Protein Atlas Accelerates Personalized Medicine in Leukemia Patients
|
By LabMedica International staff writers Posted on 30 Apr 2019 |
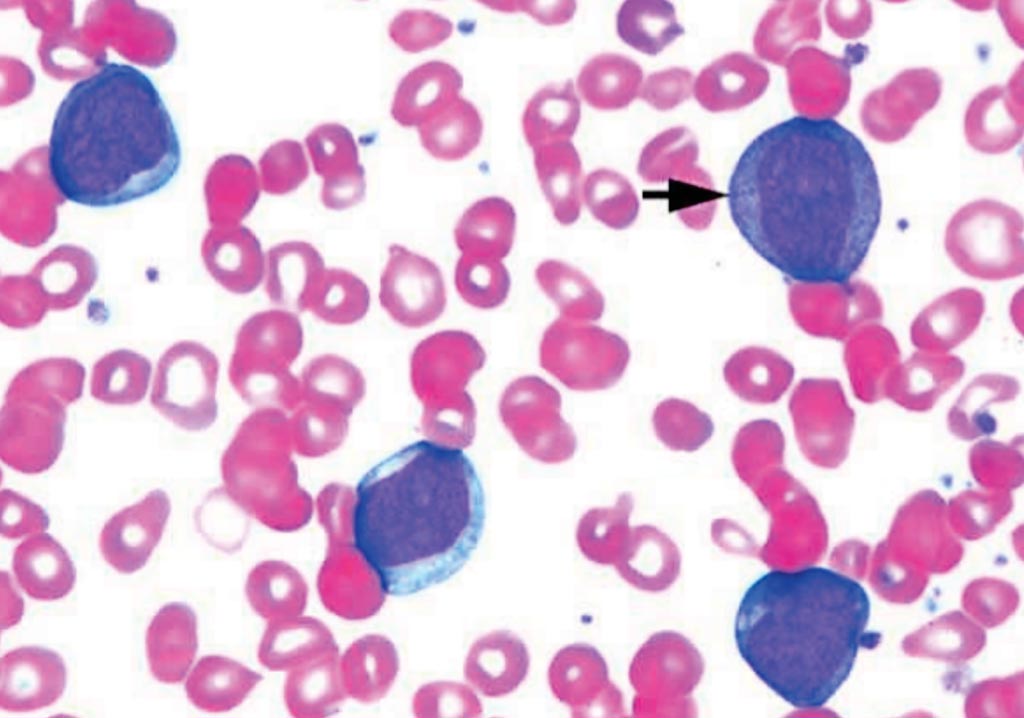
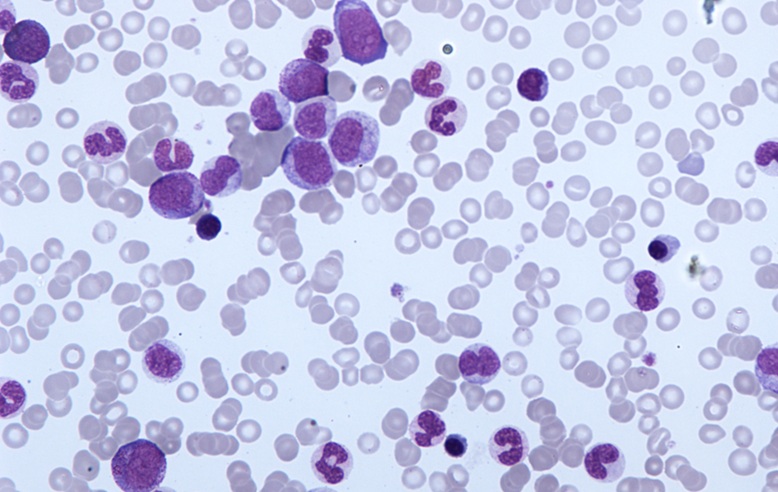
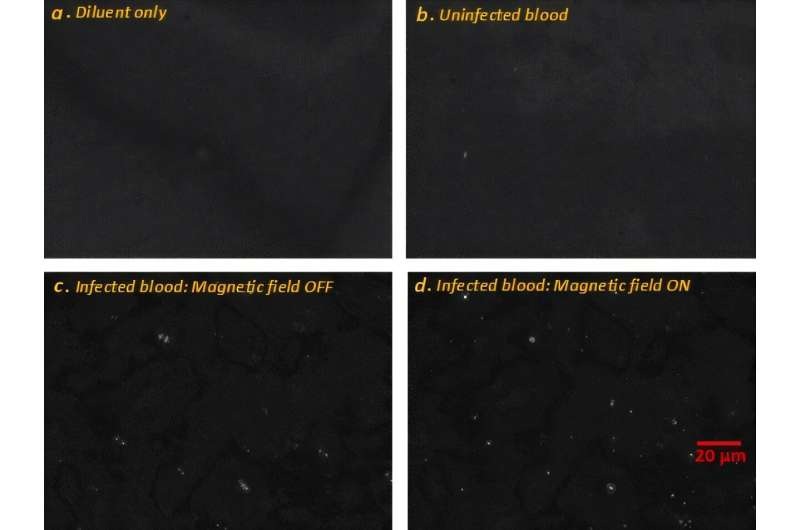
Image: Blood film of a patient with acute myelogenous leukemia defined by presence of more than 90% myeloblasts in blood and/or bone marrow (Photo courtesy of Pathpedia).
Acute myelogenous leukemia is associated with risk factors that are largely unknown and with a heterogeneous response to treatment. Only about one in four people diagnosed with acute myelogenous leukemia (AML) survive five years after the initial diagnosis.
To improve that survival rate, scientists have created an online atlas to identify and classify protein signatures present at AML diagnosis. The new protein classifications will help clinicians recommend better treatment and personalized medicine for patients suffering from this aggressive cancer, which occurs in the blood and bone marrow.
A team of scientists at the University of Texas at San Antonio (UTSA, San Antonio, TX, USA) and the University of Texas MD Anderson Cancer Center (Houston, TX, USA) examined the genetic, epigenetic and environmental diversity that occurs in cancerous cells due to AML. They analyzed proteomic screens of 205 patient biopsies and developed a new computational method called MetaGalaxy to categorize the protein signatures into 154 different patterns based on their cellular functions and pathways.
By approaching this challenge through the unique lens of developing a quantitative map for each leukemia patient from protein expression in their blood and bone marrow, rather than the standard lens of qualitative metrics and genetic risks alone, the collaborators will be able to more precisely categorize patients into risk groups and better predict their treatment outcomes. The team found 11 constellations of correlated functional patterns and 13 signatures that stratify the outcomes of patients. The scientists found limited overlap between proteomics data and both cytogenetics and genetic mutations. Moreover, leukemia cell lines show limited proteomic similarities with cells from patients with AML, suggesting that a deeper focus on patient-derived samples is needed to gain disease-relevant insights.
Amina Qutub, PhD, an associate professor and Biochemical Engineer and a senior study author said, “Acute myelogenous leukemia presents as a cancer so heterogeneous that it is often described as not one, but a collection of diseases. To decipher the clues found in proteins from blood and bone marrow of leukemia patients, we developed a new computer analysis, MetaGalaxy that identifies molecular hallmarks of leukemia. These hallmarks are analogous to the way constellations guide navigation of the stars: they provide a map to protein changes for leukemia.” The study was published on April 15, 2019, in the journal Nature Biomedical Engineering.
Related Links:
University of Texas at San Antonio
University of Texas MD Anderson Cancer Center
To improve that survival rate, scientists have created an online atlas to identify and classify protein signatures present at AML diagnosis. The new protein classifications will help clinicians recommend better treatment and personalized medicine for patients suffering from this aggressive cancer, which occurs in the blood and bone marrow.
A team of scientists at the University of Texas at San Antonio (UTSA, San Antonio, TX, USA) and the University of Texas MD Anderson Cancer Center (Houston, TX, USA) examined the genetic, epigenetic and environmental diversity that occurs in cancerous cells due to AML. They analyzed proteomic screens of 205 patient biopsies and developed a new computational method called MetaGalaxy to categorize the protein signatures into 154 different patterns based on their cellular functions and pathways.
By approaching this challenge through the unique lens of developing a quantitative map for each leukemia patient from protein expression in their blood and bone marrow, rather than the standard lens of qualitative metrics and genetic risks alone, the collaborators will be able to more precisely categorize patients into risk groups and better predict their treatment outcomes. The team found 11 constellations of correlated functional patterns and 13 signatures that stratify the outcomes of patients. The scientists found limited overlap between proteomics data and both cytogenetics and genetic mutations. Moreover, leukemia cell lines show limited proteomic similarities with cells from patients with AML, suggesting that a deeper focus on patient-derived samples is needed to gain disease-relevant insights.
Amina Qutub, PhD, an associate professor and Biochemical Engineer and a senior study author said, “Acute myelogenous leukemia presents as a cancer so heterogeneous that it is often described as not one, but a collection of diseases. To decipher the clues found in proteins from blood and bone marrow of leukemia patients, we developed a new computer analysis, MetaGalaxy that identifies molecular hallmarks of leukemia. These hallmarks are analogous to the way constellations guide navigation of the stars: they provide a map to protein changes for leukemia.” The study was published on April 15, 2019, in the journal Nature Biomedical Engineering.
Related Links:
University of Texas at San Antonio
University of Texas MD Anderson Cancer Center
Latest Molecular Diagnostics News
- Genetic Signature Predicts Myeloid Leukemia Risk in Down Syndrome
- Gene Expression Model Guides Neoadjuvant Therapy Selection in Breast Cancer
- AI Blood Test Enhances Monitoring of Liver Cirrhosis Progression
- Cancer-Related Mutations in Immune Cells Linked to Alzheimer’s
- Composite Blood Biomarkers Enable Early Detection of Common Cancers
- Machine Learning Model Uses DNA Methylation to Predict Tumor Origin in Cancers of Unknown Primary
- Blood Test Enables Early Detection and Classification of Glioma
- Multi-Biomarker Blood Test Detects Early-Stage Cancers Across Types
- New Sample-to-Answer PCR System Supports High-Throughput Infectious Disease Testing
- Framework Guides Targeted Immunotherapy Selection in Liver Cancer
- Collaboration Brings Rapid At-Home STI Testing with Virtual Follow-Up
- Blood-Based Epigenetic Signals Enable Osteosarcoma Disease Monitoring
- Host–Virus Genetic Interactions Drive Nasopharyngeal Cancer Risk
- AI-Enabled Biochip Detects microRNA Biomarkers in Minutes
- Blood Test Detects Early Pancreatic Cancer in High-Risk Patients
- Long-Read RNA Sequencing Platform Improves Rare Disease Diagnosis
Channels
Clinical Chemistry
view channel
Proteomic Data Underscore Need for Age-Specific Pediatric Reference Ranges
Serum proteins underpin many routine tests used to detect inflammation, hormonal imbalance, cardiovascular disease, and metabolic disorders. Yet pediatric interpretation often relies on adult reference... Read more
Routine Blood Count Ratio Linked to Future Alzheimer’s and Dementia Risk
Alzheimer’s disease and related dementias develop over years, making it difficult to identify at-risk patients before symptoms appear. Clinicians therefore need widely available laboratory markers that... Read more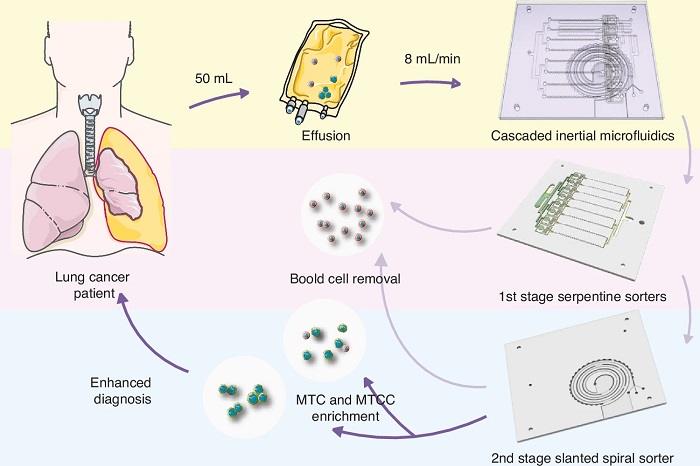
Label-Free Microfluidic Device Enriches Tumor Cells and Clusters from Pleural Effusions
Diagnosing malignancy from pleural effusion remains challenging because tumor cells are rare and clusters are easily disrupted during processing. Conventional cytology can miss malignant tumor cells and... Read moreHematology
view channel
Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
CareDx (Brisbane, CA, USA) has introduced AlloSeq Nano, a nanopore‑based HLA (human leukocyte antigen) and ABO genotyping solution unveiled at the European Federation for Immunogenetics (EFI) Conference 2026.... Read more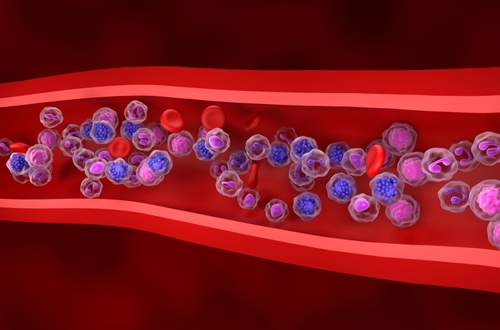
Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
Diffuse large B-cell lymphoma (DLBCL) is the most common form of non-Hodgkin lymphoma and often presents with aggressive clinical behavior. Although many patients respond to standard chemotherapy with... Read moreImmunology
view channel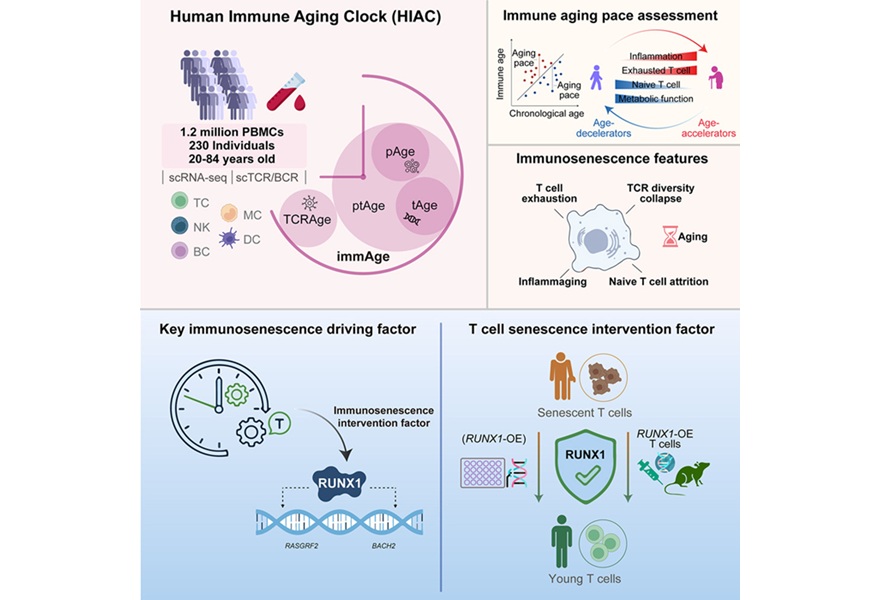
Immune Aging Clock Quantifies Immunosenescence and Identifies Therapeutic Target
Immune aging undermines host defense and contributes to multiple age-related diseases, yet its heterogeneity complicates measurement and intervention. Clinical laboratories increasingly seek objective... Read more
Study Finds Influenza Often Undiagnosed in Winter Deaths
Seasonal influenza drives substantial excess mortality, yet its contribution is often obscured when infections go undiagnosed near the time of death. Many deaths occur outside hospitals or in older adults... Read moreMicrobiology
view channel
Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
Tuberculosis remains a major global health challenge and continues to drive significant morbidity and mortality. The World Health Organization’s 2024 global report cites it as the leading cause of death... Read more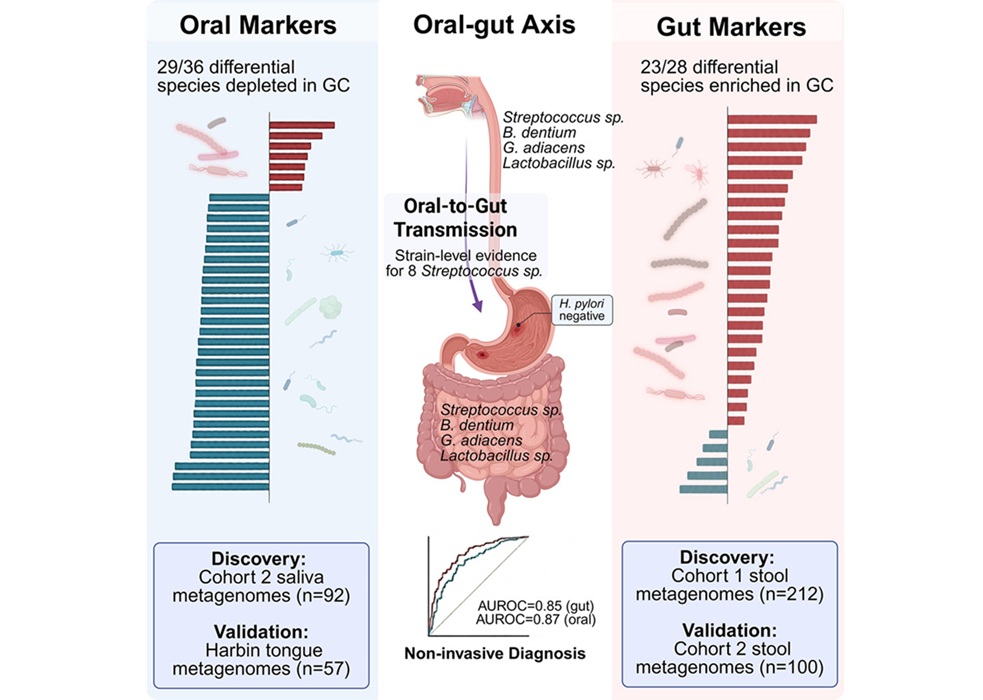
Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
Early detection of gastric cancer could be advanced by scalable screening strategies using minimally invasive sampling. Saliva collection is noninvasive and cost-effective, supporting wider adoption... Read morePathology
view channel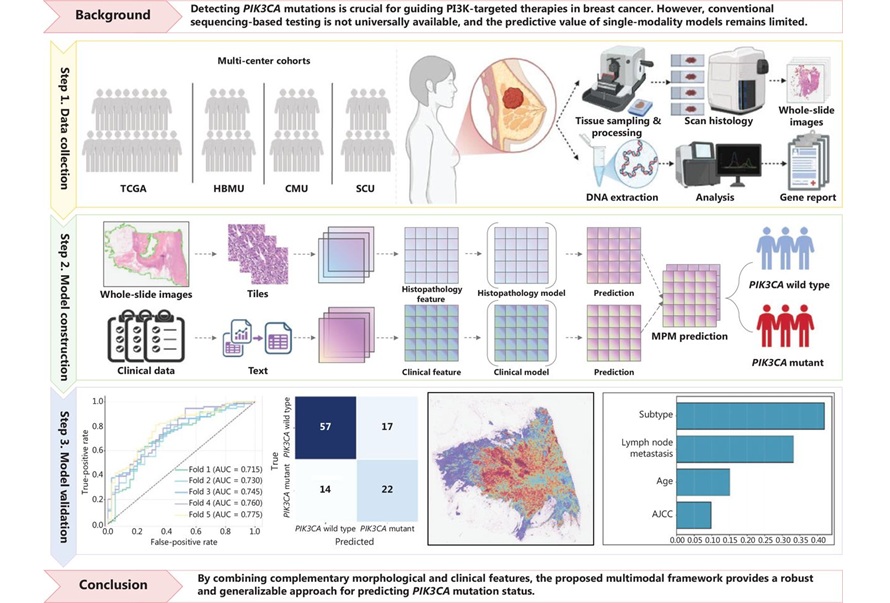
Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
PIK3CA mutations are key biomarkers for selecting phosphoinositide 3-kinase (PI3K)–targeted therapies in breast cancer, yet access to molecular testing can be inconsistent and costly. Conventional polymerase... Read more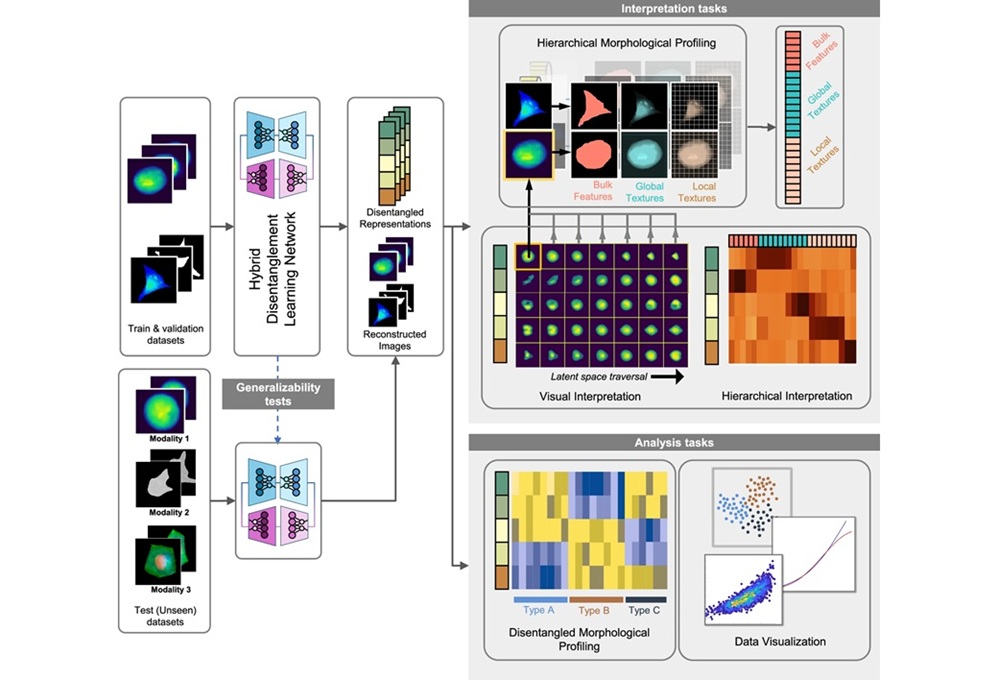
Interpretable AI Reveals Hidden Cellular Features from Microscopy Images
Microscopy images contain rich clues about cell health, but many disease-relevant morphological differences are too subtle to see and difficult to quantify consistently. Artificial intelligence (AI) has... Read moreTechnology
view channel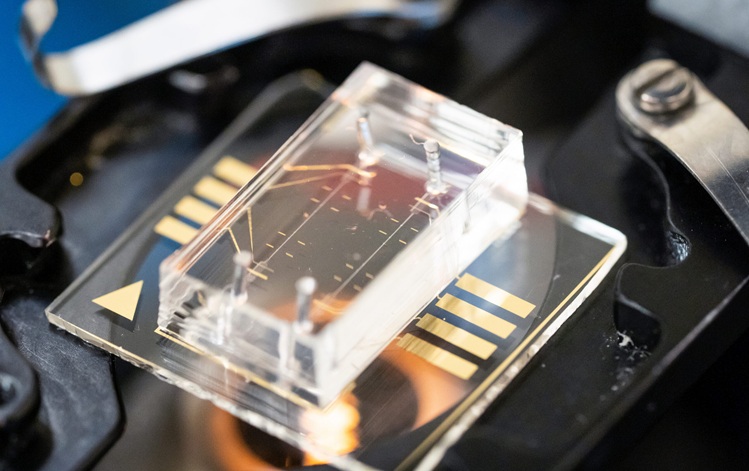
Microfluidic Single-Cell Assay Predicts Breast Cancer Risk
Risk stratification for breast cancer remains imprecise, as population-based models and breast density can over- or underestimate individual risk, potentially leading to over- or under-screening.... Read more
AI Tool Predicts Non-Response to Targeted Therapy in Colorectal Cancer
Advanced bowel cancer remains difficult to treat, and many patients receive targeted therapies that do not help them but still cause harm. Clinicians need reliable ways to identify likely responders before... Read moreIndustry
view channel
Collaboration Expands Access to Rapid Metagenomic Diagnostics for Complex Infections
Hospitals are seeing rising rates of complicated and healthcare-associated infections, especially in immunocompromised patients, intensifying the need for rapid, comprehensive pathogen detection.... Read more