may lead... Read moreHematology
view channel
Next Generation Instrument Screens for Hemoglobin Disorders in Newborns
Hemoglobinopathies, the most widespread inherited conditions globally, affect about 7% of the population as carriers, with 2.7% of newborns being born with these conditions. The spectrum of clinical manifestations... Read more
First 4-in-1 Nucleic Acid Test for Arbovirus Screening to Reduce Risk of Transfusion-Transmitted Infections
Arboviruses represent an emerging global health threat, exacerbated by climate change and increased international travel that is facilitating their spread across new regions. Chikungunya, dengue, West... Read more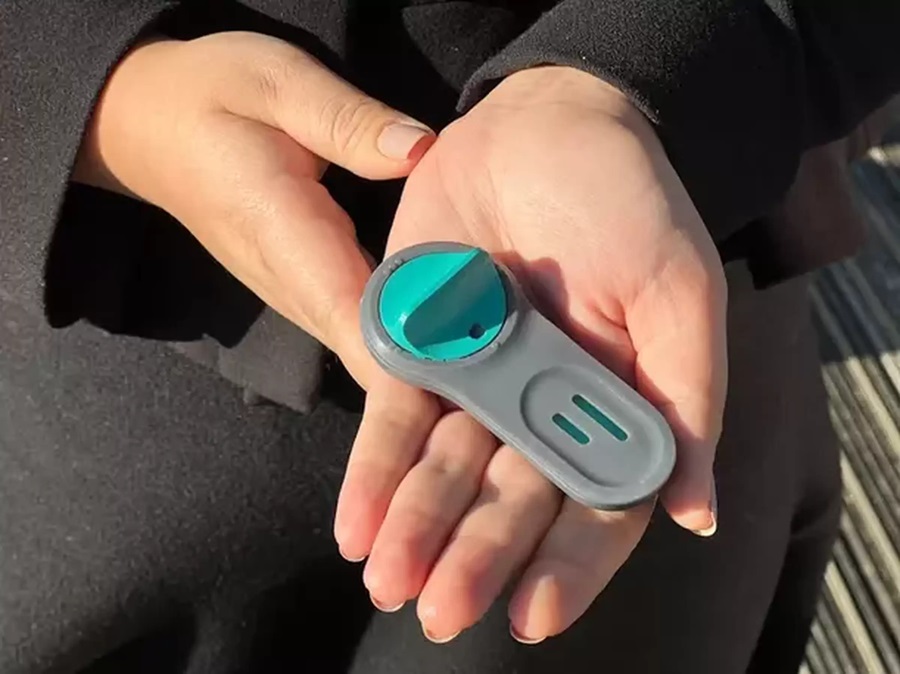
POC Finger-Prick Blood Test Determines Risk of Neutropenic Sepsis in Patients Undergoing Chemotherapy
Neutropenia, a decrease in neutrophils (a type of white blood cell crucial for fighting infections), is a frequent side effect of certain cancer treatments. This condition elevates the risk of infections,... Read more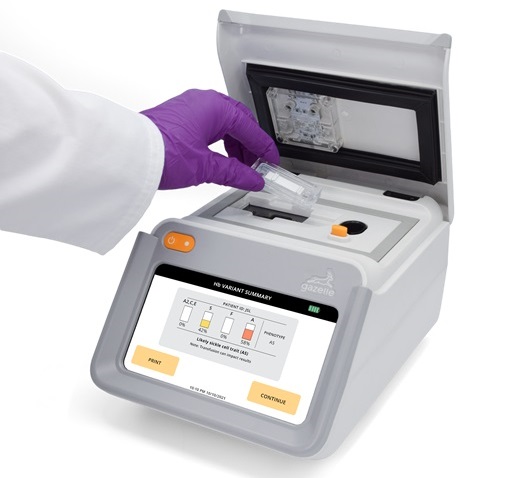
First Affordable and Rapid Test for Beta Thalassemia Demonstrates 99% Diagnostic Accuracy
Hemoglobin disorders rank as some of the most prevalent monogenic diseases globally. Among various hemoglobin disorders, beta thalassemia, a hereditary blood disorder, affects about 1.5% of the world's... Read moreImmunology
view channel
Diagnostic Blood Test for Cellular Rejection after Organ Transplant Could Replace Surgical Biopsies
Transplanted organs constantly face the risk of being rejected by the recipient's immune system which differentiates self from non-self using T cells and B cells. T cells are commonly associated with acute... Read more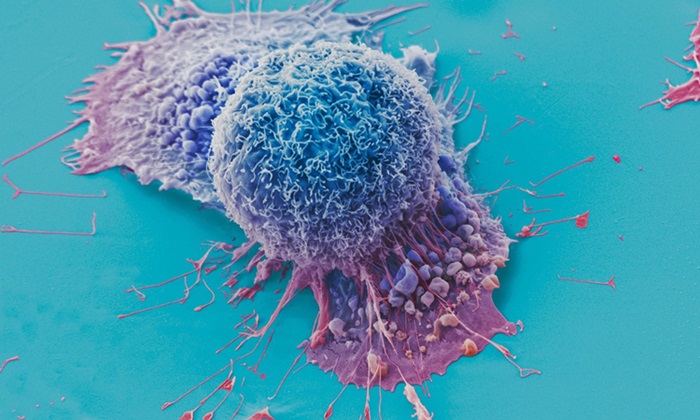
AI Tool Precisely Matches Cancer Drugs to Patients Using Information from Each Tumor Cell
Current strategies for matching cancer patients with specific treatments often depend on bulk sequencing of tumor DNA and RNA, which provides an average profile from all cells within a tumor sample.... Read more
Genetic Testing Combined With Personalized Drug Screening On Tumor Samples to Revolutionize Cancer Treatment
Cancer treatment typically adheres to a standard of care—established, statistically validated regimens that are effective for the majority of patients. However, the disease’s inherent variability means... Read moreMicrobiology
view channelEnhanced Rapid Syndromic Molecular Diagnostic Solution Detects Broad Range of Infectious Diseases
GenMark Diagnostics (Carlsbad, CA, USA), a member of the Roche Group (Basel, Switzerland), has rebranded its ePlex® system as the cobas eplex system. This rebranding under the globally renowned cobas name... Read more
Clinical Decision Support Software a Game-Changer in Antimicrobial Resistance Battle
Antimicrobial resistance (AMR) is a serious global public health concern that claims millions of lives every year. It primarily results from the inappropriate and excessive use of antibiotics, which reduces... Read more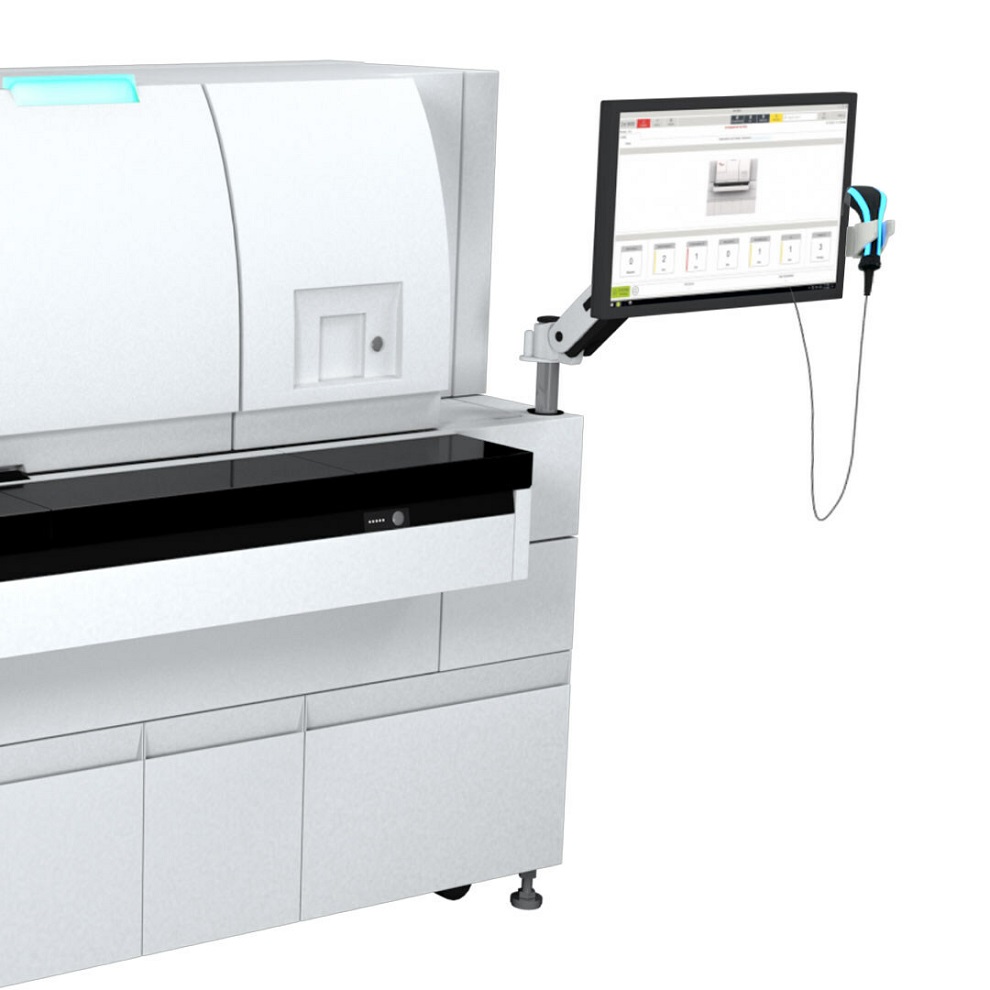
New CE-Marked Hepatitis Assays to Help Diagnose Infections Earlier
According to the World Health Organization (WHO), an estimated 354 million individuals globally are afflicted with chronic hepatitis B or C. These viruses are the leading causes of liver cirrhosis, liver... Read more
1 Hour, Direct-From-Blood Multiplex PCR Test Identifies 95% of Sepsis-Causing Pathogens
Sepsis contributes to one in every three hospital deaths in the US, and globally, septic shock carries a mortality rate of 30-40%. Diagnosing sepsis early is challenging due to its non-specific symptoms... Read morePathology
view channel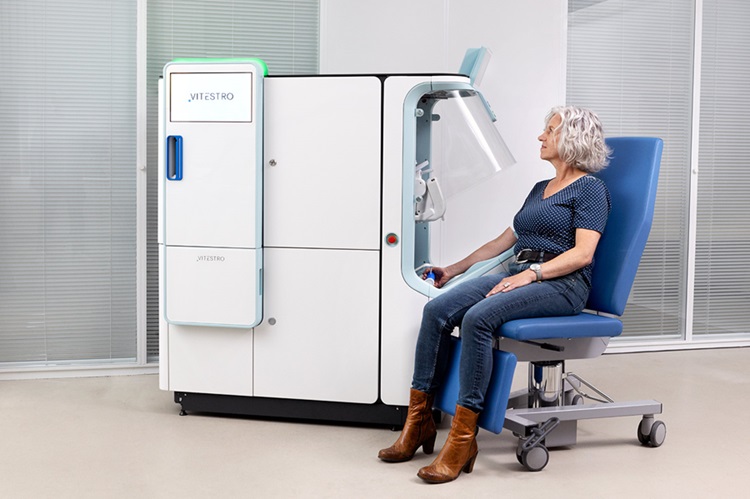
Robotic Blood Drawing Device to Revolutionize Sample Collection for Diagnostic Testing
Blood drawing is performed billions of times each year worldwide, playing a critical role in diagnostic procedures. Despite its importance, clinical laboratories are dealing with significant staff shortages,... Read more.jpg)
Use of DICOM Images for Pathology Diagnostics Marks Significant Step towards Standardization
Digital pathology is rapidly becoming a key aspect of modern healthcare, transforming the practice of pathology as laboratories worldwide adopt this advanced technology. Digital pathology systems allow... Read more
First of Its Kind Universal Tool to Revolutionize Sample Collection for Diagnostic Tests
The COVID pandemic has dramatically reshaped the perception of diagnostics. Post the pandemic, a groundbreaking device that combines sample collection and processing into a single, easy-to-use disposable... Read moreTechnology
view channel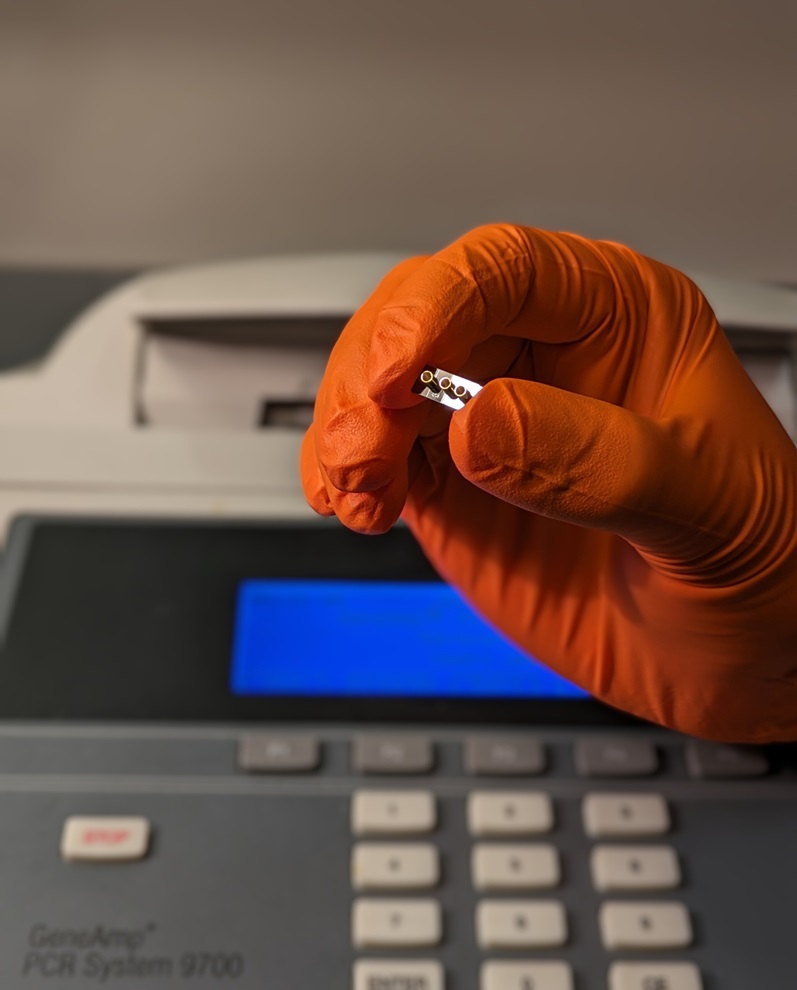
New Diagnostic System Achieves PCR Testing Accuracy
While PCR tests are the gold standard of accuracy for virology testing, they come with limitations such as complexity, the need for skilled lab operators, and longer result times. They also require complex... Read more
DNA Biosensor Enables Early Diagnosis of Cervical Cancer
Molybdenum disulfide (MoS2), recognized for its potential to form two-dimensional nanosheets like graphene, is a material that's increasingly catching the eye of the scientific community.... Read more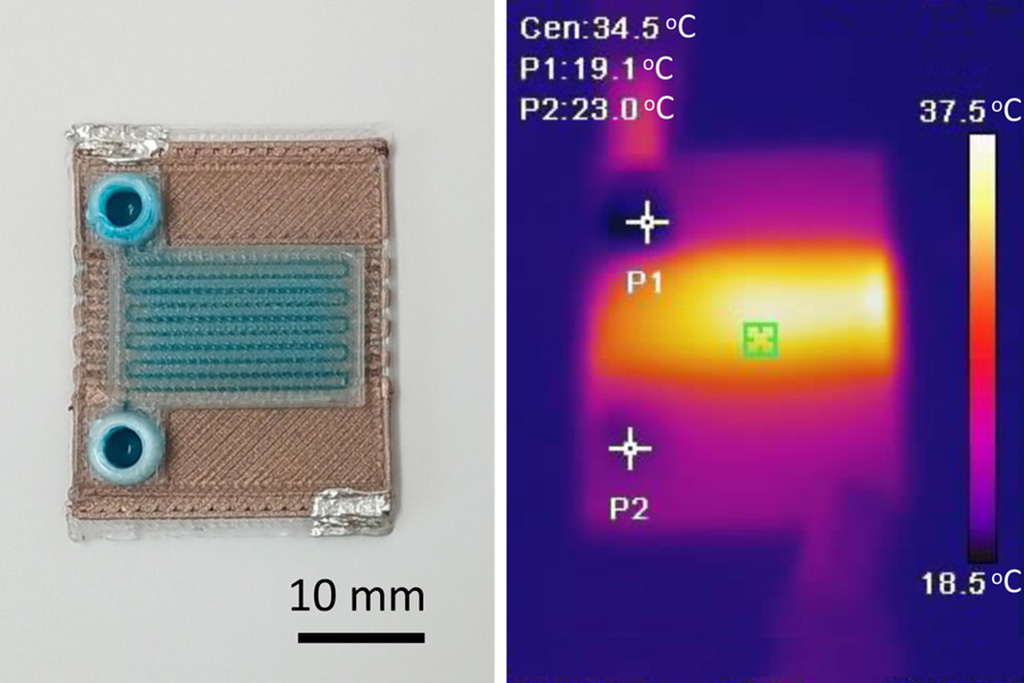
Self-Heating Microfluidic Devices Can Detect Diseases in Tiny Blood or Fluid Samples
Microfluidics, which are miniature devices that control the flow of liquids and facilitate chemical reactions, play a key role in disease detection from small samples of blood or other fluids.... Read more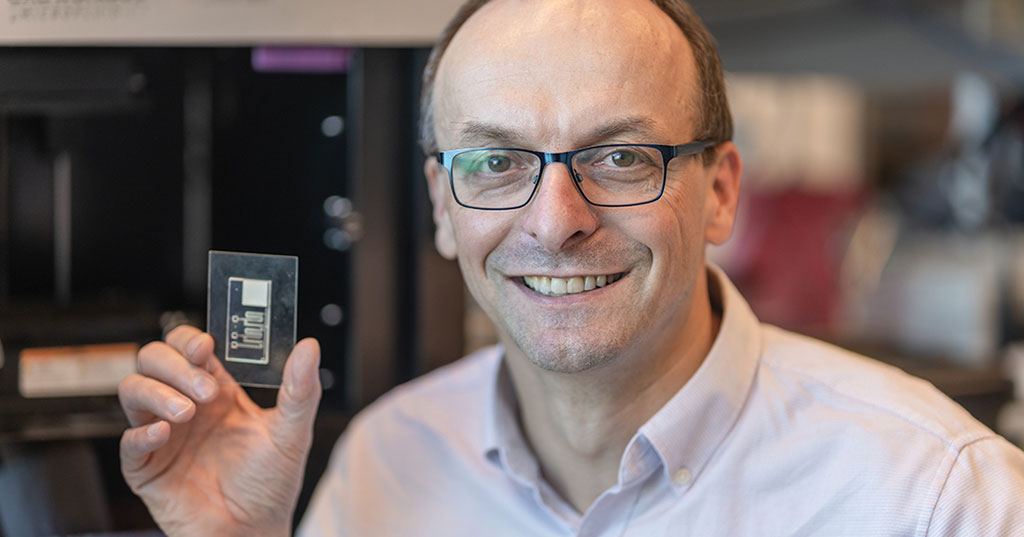
Breakthrough in Diagnostic Technology Could Make On-The-Spot Testing Widely Accessible
Home testing gained significant importance during the COVID-19 pandemic, yet the availability of rapid tests is limited, and most of them can only drive one liquid across the strip, leading to continued... Read moreIndustry
view channel_1.jpg)
Thermo Fisher and Bio-Techne Enter Into Strategic Distribution Agreement for Europe
Thermo Fisher Scientific (Waltham, MA USA) has entered into a strategic distribution agreement with Bio-Techne Corporation (Minneapolis, MN, USA), resulting in a significant collaboration between two industry... Read more
ECCMID Congress Name Changes to ESCMID Global
Over the last few years, the European Society of Clinical Microbiology and Infectious Diseases (ESCMID, Basel, Switzerland) has evolved remarkably. The society is now stronger and broader than ever before... Read more