Novel Method Reclaims Resolution of Single-Cell RNA-Seq
|
By LabMedica International staff writers Posted on 28 Oct 2020 |
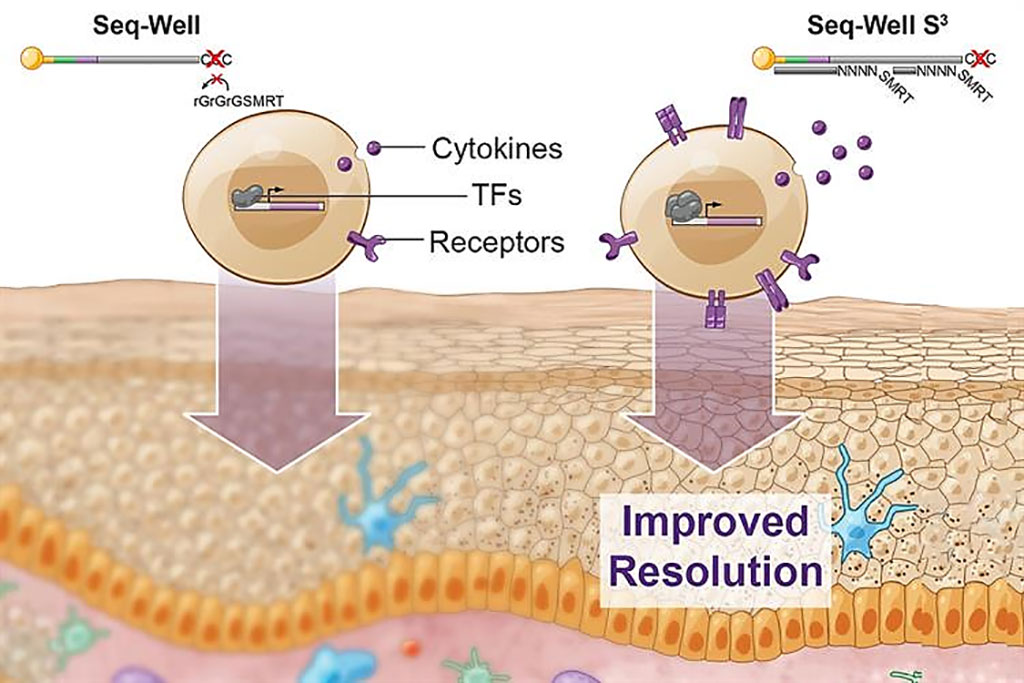
Image: Scientists have greatly boosted the amount of information that can be obtained using Seq-Well S3, a technique for rapidly sequencing RNA from single cells (Photo courtesy of MIT).
Single-cell RNA sequencing (scRNA-seq) is a powerful tool to characterize cells. Current scRNA-seq platforms, despite offering high throughput, are inefficient and provide low resolution among distinct cell states and molecular features.
Most high-throughput scRNA-seq methods rely on barcoding of cellular components to recover single-cell transcriptomes for thousands of cells at once. This is achieved by isolating uniquely barcoded poly-dT oligonucleotides that can capture and tag cellular messenger RNA (mRNA) during reverse transcription. In a second step, an additional oligonucleotide priming site is added to newly synthesized complementary DNA (cDNA) to enable polymerase chain reaction (PCR)-based amplification.
Medical Biochemists at the Massachusetts Institute of Technology (Cambridge, MA, USA) and their associates developed Seq-Well S3 ("Second-Strand Synthesis") as a massively parallel scRNA-seq protocol that uses a randomly primed second-strand synthesis to recover cDNA molecules to facilitate template-switching. This generates double-stranded cDNA that is labeled on one end with the SMART sequence and its reverse complement on the other, making it more accessible for PCR enzymes to amplify the molecules.
To perform the study skin biopsies were obtained from a total of 16 patients at the University of California, Los Angeles and University of Southern California Hansen’s Clinic, while an additional three samples were obtained from the University of Michigan. The team utilized Seq-Well, a massively parallel, low-input scRNA-seq platform for clinical samples, to capture the transcriptome of single cells. The team performed Templated Second-Strand Synthesis, PCR Amplification, Optimization of Second-Strand Synthesis, CD4+ T Cell comparisons of 10x Genomics, Pleasanton, CA, USA), Seq-Well S3, and Smart-Seq2, DNA Sequencing and Alignment of peripheral blood mononuclear cells (PBMC) optimization samples, and tissue immunofluorescence staining.
In total, the scientists processed 19 skin biopsies and retained over 38,000 high-quality single-cell transcriptomes using Seq-Well S3. They were able to recover 15 primary cell types. To further define biological features, the team used the method to examine subpopulations of T cells, myeloid cells, endothelial cells, dermal fibroblasts, and keratinocytes in each inflammatory condition. The team found, for example, regulatory T cells, dysfunctional NR4A1-expressing T cells, and senescent SESN3+ T cells were over-represented, potentially reflecting T-cell dysfunction in psoriasis pathology.
The team also distinguished patterns associated with multiple diseases by looking across different inflammatory skin conditions, revealing common and unique features. For instance, they found that a group of natural killer cells, γΔ T cells, and a sub-cluster of immature cytotoxic T cells are derived from leprosy and granuloma annulare, indicating common T-cell programming in both forms of inflammation.
Alex Shalek, PhD, associate professor of chemistry at MIT and a senior author of the study, said, “"It's become clear that these technologies have transformative potential for understanding complex biological systems. If we look across a range of different datasets, we can really understand the landscape of health and disease, and that can give us information as to what therapeutic strategies we might employ.” The study was published on October 13, 2020 in the journal Immunity.
Related Links:
Massachusetts Institute of Technology
10x Genomics
Most high-throughput scRNA-seq methods rely on barcoding of cellular components to recover single-cell transcriptomes for thousands of cells at once. This is achieved by isolating uniquely barcoded poly-dT oligonucleotides that can capture and tag cellular messenger RNA (mRNA) during reverse transcription. In a second step, an additional oligonucleotide priming site is added to newly synthesized complementary DNA (cDNA) to enable polymerase chain reaction (PCR)-based amplification.
Medical Biochemists at the Massachusetts Institute of Technology (Cambridge, MA, USA) and their associates developed Seq-Well S3 ("Second-Strand Synthesis") as a massively parallel scRNA-seq protocol that uses a randomly primed second-strand synthesis to recover cDNA molecules to facilitate template-switching. This generates double-stranded cDNA that is labeled on one end with the SMART sequence and its reverse complement on the other, making it more accessible for PCR enzymes to amplify the molecules.
To perform the study skin biopsies were obtained from a total of 16 patients at the University of California, Los Angeles and University of Southern California Hansen’s Clinic, while an additional three samples were obtained from the University of Michigan. The team utilized Seq-Well, a massively parallel, low-input scRNA-seq platform for clinical samples, to capture the transcriptome of single cells. The team performed Templated Second-Strand Synthesis, PCR Amplification, Optimization of Second-Strand Synthesis, CD4+ T Cell comparisons of 10x Genomics, Pleasanton, CA, USA), Seq-Well S3, and Smart-Seq2, DNA Sequencing and Alignment of peripheral blood mononuclear cells (PBMC) optimization samples, and tissue immunofluorescence staining.
In total, the scientists processed 19 skin biopsies and retained over 38,000 high-quality single-cell transcriptomes using Seq-Well S3. They were able to recover 15 primary cell types. To further define biological features, the team used the method to examine subpopulations of T cells, myeloid cells, endothelial cells, dermal fibroblasts, and keratinocytes in each inflammatory condition. The team found, for example, regulatory T cells, dysfunctional NR4A1-expressing T cells, and senescent SESN3+ T cells were over-represented, potentially reflecting T-cell dysfunction in psoriasis pathology.
The team also distinguished patterns associated with multiple diseases by looking across different inflammatory skin conditions, revealing common and unique features. For instance, they found that a group of natural killer cells, γΔ T cells, and a sub-cluster of immature cytotoxic T cells are derived from leprosy and granuloma annulare, indicating common T-cell programming in both forms of inflammation.
Alex Shalek, PhD, associate professor of chemistry at MIT and a senior author of the study, said, “"It's become clear that these technologies have transformative potential for understanding complex biological systems. If we look across a range of different datasets, we can really understand the landscape of health and disease, and that can give us information as to what therapeutic strategies we might employ.” The study was published on October 13, 2020 in the journal Immunity.
Related Links:
Massachusetts Institute of Technology
10x Genomics
Latest Immunology News
- Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
- Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
- Routine TB Screening Test May Reveal Immune Aging and Mortality Risk
- Biomarkers and Molecular Testing Advance Precision Allergy Care
- Point-of-Care Tests Could Expand Access to Mpox Diagnosis
- T-Cell Senescence Profiling May Predict CAR T Responses
- Finger-Prick Lateral Flow Test Detects Sepsis Biomarkers at Point of Care
- Study Highlights Low Sensitivity of Current Lyme Tests in Early Infection
- Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
- Study Finds Influenza Often Undiagnosed in Winter Deaths
- Combined Screening Approach Identifies Early Leprosy Cases
- Antibody Blood Test Identifies Active TB and Distinguishes Latent Infection
- FDA Approval Expands Use of PD-L1 Companion Diagnostic in Esophageal and GEJ Carcinomas
- Study Identifies Inflammatory Pathway Driving Immunotherapy Resistance in Bladder Cancer
- Microfluidic Chip Detects Cancer Recurrence from Immune Response Signals
- Cancer Mutation ‘Fingerprints’ to Improve Prediction of Immunotherapy Response
Channels
Clinical Chemistry
view channel
Fluid Biomarker Improves Diagnosis and Monitoring of Primary CNS Lymphoma
Primary central nervous system lymphoma (PCNSL) is a rare malignancy of the brain, spinal cord, and eyes with delayed diagnosis and poor outcomes. Current fluid-based testing using interleukin measurements... Read more
New CA19-9 Cutoff Value Helps Identify High-Risk Pancreatic Cancer Patients
Pancreatic ductal adenocarcinoma (PDAC) is frequently diagnosed at an advanced stage and remains one of the most lethal solid tumors. Clinicians commonly use serum carbohydrate antigen 19-9 (CA19-9) to... Read moreHematology
view channel
Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
Iron deficiency in school-age children can affect brain development, learning, growth, and physical performance, yet early deficiency may be missed when screening focuses mainly on anemia.... Read more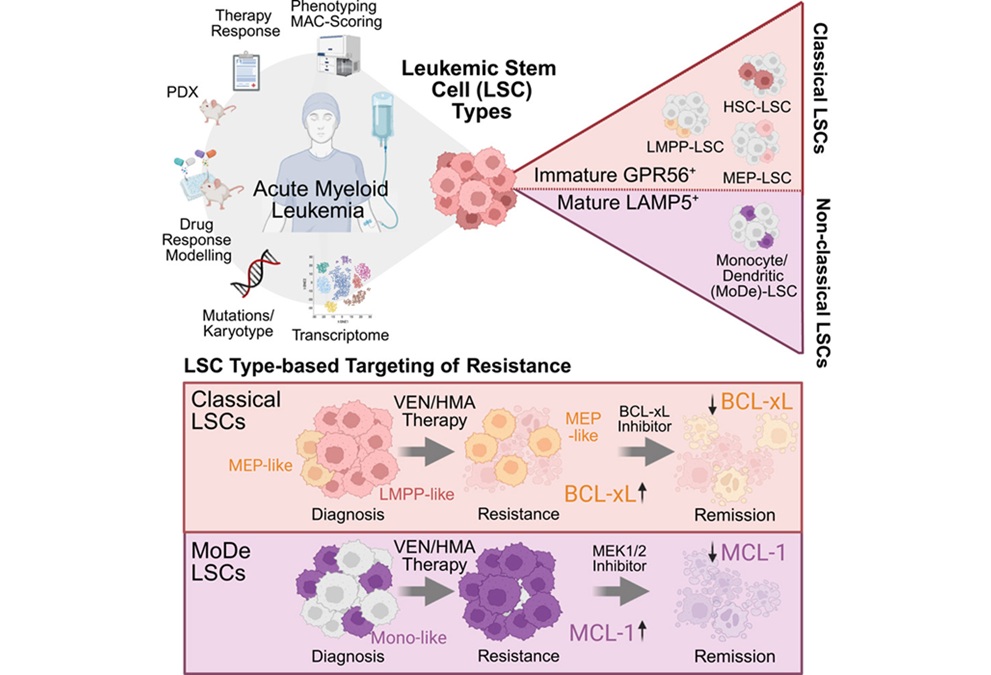
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read moreImmunology
view channel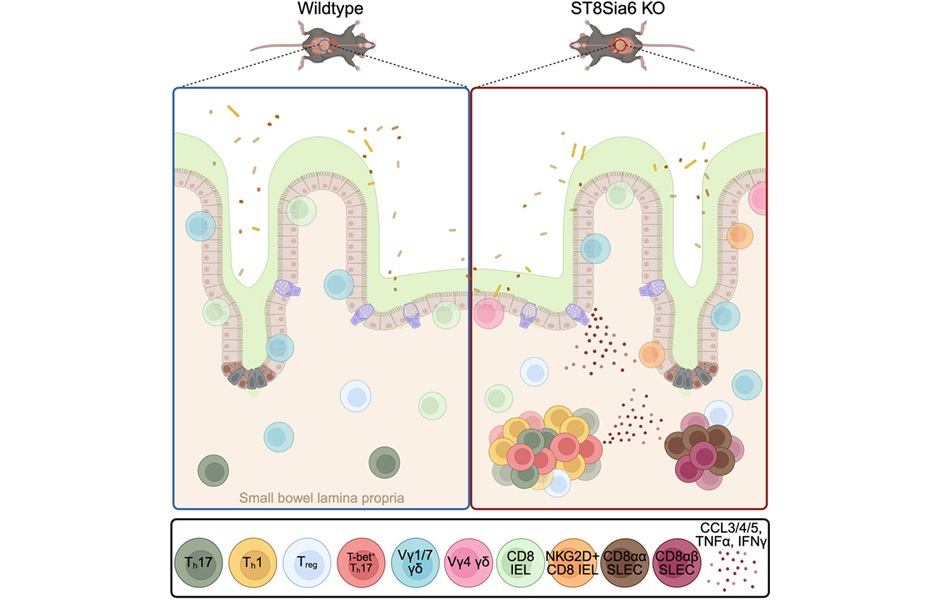
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read more
Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
Lung transplant recipients face some of the highest rates of acute cellular rejection, and routine surveillance often relies on repeated surgical biopsies. These procedures can cause complications such... Read moreMicrobiology
view channel
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read more
New AMR Assay Supports Rapid Infection Control Screening in Hospitals
As antimicrobial resistance spreads worldwide, healthcare-associated infections are placing a growing burden on hospitals, increasing the need for faster and broader diagnostic solutions.... Read morePathology
view channel
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read more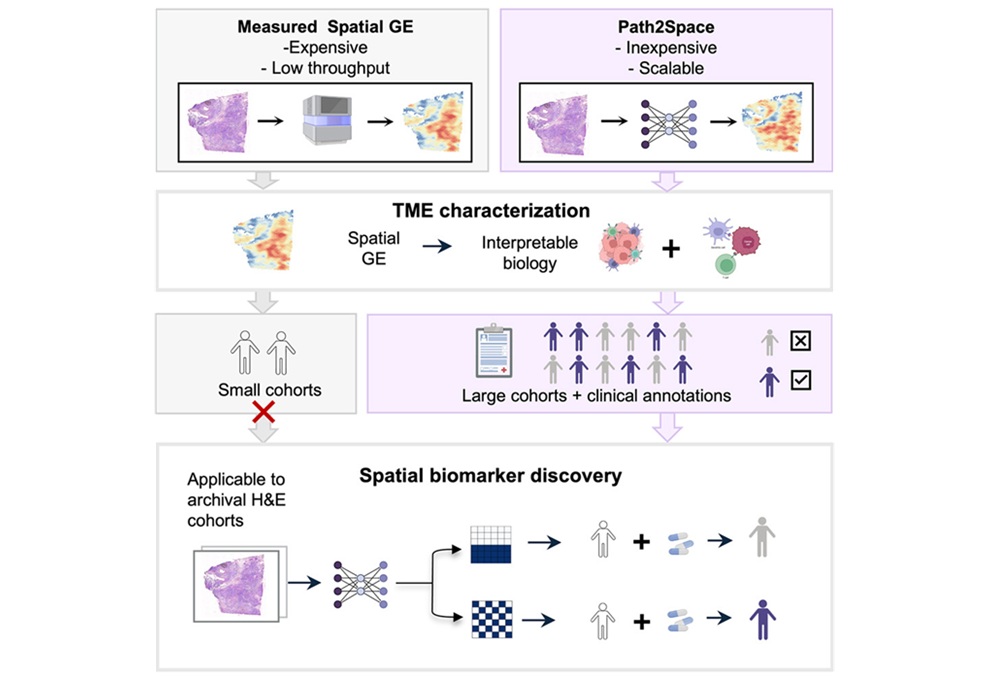
Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
Gene expression profiling can inform tumor biology and treatment selection, but spatial assays remain costly and time-consuming. Results can take weeks and cost thousands of dollars, limiting large-scale... Read moreTechnology
view channel
AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
Clinical laboratories and research groups face increasingly complex molecular workflows and expanding technical documentation spread across multiple systems. Fragmented digital tools can slow experiment... Read more
AI Tool Automates Validation of Laboratory Software Configuration Changes
Regulated laboratories face heavy documentation and requalification demands when software configurations change, slowing improvements and discouraging beneficial updates. A new capability now automates... Read moreIndustry
view channel
Natera to Present Data on MRD-Guided Cancer Care at ASCO 2026
Natera, Inc. (Austin, TX, USA), a company focused on cell-free DNA testing and precision medicine, announced an oncology data program for the 2026 American Society of Clinical Oncology (ASCO) Annual Meeting,... Read more