New Genetic Risk Factors Identified for Peanut Allergy
|
By LabMedica International staff writers Posted on 25 Oct 2017 |
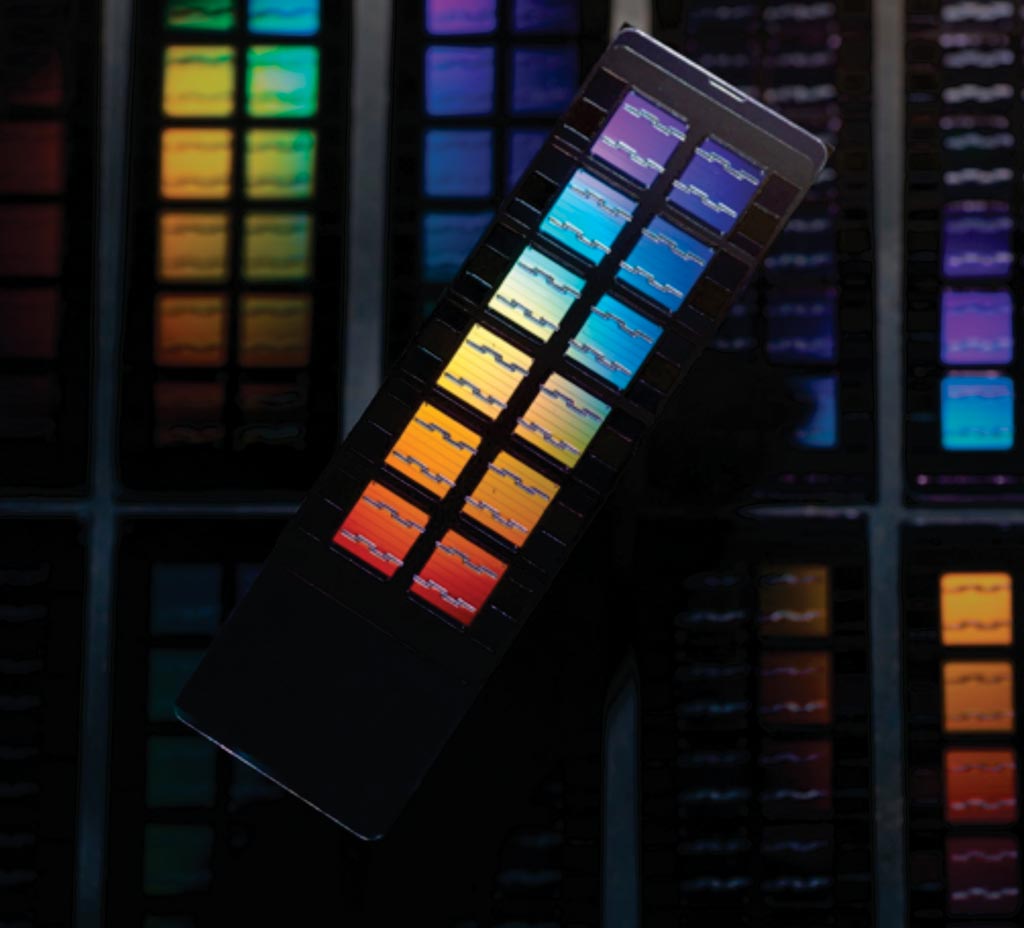

Image: Whole genome genotyping arrays are an important tool for discovering variants that contribute to human disease (Photo courtesy of Megan Smolenyak, MBA).
Peanut allergy develops in early life and is rarely outgrown. Roughly 1% of Canadian adults and between 2% and 3% of Canadian children are affected, and the symptoms can be severe and even life threatening.
A new gene associated with peanut allergy has been revealed, offering further evidence that genes play a role in the development of food allergies and opening the door to future studies, improved diagnostics and new treatment options.
An international team of scientists collaborating with those at the University of British Columbia (Vancouver, BC, Canada) scanned more than 7.5 million genetic locations in the DNA of 850 people with peanut allergy and nearly 1,000 people without it, through a genome-wide association study (GWAS), to search for markers that might be linked to food allergy. They recruited the peanut allergy participants from the Canadian Peanut Allergy Registry. The team also conducted a fresh analysis of results pooled from six other genetic studies of populations in North America, Australia, Germany, and the Netherlands. Genotyping of 1,974 individuals (987 cases, 987 controls) was conducted on the Illumina Omni 2.5M+Exome 8v1.1 chip.
The scientists reported that their study is the first to associate the EMSY, BRCA2 Interacting Transcriptional Repressor (EMSY) locus with food allergy, and these findings suggest that the gene plays an important role in the development of not just food allergy but also general allergic predisposition. The gene, called c11orf30/EMSY (EMSY), is already known to play a role in other allergy-related conditions, such as eczema, asthma, and allergic rhinitis. The team also found evidence that five other genetic locations might be involved.
Denise Daley, PhD, an associate professor and senior author of the study, said, “Food allergy is the result of both genetic and environmental factors, but there are surprisingly few data regarding the genetic basis of this condition. The discovery of this genetic link gives us a fuller picture of the causes of food allergies, and this could eventually help doctors identify children at risk.” The study was published on November 10, 2017, in the Journal of Allergy and Clinical Immunology.
Related Links:
University of British Columbia
A new gene associated with peanut allergy has been revealed, offering further evidence that genes play a role in the development of food allergies and opening the door to future studies, improved diagnostics and new treatment options.
An international team of scientists collaborating with those at the University of British Columbia (Vancouver, BC, Canada) scanned more than 7.5 million genetic locations in the DNA of 850 people with peanut allergy and nearly 1,000 people without it, through a genome-wide association study (GWAS), to search for markers that might be linked to food allergy. They recruited the peanut allergy participants from the Canadian Peanut Allergy Registry. The team also conducted a fresh analysis of results pooled from six other genetic studies of populations in North America, Australia, Germany, and the Netherlands. Genotyping of 1,974 individuals (987 cases, 987 controls) was conducted on the Illumina Omni 2.5M+Exome 8v1.1 chip.
The scientists reported that their study is the first to associate the EMSY, BRCA2 Interacting Transcriptional Repressor (EMSY) locus with food allergy, and these findings suggest that the gene plays an important role in the development of not just food allergy but also general allergic predisposition. The gene, called c11orf30/EMSY (EMSY), is already known to play a role in other allergy-related conditions, such as eczema, asthma, and allergic rhinitis. The team also found evidence that five other genetic locations might be involved.
Denise Daley, PhD, an associate professor and senior author of the study, said, “Food allergy is the result of both genetic and environmental factors, but there are surprisingly few data regarding the genetic basis of this condition. The discovery of this genetic link gives us a fuller picture of the causes of food allergies, and this could eventually help doctors identify children at risk.” The study was published on November 10, 2017, in the Journal of Allergy and Clinical Immunology.
Related Links:
University of British Columbia
Latest Molecular Diagnostics News
- Blood Test Predicts Immunotherapy Response in Head and Neck Cancer
- New PCR Assay Supports Bundibugyo Ebola Outbreak Surveillance
- Blood-Based RNA Test May Predict Chemotherapy Sensitivity in Lung Cancer
- Plasma Protein Signature Predicts Lung Cancer Risk Up to Five Years Ahead
- Circulating Tumor DNA Testing Guides Chemotherapy, Reduces Relapse in Colon Cancer
- Researchers Uncover Distinct Chromosome Signature in Aggresive ALT Cancers
- Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
- Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
- Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
- Tumor Mutation Marker Helps Refine Lung Cancer Prognosis and Guide Therapy Selection
- Multi-Cancer Test Boosts Detection When Added to Standard Screening
- Blood-Based MRD Monitoring Supports Relapse Prevention in Leukemia
- Genomic Test Predicts Chemotherapy Benefit in Metastatic Prostate Cancer
- Blood Protein Markers Flag Multiple Sclerosis Risk Years Before Diagnosis
- Digital PCR Assays Support Surveillance of Bundibugyo Ebolavirus Outbreak
- Updated Guidance Prioritizes Stool-Based Colorectal Cancer Screening Tests
Channels
Clinical Chemistry
view channel
Urinary Biomarker Assay Predicts Kidney Disease Progression Beyond Standard Measures
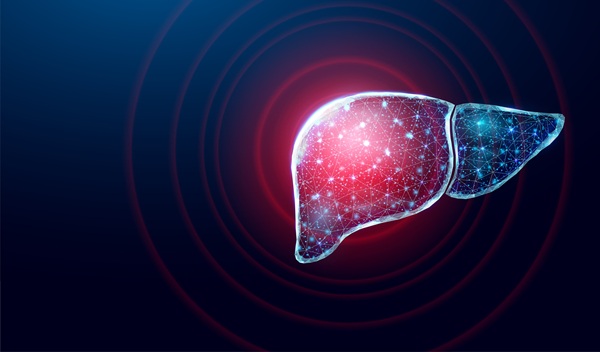
Many patients with type 2 diabetes and chronic kidney disease continue to experience progressive renal decline, yet conventional markers such as albuminuria and estimated glomerular filtration rate (eGFR)... Read more
Saliva-Based Test Detects Biochemical Signs of Sleep Loss
Acute sleep loss impairs cognition and motor skills, raising safety risks that resemble alcohol intoxication. Clinicians currently lack an objective biochemical test to determine when someone is dangerously... Read more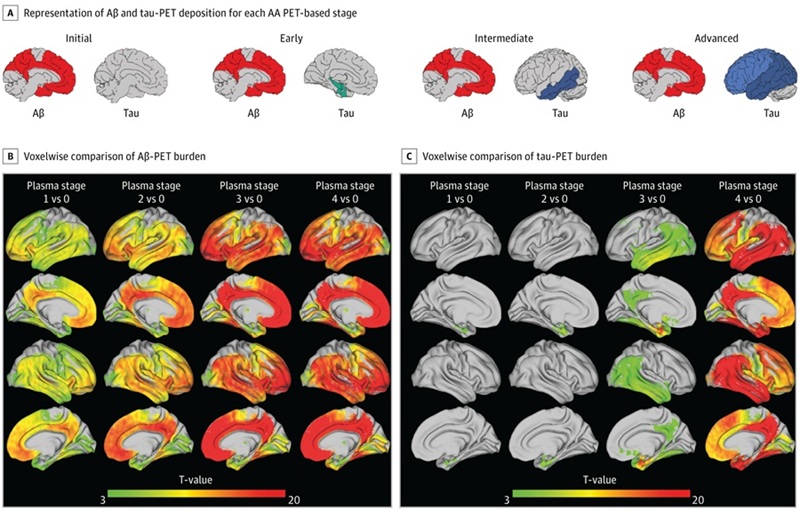
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreMolecular Diagnostics
view channel
Blood-Based RNA Test May Predict Chemotherapy Sensitivity in Lung Cancer
Lung cancer care increasingly relies on biomarker-guided patient stratification, but tissue biopsy can be impractical and treatment selection remains difficult for many patients. Blood-based assays that... Read more
Blood Test Predicts Immunotherapy Response in Head and Neck Cancer
Head and neck squamous cell carcinoma affects hundreds of thousands of people worldwide each year, yet response rates to immunotherapy remain low. Clinicians lack reliable, minimally invasive tools to... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more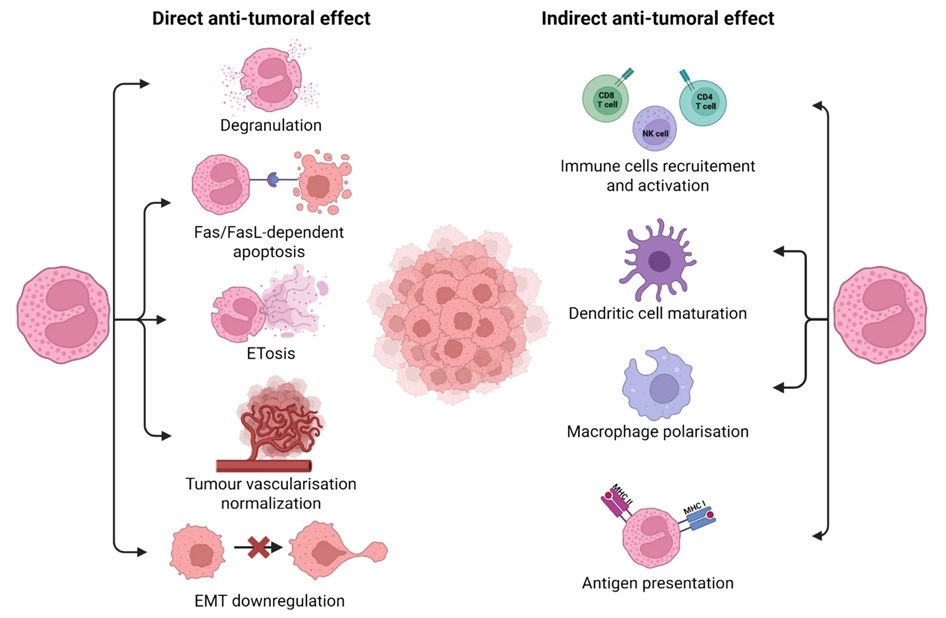
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreMicrobiology
view channel
New Culture Medium Speeds C. difficile Resistance Detection and Reduces Costs
Clostridioides difficile infections remain a persistent threat in hospitals and communities, affecting about 500,000 people in the United States each year. Severe cases can be fatal within 30 days of diagnosis,... Read more
Automated Blood Culture System Speeds Detection of Bloodstream Infections
Bloodstream infections and sepsis require rapid laboratory detection to guide targeted antimicrobial therapy and reduce mortality. Conventional blood culture workflows can delay actionable results by critical... Read morePathology
view channel
AI Pathology Tool Predicts Meningioma Recurrence from Routine Slides
Meningiomas are the most common primary brain tumors in adults, yet their course ranges from indolent to highly recurrent disease. Estimating an individual patient’s recurrence risk often requires advanced... Read more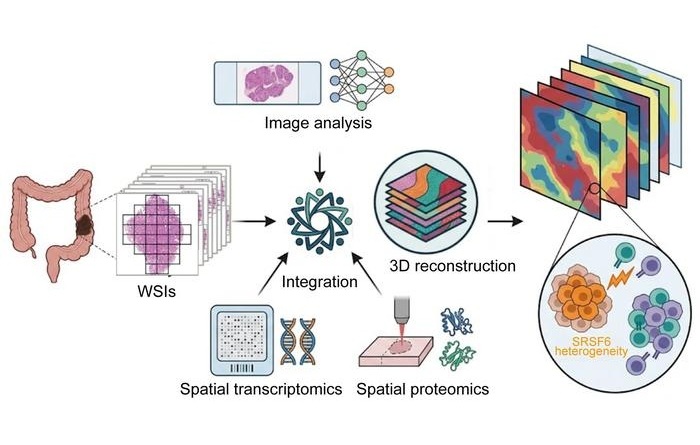
3D Spatial Multi-Omics Maps Intra-Tumor Diversity in Colorectal Cancer
Colorectal cancer remains a leading cause of cancer death, and clinical decision-making is complicated by marked intra-tumor heterogeneity. Conventional bulk sequencing averages molecular signals across... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel
Genetic Testing Program Expands Detection of Alpha-1 Antitrypsin Deficiency
Alpha-1 Antitrypsin Deficiency (AATD) is a progressive genetic condition, the leading known genetic risk factor for chronic obstructive pulmonary disease (COPD), and a cause of liver disease in both children... Read more