New Gene Mutation Associated with Fanconi Anemia
|
By LabMedica International staff writers Posted on 25 Jul 2017 |
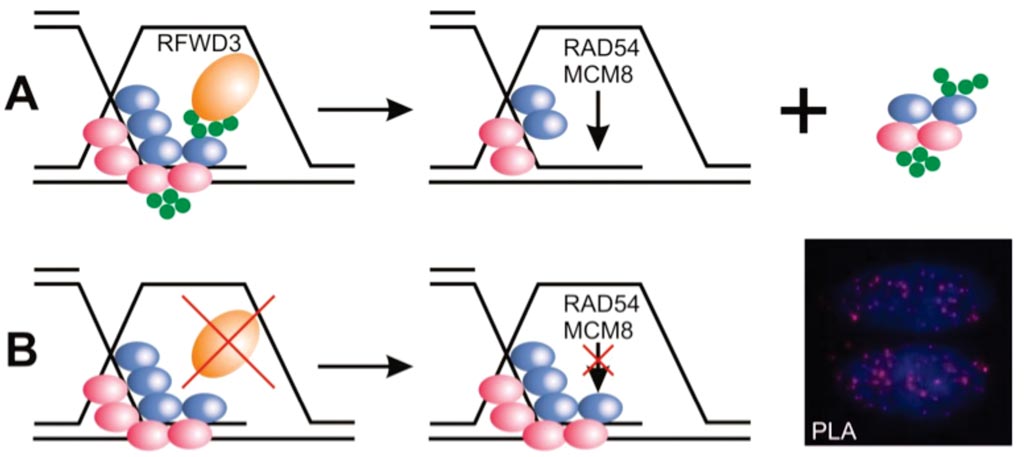
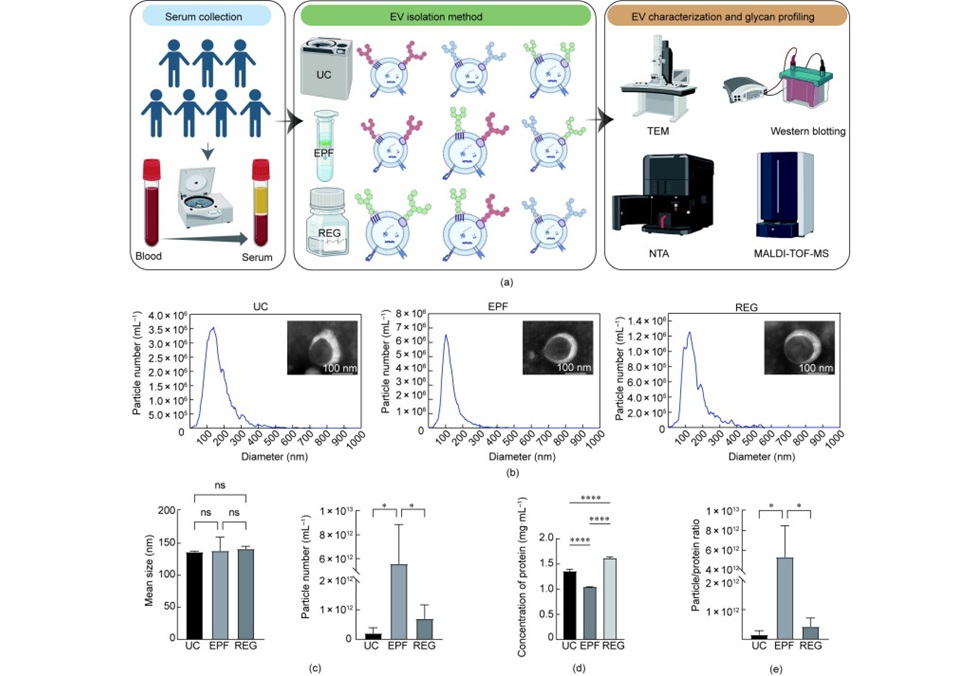
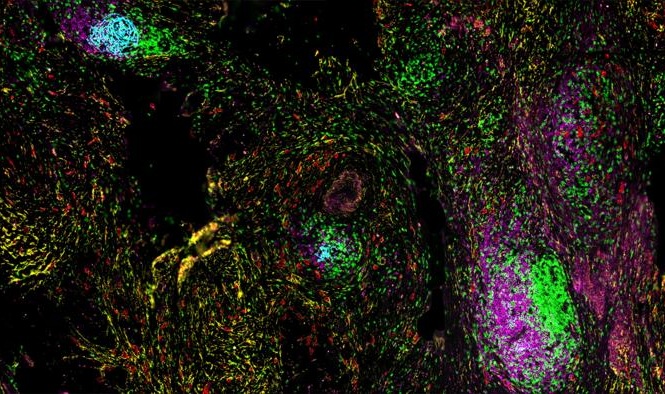
Image: The enzyme E3 ubiquitin ligase (RFWD3) helps target other proteins on single-stranded DNA for degradation. B: Cells lacking RFWD3 show DNA repair defects (Photo courtesy of Julius-Maximilians-Universität).
Fanconi anemia is a rare genetic disease characterized by bone marrow failure heralded by low platelet counts and unusually large red blood cells. Mutations in over 20 genes have been identified as causative for Fanconi anemia, which encode proteins commonly involved in DNA repair mechanisms.
Safeguarding of the genome is essential for the suppression of oncogenesis and for stem cell maintenance in many organisms, and is warranted by DNA repair mechanisms. If they fail on a genetic basis, DNA repair disorders occur and Fanconi anemia (FA) is such a rare human genomic instability disease.
Scientists at the Julius-Maximilians-Universität (Würzburg, Germany) reports classical Fanconi anemia symptoms in a 12-year-old individual without mutations in any of the known Fanconi anemia genes. The patient presented with congenital abnormalities characteristic of FA and had a typical FA phenotype and compound heterozygous mutations in the E3 ubiquitin ligase (RFWD3).
The team used various techniques in the study including DNA sample preparation, polymerase chain reaction (PCR), Sanger sequencing, and whole exome sequencing, on a HighSeq2000 instrument. Other methods used were generation of stable lentiviral producer lines and transduction of adherent cells, DT40 gene targeting, cell cycle, cell survival, and chromosomal studies, where cell numbers were determined using a Nucleo Counter NC-250. Immunofluorescence studies were analyzed on a BIOREVO BZ-9000 microscope.
Sequencing of this individual's genome detected mutations in both alleles of the gene RFWD3, which encodes an enzyme that helps target other proteins on single-stranded DNA for degradation. This process is impaired in patient's cells, which rendered them more sensitive to chromosome breakage and DNA damage, compared to cells from healthy individuals. Other cells either lacking RFWD3 or genetically engineered with the patient's missense mutation showed similar DNA repair defects, which were rescued by expression of wild-type RFWD3. Together, these findings support the identification of RFWD3 as a Fanconi anemia gene. The study was published on July 10, 2017, in the Journal of Clinical Investigation.
Related Links:
Julius-Maximilians-Universität
Safeguarding of the genome is essential for the suppression of oncogenesis and for stem cell maintenance in many organisms, and is warranted by DNA repair mechanisms. If they fail on a genetic basis, DNA repair disorders occur and Fanconi anemia (FA) is such a rare human genomic instability disease.
Scientists at the Julius-Maximilians-Universität (Würzburg, Germany) reports classical Fanconi anemia symptoms in a 12-year-old individual without mutations in any of the known Fanconi anemia genes. The patient presented with congenital abnormalities characteristic of FA and had a typical FA phenotype and compound heterozygous mutations in the E3 ubiquitin ligase (RFWD3).
The team used various techniques in the study including DNA sample preparation, polymerase chain reaction (PCR), Sanger sequencing, and whole exome sequencing, on a HighSeq2000 instrument. Other methods used were generation of stable lentiviral producer lines and transduction of adherent cells, DT40 gene targeting, cell cycle, cell survival, and chromosomal studies, where cell numbers were determined using a Nucleo Counter NC-250. Immunofluorescence studies were analyzed on a BIOREVO BZ-9000 microscope.
Sequencing of this individual's genome detected mutations in both alleles of the gene RFWD3, which encodes an enzyme that helps target other proteins on single-stranded DNA for degradation. This process is impaired in patient's cells, which rendered them more sensitive to chromosome breakage and DNA damage, compared to cells from healthy individuals. Other cells either lacking RFWD3 or genetically engineered with the patient's missense mutation showed similar DNA repair defects, which were rescued by expression of wild-type RFWD3. Together, these findings support the identification of RFWD3 as a Fanconi anemia gene. The study was published on July 10, 2017, in the Journal of Clinical Investigation.
Related Links:
Julius-Maximilians-Universität
Latest Molecular Diagnostics News
- Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
- Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
- Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
- Tumor Mutation Marker Helps Refine Lung Cancer Prognosis and Guide Therapy Selection
- Multi-Cancer Test Boosts Detection When Added to Standard Screening
- Blood-Based MRD Monitoring Supports Relapse Prevention in Leukemia
- Genomic Test Predicts Chemotherapy Benefit in Metastatic Prostate Cancer
- Blood Protein Markers Flag Multiple Sclerosis Risk Years Before Diagnosis
- Digital PCR Assays Support Surveillance of Bundibugyo Ebolavirus Outbreak
- Updated Guidance Prioritizes Stool-Based Colorectal Cancer Screening Tests
- Blood-Based Proteomic Test May Predict Treatment Response in Non-Small Cell Lung Cancer
- Position Statements Outline Evidence Standards for Multi-Cancer Detection Tests
- Ultrasensitive MRD Blood Test Detects Early Breast Cancer Recurrence
- Gene Fusion Patterns May Flag High Risk Solitary Fibrous Tumors
- New RNA Origami Method Supports Faster Targeted Testing for Repeat Expansion Disorders
- FDA Approves Expanded Liquid Biopsy Panel for Advanced Cancer Profiling
Channels
Clinical Chemistry
view channel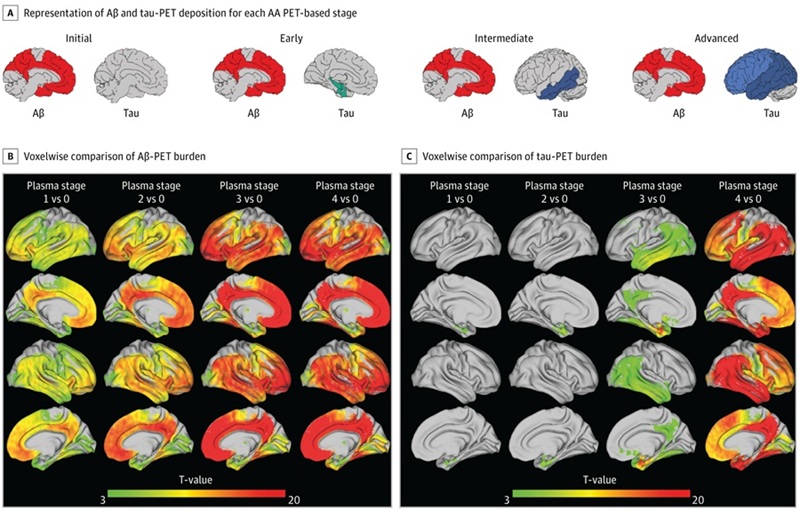
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more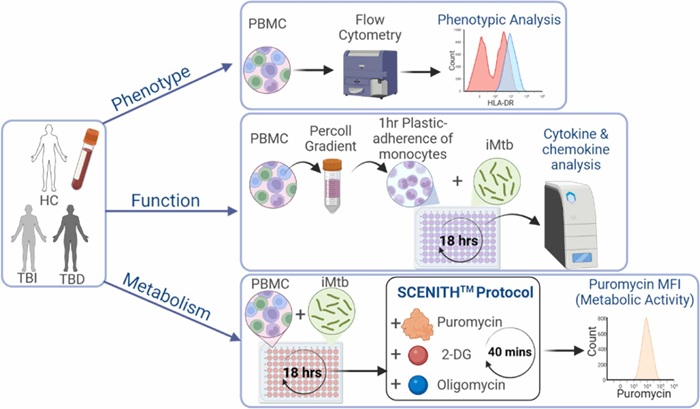
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read moreMicrobiology
view channel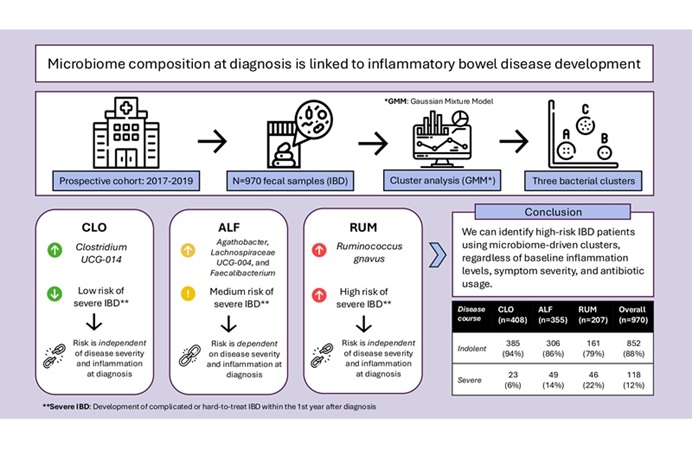
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more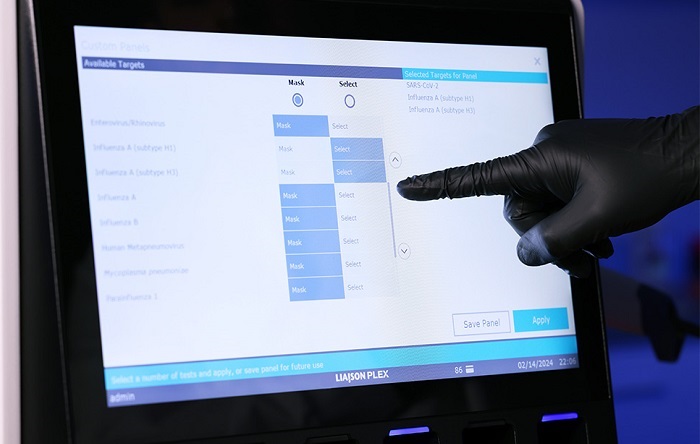
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel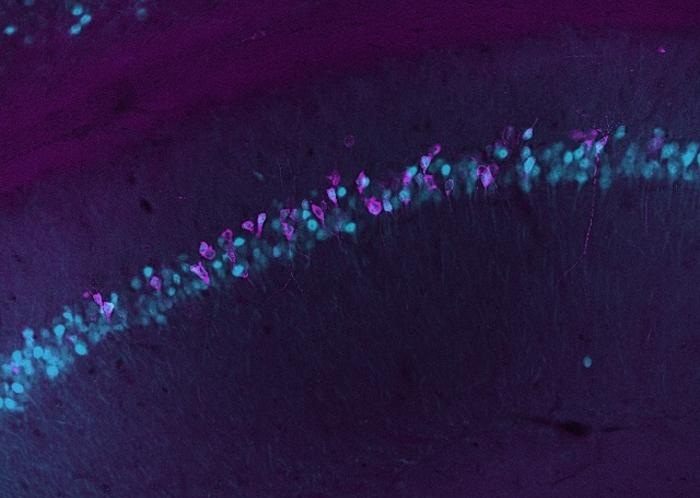
Blood-Based Method Tracks Gene Activity in the Living Brain
Real-time measurement of gene activity in the brain has been limited by assays requiring destructive tissue sampling. Tracking active genes could reveal how the body responds to environmental factors,... Read more
FDA Approval Expands Automated PD-L1 Testing Across Solid Tumors
Clinical laboratories play a central role in guiding immunotherapy by reporting programmed death ligand-1 (PD‑L1) status across multiple solid tumors. Many sites are standardizing this work on fully automated... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel