Multiple Microbiological Tests Needed for Underweight Newborns
|
By LabMedica International staff writers Posted on 03 Mar 2013 |
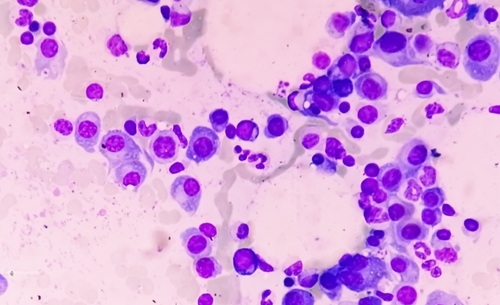
Image: The oral bacteria Fusobacterium nucleatum (Photo courtesy of HealthyDent).
Cultures commonly used to detect bacterial infections in low birth-weight newborns with early onset sepsis may fail to detect some microorganisms.
There is a need for multiple detection methods, such as DNA genomic analyses and other independent culture technologies, to identify bacteria that routine culturing may miss.
Scientists at Case Western Reserve University (Cleveland, OH, USA) performed a comparative microbial analysis of paired amniotic fluid (AF) and cord blood (CB) from pregnancies complicated by preterm birth and early-onset neonatal sepsis. The biological samples from 44 women were collected from September 2004 to February 2009.
Amniotic fluid (AF) was cultured for aerobic and anaerobic bacteria, Ureaplasma and Mycoplasma species. DNA was extracted from AF or CB serum. To identify the species amplified by polymerase chain reaction (PCR) and to ensure that the PCR amplicons were indeed bacterial ribosomal ribonucleic acid (rRNA) genes rather than artifacts, the PCR products were cloned into the pCR8 vector (Invitrogen, Carlsbad, CA, USA).
The investigators found more than 20 bacterial species not discovered using standard culturing. Some of the uncultured species appeared in both the cord blood and amniotic fluid samples. The uncultured bacteria were detected with DNA genomic analysis that had been used in a prior study that discovered the link between oral bacteria that causes still- or premature-births due to infected amniotic fluid that is supposed to be a sterile environment.
Yiping Han, PhD, the professor of Periodontics and Reproductive Biology at the Case Western, said, "Culture independent technology has broadened our scope of understanding human pathogens. DNA testing techniques were able for the first time to detect the oral bacteria Fusobacterium nucleatum, Bergeyella, and Sneathia sanguinegens that brought on early neonatal sepsis and put newborns at risk of dying shortly after birth. Among these, F. nucleatum was found at the same high frequency as the well-known Escherichia coli, putting the former on the same importance scale as the latter." The study was published on February 20, 2013, in the journal Public Library of Science ONE.
Related Links:
Case Western Reserve University
Invitrogen
There is a need for multiple detection methods, such as DNA genomic analyses and other independent culture technologies, to identify bacteria that routine culturing may miss.
Scientists at Case Western Reserve University (Cleveland, OH, USA) performed a comparative microbial analysis of paired amniotic fluid (AF) and cord blood (CB) from pregnancies complicated by preterm birth and early-onset neonatal sepsis. The biological samples from 44 women were collected from September 2004 to February 2009.
Amniotic fluid (AF) was cultured for aerobic and anaerobic bacteria, Ureaplasma and Mycoplasma species. DNA was extracted from AF or CB serum. To identify the species amplified by polymerase chain reaction (PCR) and to ensure that the PCR amplicons were indeed bacterial ribosomal ribonucleic acid (rRNA) genes rather than artifacts, the PCR products were cloned into the pCR8 vector (Invitrogen, Carlsbad, CA, USA).
The investigators found more than 20 bacterial species not discovered using standard culturing. Some of the uncultured species appeared in both the cord blood and amniotic fluid samples. The uncultured bacteria were detected with DNA genomic analysis that had been used in a prior study that discovered the link between oral bacteria that causes still- or premature-births due to infected amniotic fluid that is supposed to be a sterile environment.
Yiping Han, PhD, the professor of Periodontics and Reproductive Biology at the Case Western, said, "Culture independent technology has broadened our scope of understanding human pathogens. DNA testing techniques were able for the first time to detect the oral bacteria Fusobacterium nucleatum, Bergeyella, and Sneathia sanguinegens that brought on early neonatal sepsis and put newborns at risk of dying shortly after birth. Among these, F. nucleatum was found at the same high frequency as the well-known Escherichia coli, putting the former on the same importance scale as the latter." The study was published on February 20, 2013, in the journal Public Library of Science ONE.
Related Links:
Case Western Reserve University
Invitrogen
Latest Microbiology News
- Gut Microbiome Signatures Help Identify Risk of IBD Progression
- FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
- New AMR Assay Supports Rapid Infection Control Screening in Hospitals
- Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- Rapid Color Test Stratifies Virulent and Resistant Staph Strains
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Syndromic Panel Enables Rapid Identification of Bloodstream Infections
Channels
Clinical Chemistry
view channel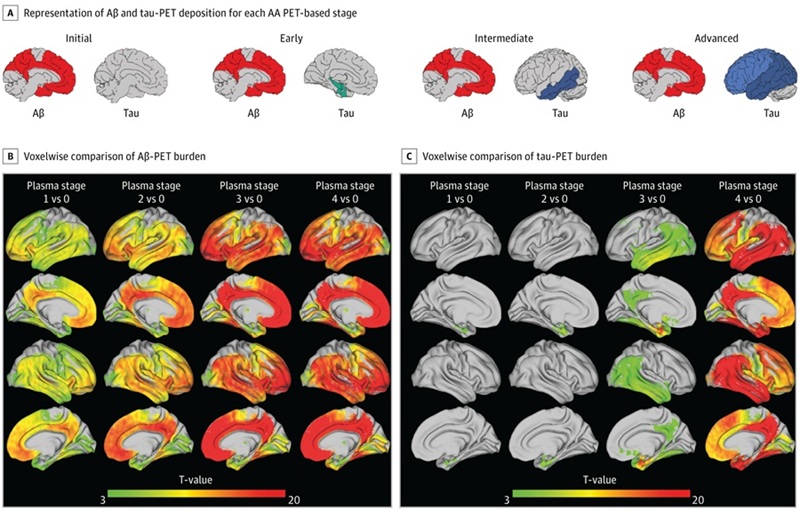
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreMolecular Diagnostics
view channel
Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
Selecting adjuvant therapy for early-stage hormone receptor-positive breast cancer depends on accurately assessing long-term risk of distant recurrence. Clinical features alone can leave uncertainty about... Read more
Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
Many cancers deviate from the normal chromosome number, but the clinical impact of extreme chromosome loss remains unclear. This widespread genomic disruption is associated with aggressive disease and... Read more
Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
Sarcomas remain difficult to monitor during immunotherapy, as low tumor mutation burden can limit traditional circulating tumor DNA approaches and repeat tissue biopsies are often impractical in advanced disease.... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more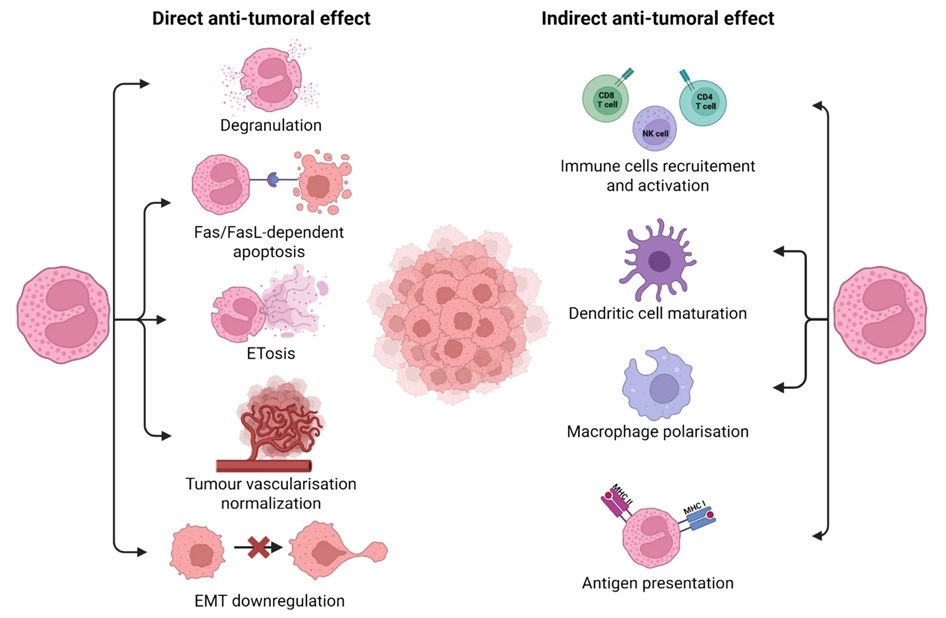
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more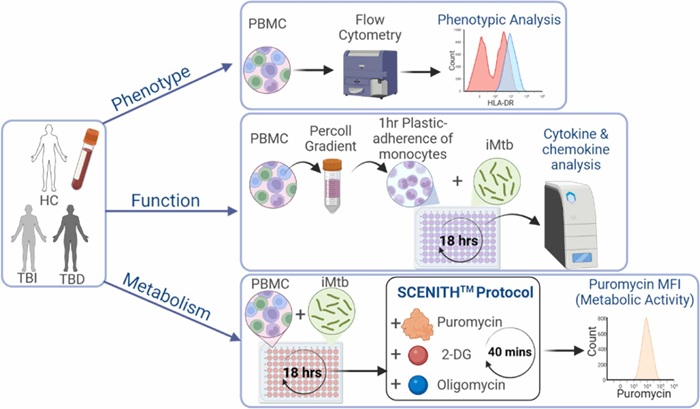
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read morePathology
view channel
Blood-Based Method Tracks Gene Activity in the Living Brain
Real-time measurement of gene activity in the brain has been limited by assays requiring destructive tissue sampling. Tracking active genes could reveal how the body responds to environmental factors,... Read more
FDA Approval Expands Automated PD-L1 Testing Across Solid Tumors
Clinical laboratories play a central role in guiding immunotherapy by reporting programmed death ligand-1 (PD‑L1) status across multiple solid tumors. Many sites are standardizing this work on fully automated... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel