Biomarkers Could Give Cancer Patients Better Survival Estimates
|
By LabMedica International staff writers Posted on 21 Jun 2016 |
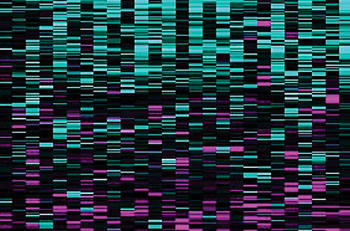
Image: A SURVIV analysis of breast cancer isoforms developed at UCLA. Blue lines are associated with longer survival times, and magenta lines with shorter survival times (Photo courtesy of Professor Yi Xing).
Cancer patients are often told by their doctors approximately how long they have to live, and how well they will respond to treatments, but there is a way to improve the accuracy of doctors' predictions.
A new method has been developed that could eventually lead to a way to do just that, using data about patients' genetic sequences to produce more reliable projections for survival time and how they might respond to possible treatments.
Scientists at the University of California-Los Angeles (UCLA, CA, USA) and their colleagues have developed a method that analyzes various gene isoforms using data from ribonucleic acid (RNA) molecules in cancer specimens. These isoforms are combinations of genetic sequences that can produce an enormous variety of RNAs and proteins from a single gene.
That process, called RNA sequencing, or RNA-seq, reveals the presence and quantity of RNA molecules in a biological sample. In the method developed, the scientists analyzed the ratios of slightly different genetic sequences within the isoforms, enabling them to detect important but subtle differences in the genetic sequences. In contrast, the conventional analysis aggregates all of the isoforms together, meaning that the technique misses important differences within the isoforms.
The scientists studied tissues from 2,684 people with cancer whose samples were part of the National Institutes of Health's Cancer Genome Atlas, and they spent more than two years developing the algorithm for SURVIV (for "survival analysis of mRNA isoform variation"). The team has identified some 200 isoforms that are associated with survival time for people with breast cancer; some predict longer survival times, others are linked to shorter times. Armed with that knowledge, the scientists might eventually be able to target the isoforms associated with shorter survival times in order to suppress them and fight disease. They evaluated the performance of survival predictors using a metric called C-index and found that across the six different types of cancer they analyzed, their isoform-based predictions performed consistently better than the conventional gene-based predictions.
Yi Xing, PhD, an assistant professor and senior author of the study, said, “Our finding suggests that isoform ratios provide a more robust molecular signature of cancer patients in large-scale RNA-seq datasets. In cancer, sometimes a single gene produces two isoforms, one of which promotes metastasis and one of which represses metastasis.” The study was published on June 9, 2016, in the journal Nature Communications.
Related Links:
University of California-Los Angeles
A new method has been developed that could eventually lead to a way to do just that, using data about patients' genetic sequences to produce more reliable projections for survival time and how they might respond to possible treatments.
Scientists at the University of California-Los Angeles (UCLA, CA, USA) and their colleagues have developed a method that analyzes various gene isoforms using data from ribonucleic acid (RNA) molecules in cancer specimens. These isoforms are combinations of genetic sequences that can produce an enormous variety of RNAs and proteins from a single gene.
That process, called RNA sequencing, or RNA-seq, reveals the presence and quantity of RNA molecules in a biological sample. In the method developed, the scientists analyzed the ratios of slightly different genetic sequences within the isoforms, enabling them to detect important but subtle differences in the genetic sequences. In contrast, the conventional analysis aggregates all of the isoforms together, meaning that the technique misses important differences within the isoforms.
The scientists studied tissues from 2,684 people with cancer whose samples were part of the National Institutes of Health's Cancer Genome Atlas, and they spent more than two years developing the algorithm for SURVIV (for "survival analysis of mRNA isoform variation"). The team has identified some 200 isoforms that are associated with survival time for people with breast cancer; some predict longer survival times, others are linked to shorter times. Armed with that knowledge, the scientists might eventually be able to target the isoforms associated with shorter survival times in order to suppress them and fight disease. They evaluated the performance of survival predictors using a metric called C-index and found that across the six different types of cancer they analyzed, their isoform-based predictions performed consistently better than the conventional gene-based predictions.
Yi Xing, PhD, an assistant professor and senior author of the study, said, “Our finding suggests that isoform ratios provide a more robust molecular signature of cancer patients in large-scale RNA-seq datasets. In cancer, sometimes a single gene produces two isoforms, one of which promotes metastasis and one of which represses metastasis.” The study was published on June 9, 2016, in the journal Nature Communications.
Related Links:
University of California-Los Angeles
Latest Pathology News
- AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
- AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
- Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
- AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
- FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
- New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
- AI System Analyzes Routine Pathology Slides to Predict Cancer Outcomes
- New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease
- Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
- Interpretable AI Reveals Hidden Cellular Features from Microscopy Images
- Tumor Immune Structure Predicts Response to Immunotherapy in Melanoma
- Plug-and-Play AI Pathology System Classifies Multiple Cancers from Few Slides
- AI-Based Assays Support Risk Stratification in Prostate and Breast Cancer
- AI Pathology Model Predicts Immunotherapy Response in Lung Cancer
- Study Reveals Moleclar Mechanism Driving Aggressive Skin Cancer
- AI Precision Tests Deliver Cancer Risk Insights from Routine H&E Slides
Channels
Clinical Chemistry
view channel
Urine-Based Test Shows Promise for Autism Screening in Children
Autism spectrum disorder (ASD) is commonly diagnosed through behavioral assessments, which can involve long waits that delay intervention. Earlier identification is linked to better developmental outcomes,... Read more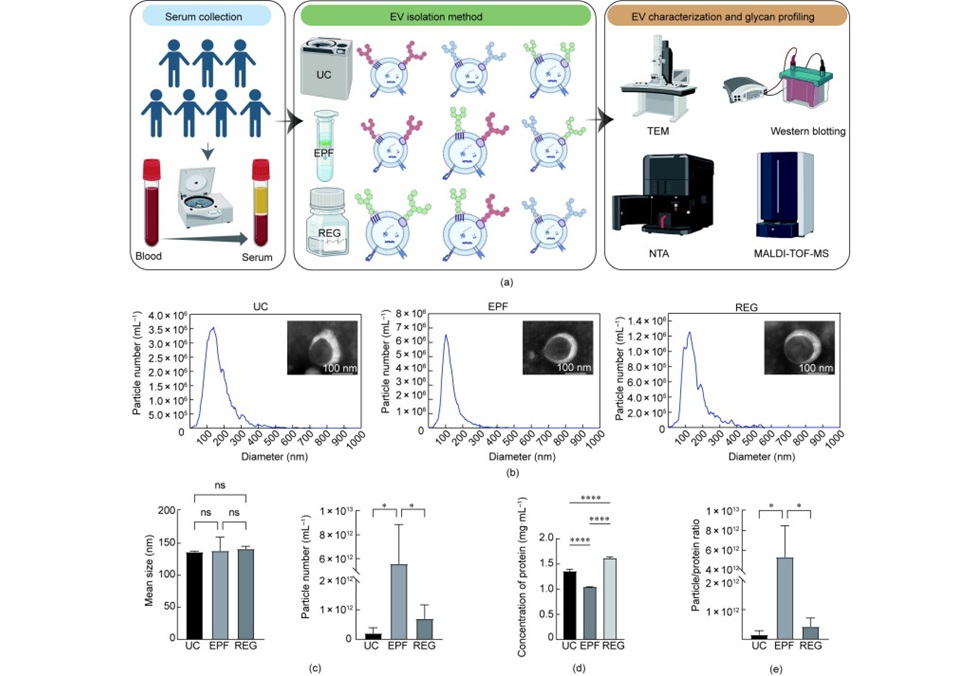
Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
Childhood epilepsy remains a major neurological disorder with unmet needs for accurate, non-invasive biomarkers, as conventional tests such as electroencephalography and neuroimaging can have limited sensitivity... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more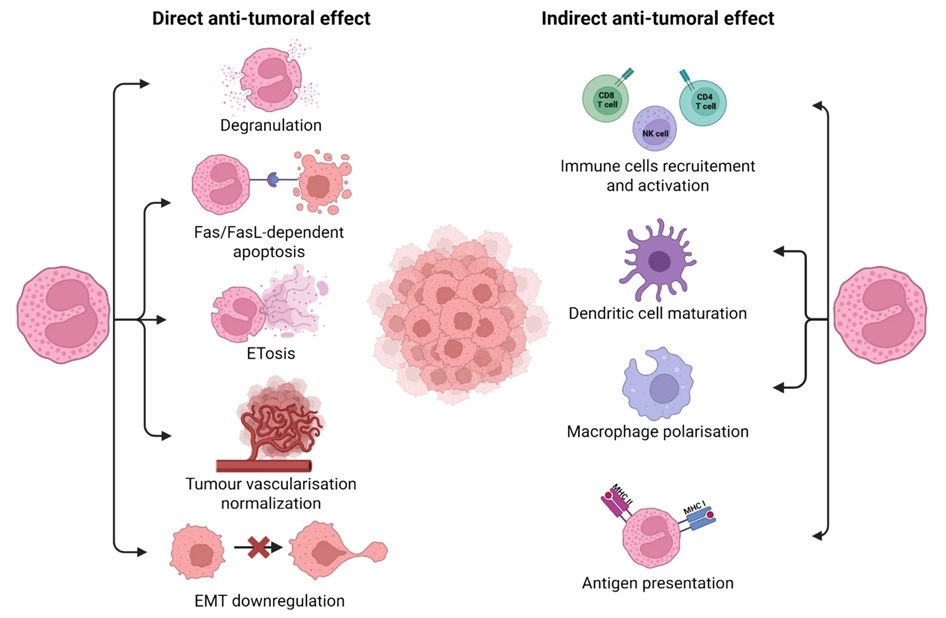
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel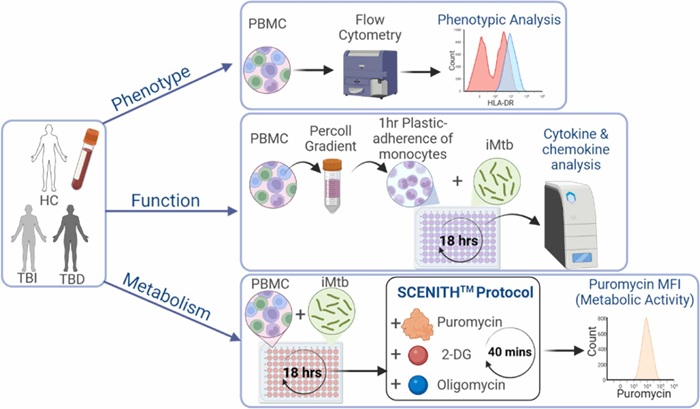
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read more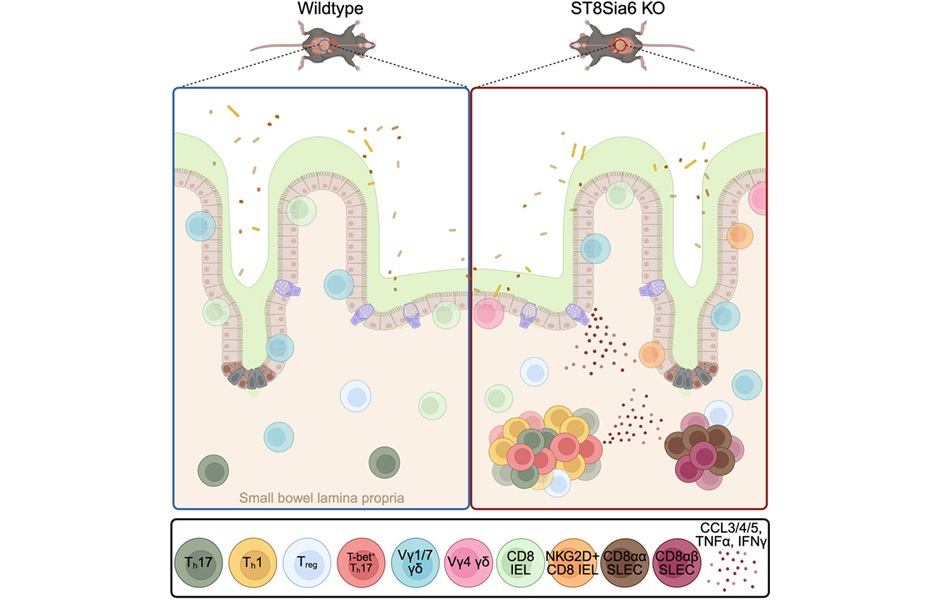
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read moreMicrobiology
view channel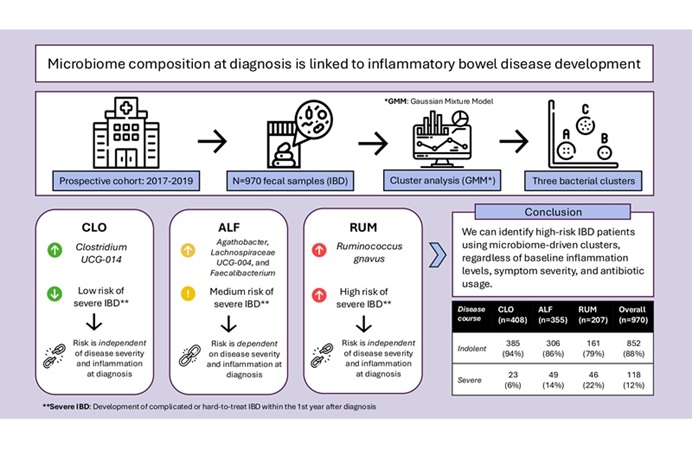
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel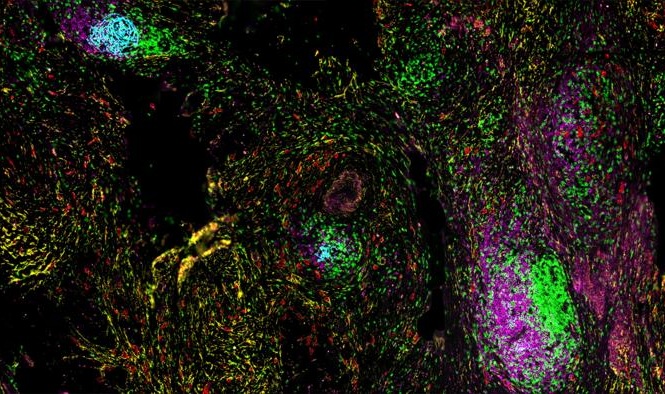
AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
Tertiary lymphoid structures are emerging as important indicators of antitumor immunity, but their heterogeneity and spatial context within tumors remain difficult to capture through routine diagnostics.... Read more
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read moreTechnology
view channel
Mailed Screening Kits Help Reduce Colorectal Cancer Screening Gaps
Colorectal cancer screening is a longstanding preventive priority, yet participation and follow-up remain uneven across patient groups. Safety‑net primary care settings often face barriers that limit screening... Read more
Algorithm Panel Aids Liver Fibrosis Assessment and Liver Cancer Surveillance
Chronic liver disease is common and often progresses silently, increasing the risk of cirrhosis and hepatocellular carcinoma when not detected early. With an estimated 1.5 billion people affected worldwide... Read moreIndustry
view channelWerfen and Oxford Nanopore Collaborate on Transplant Assay Development
Werfen (Barcelona, Spain), a global specialized diagnostics company, has announced a strategic collaboration with Oxford Nanopore Technologies (Oxford, UK), which develops nanopore-based sequencing technology,... Read more