Molecular Methods Differentiate E. coli Outbreaks
|
By LabMedica International staff writers Posted on 09 Dec 2015 |
The public health importance of Shiga toxin-producing Escherichia coli (STEC) infections, epidemiological and molecular surveillance systems are essential for early outbreak detection and in recent years, rapid advancements in the use of molecular typing methods have improved STEC surveillance and outbreak detection.
The application of these tools helps to identify disease clusters, refine outbreak case definitions, facilitate case finding, and link human cases to environmental sources. In order to achieve these outcomes, molecular typing assays must possess the discriminatory power required to distinguish between related and nonrelated bacterial isolates, have high reproducibility, and be easy to perform.
A team of scientists led by those at the Alberta Provincial Laboratory for Public Health (Calgary, AB, Canada) assessed the capacity of either high-throughput polymerase chain reaction (PCR) of 49 virulence genes, core-genome single nt variants (SNVs) or k-mer clustering to discriminate between outbreak-associated and sporadic E. coli O157:H7 isolates. Three outbreaks and multiple sporadic isolates from the province of Alberta (Canada) were included in the study. Two of the outbreaks occurred concurrently in 2014 and one occurred in 2012.
The BioMark real-time PCR system (Fluidigm; South San Francisco, CA, USA) was used for real-time PCR amplification of 49 genetic markers using 48.48 dynamic arrays. Amplifications were performed using the dyes, 6-carboxyfluorescein (FAM)- and 6-carboxy-2’, 4, 4’, 5’, 7, 7’-hexachlorofluorescein succinimidyl ester (HEX)-labelled TaqMan probes. All amplification assays included both positive and negative controls. Detection of Stx genes, stx1 and stx2 was determined using a real-time multiplex PCR assay consisting of two separate reactions run on the ABI Prism 7500FAST Sequence Detection System (Life Technologies, Inc.; Burlington, ON, Canada).
Pulsed-field gel electrophoresis (PFGE) and multilocus variable-number tandem repeat analysis (MLVA) were employed as comparator typing methods. The virulence gene profiles of isolates from the 2012 and 2014 Alberta outbreak events and contemporary sporadic isolates were mostly identical; therefore the set of virulence genes chosen in the study were not discriminatory enough to distinguish between outbreak clusters. Concordant with PFGE and MLVA results, core genome single-nucleotide variations (SNV) and k-mer phylogenies clustered isolates from the 2012 and 2014 outbreaks as distinct events. The k-mer phylogenies demonstrated increased discriminatory power compared with core SNV phylogenies.
The authors concluded that whole genome sequencing (WGS) holds significant potential to replace current gold-standard typing methods such as PFGE for the routine surveillance and detection of enteric outbreaks. This shift will be driven by the advantages offered by WGS such as increased discriminatory power and genetic resolution. The study was published on November 26, 2015, in the journal Eurosurveillance.
Related Links:
Alberta Provincial Laboratory for Public Health
Fluidigm
Life Technologies, Inc.
The application of these tools helps to identify disease clusters, refine outbreak case definitions, facilitate case finding, and link human cases to environmental sources. In order to achieve these outcomes, molecular typing assays must possess the discriminatory power required to distinguish between related and nonrelated bacterial isolates, have high reproducibility, and be easy to perform.
A team of scientists led by those at the Alberta Provincial Laboratory for Public Health (Calgary, AB, Canada) assessed the capacity of either high-throughput polymerase chain reaction (PCR) of 49 virulence genes, core-genome single nt variants (SNVs) or k-mer clustering to discriminate between outbreak-associated and sporadic E. coli O157:H7 isolates. Three outbreaks and multiple sporadic isolates from the province of Alberta (Canada) were included in the study. Two of the outbreaks occurred concurrently in 2014 and one occurred in 2012.
The BioMark real-time PCR system (Fluidigm; South San Francisco, CA, USA) was used for real-time PCR amplification of 49 genetic markers using 48.48 dynamic arrays. Amplifications were performed using the dyes, 6-carboxyfluorescein (FAM)- and 6-carboxy-2’, 4, 4’, 5’, 7, 7’-hexachlorofluorescein succinimidyl ester (HEX)-labelled TaqMan probes. All amplification assays included both positive and negative controls. Detection of Stx genes, stx1 and stx2 was determined using a real-time multiplex PCR assay consisting of two separate reactions run on the ABI Prism 7500FAST Sequence Detection System (Life Technologies, Inc.; Burlington, ON, Canada).
Pulsed-field gel electrophoresis (PFGE) and multilocus variable-number tandem repeat analysis (MLVA) were employed as comparator typing methods. The virulence gene profiles of isolates from the 2012 and 2014 Alberta outbreak events and contemporary sporadic isolates were mostly identical; therefore the set of virulence genes chosen in the study were not discriminatory enough to distinguish between outbreak clusters. Concordant with PFGE and MLVA results, core genome single-nucleotide variations (SNV) and k-mer phylogenies clustered isolates from the 2012 and 2014 outbreaks as distinct events. The k-mer phylogenies demonstrated increased discriminatory power compared with core SNV phylogenies.
The authors concluded that whole genome sequencing (WGS) holds significant potential to replace current gold-standard typing methods such as PFGE for the routine surveillance and detection of enteric outbreaks. This shift will be driven by the advantages offered by WGS such as increased discriminatory power and genetic resolution. The study was published on November 26, 2015, in the journal Eurosurveillance.
Related Links:
Alberta Provincial Laboratory for Public Health
Fluidigm
Life Technologies, Inc.
Read the full article by registering today, it's FREE! 

Register now for FREE to LabMedica.com and get access to news and events that shape the world of Clinical Laboratory Medicine. 
- Free digital version edition of LabMedica International sent by email on regular basis
- Free print version of LabMedica International magazine (available only outside USA and Canada).
- Free and unlimited access to back issues of LabMedica International in digital format
- Free LabMedica International Newsletter sent every week containing the latest news
- Free breaking news sent via email
- Free access to Events Calendar
- Free access to LinkXpress new product services
- REGISTRATION IS FREE AND EASY!
Sign in: Registered website members
Sign in: Registered magazine subscribers
Latest Microbiology News
- Comprehensive Review Identifies Gut Microbiome Signatures Associated With Alzheimer’s Disease
- AI-Powered Platform Enables Rapid Detection of Drug-Resistant C. Auris Pathogens
- New Test Measures How Effectively Antibiotics Kill Bacteria
- New Antimicrobial Stewardship Standards for TB Care to Optimize Diagnostics
- New UTI Diagnosis Method Delivers Antibiotic Resistance Results 24 Hours Earlier
- Breakthroughs in Microbial Analysis to Enhance Disease Prediction
- Blood-Based Diagnostic Method Could Identify Pediatric LRTIs
- Rapid Diagnostic Test Matches Gold Standard for Sepsis Detection
- Rapid POC Tuberculosis Test Provides Results Within 15 Minutes
- Rapid Assay Identifies Bloodstream Infection Pathogens Directly from Patient Samples
- Blood-Based Molecular Signatures to Enable Rapid EPTB Diagnosis
- 15-Minute Blood Test Diagnoses Life-Threatening Infections in Children
- High-Throughput Enteric Panels Detect Multiple GI Bacterial Infections from Single Stool Swab Sample
- Fast Noninvasive Bedside Test Uses Sugar Fingerprint to Detect Fungal Infections
- Rapid Sepsis Diagnostic Device to Enable Personalized Critical Care for ICU Patients
- Microfluidic Platform Assesses Neutrophil Function in Sepsis Patients
Channels
Clinical Chemistry
view channel
New PSA-Based Prognostic Model Improves Prostate Cancer Risk Assessment
Prostate cancer is the second-leading cause of cancer death among American men, and about one in eight will be diagnosed in their lifetime. Screening relies on blood levels of prostate-specific antigen... Read more
Extracellular Vesicles Linked to Heart Failure Risk in CKD Patients
Chronic kidney disease (CKD) affects more than 1 in 7 Americans and is strongly associated with cardiovascular complications, which account for more than half of deaths among people with CKD.... Read moreMolecular Diagnostics
view channel
Diagnostic Device Predicts Treatment Response for Brain Tumors Via Blood Test
Glioblastoma is one of the deadliest forms of brain cancer, largely because doctors have no reliable way to determine whether treatments are working in real time. Assessing therapeutic response currently... Read more
Blood Test Detects Early-Stage Cancers by Measuring Epigenetic Instability
Early-stage cancers are notoriously difficult to detect because molecular changes are subtle and often missed by existing screening tools. Many liquid biopsies rely on measuring absolute DNA methylation... Read more
“Lab-On-A-Disc” Device Paves Way for More Automated Liquid Biopsies
Extracellular vesicles (EVs) are tiny particles released by cells into the bloodstream that carry molecular information about a cell’s condition, including whether it is cancerous. However, EVs are highly... Read more
Blood Test Identifies Inflammatory Breast Cancer Patients at Increased Risk of Brain Metastasis
Brain metastasis is a frequent and devastating complication in patients with inflammatory breast cancer, an aggressive subtype with limited treatment options. Despite its high incidence, the biological... Read moreHematology
view channel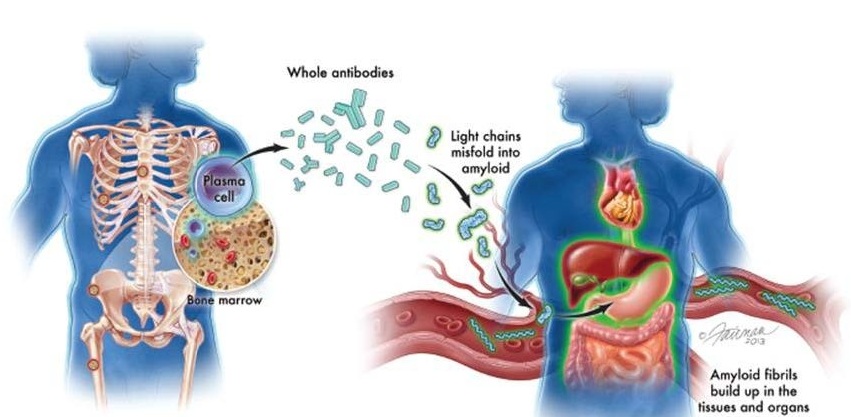
New Guidelines Aim to Improve AL Amyloidosis Diagnosis
Light chain (AL) amyloidosis is a rare, life-threatening bone marrow disorder in which abnormal amyloid proteins accumulate in organs. Approximately 3,260 people in the United States are diagnosed... Read more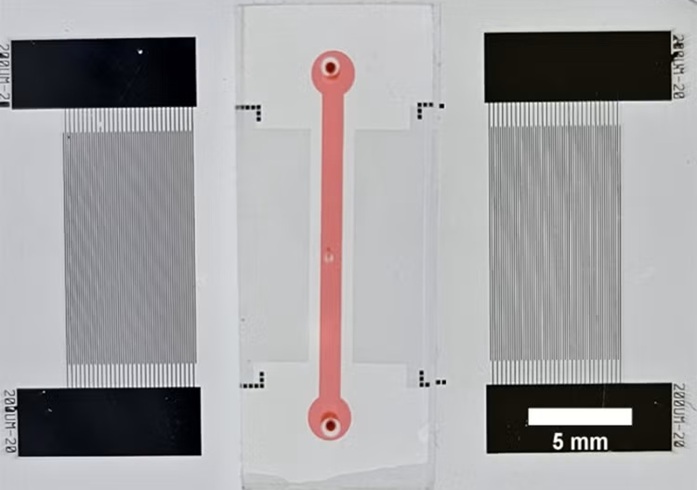
Fast and Easy Test Could Revolutionize Blood Transfusions
Blood transfusions are a cornerstone of modern medicine, yet red blood cells can deteriorate quietly while sitting in cold storage for weeks. Although blood units have a fixed expiration date, cells from... Read more
Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
High-volume hemostasis sections must sustain rapid turnaround while managing reruns and reflex testing. Manual tube handling and preanalytical checks can strain staff time and increase opportunities for error.... Read more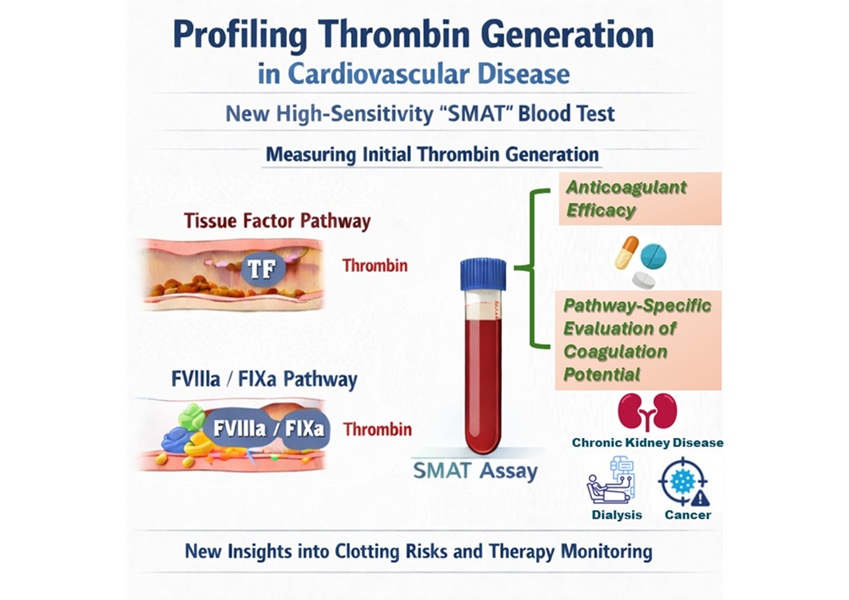
High-Sensitivity Blood Test Improves Assessment of Clotting Risk in Heart Disease Patients
Blood clotting is essential for preventing bleeding, but even small imbalances can lead to serious conditions such as thrombosis or dangerous hemorrhage. In cardiovascular disease, clinicians often struggle... Read moreImmunology
view channelBlood Test Identifies Lung Cancer Patients Who Can Benefit from Immunotherapy Drug
Small cell lung cancer (SCLC) is an aggressive disease with limited treatment options, and even newly approved immunotherapies do not benefit all patients. While immunotherapy can extend survival for some,... Read more
Whole-Genome Sequencing Approach Identifies Cancer Patients Benefitting From PARP-Inhibitor Treatment
Targeted cancer therapies such as PARP inhibitors can be highly effective, but only for patients whose tumors carry specific DNA repair defects. Identifying these patients accurately remains challenging,... Read more
Ultrasensitive Liquid Biopsy Demonstrates Efficacy in Predicting Immunotherapy Response
Immunotherapy has transformed cancer treatment, but only a small proportion of patients experience lasting benefit, with response rates often remaining between 10% and 20%. Clinicians currently lack reliable... Read morePathology
view channel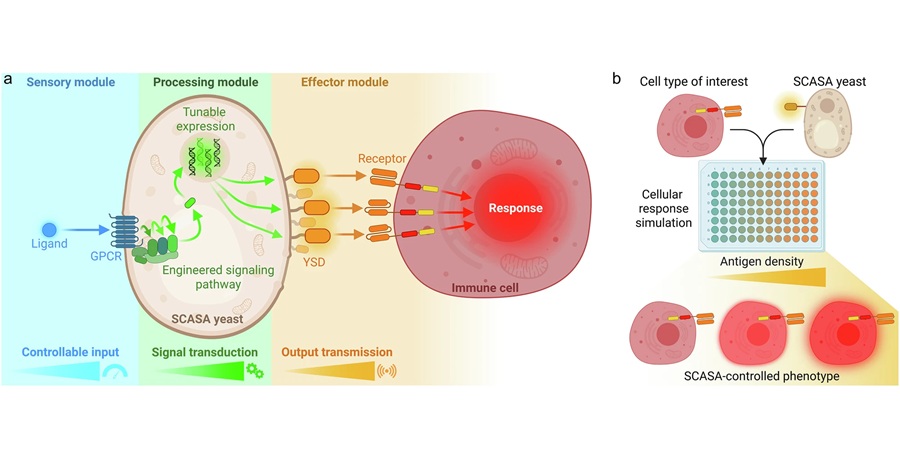
Engineered Yeast Cells Enable Rapid Testing of Cancer Immunotherapy
Developing new cancer immunotherapies is a slow, costly, and high-risk process, particularly for CAR T cell treatments that must precisely recognize cancer-specific antigens. Small differences in tumor... Read more
First-Of-Its-Kind Test Identifies Autism Risk at Birth
Autism spectrum disorder is treatable, and extensive research shows that early intervention can significantly improve cognitive, social, and behavioral outcomes. Yet in the United States, the average age... Read moreTechnology
view channel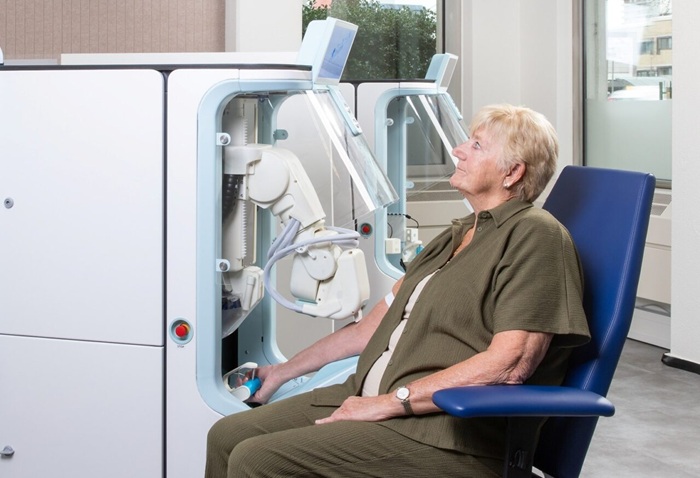
Robotic Technology Unveiled for Automated Diagnostic Blood Draws
Routine diagnostic blood collection is a high‑volume task that can strain staffing and introduce human‑dependent variability, with downstream implications for sample quality and patient experience.... Read more
ADLM Launches First-of-Its-Kind Data Science Program for Laboratory Medicine Professionals
Clinical laboratories generate billions of test results each year, creating a treasure trove of data with the potential to support more personalized testing, improve operational efficiency, and enhance patient care.... Read moreAptamer Biosensor Technology to Transform Virus Detection
Rapid and reliable virus detection is essential for controlling outbreaks, from seasonal influenza to global pandemics such as COVID-19. Conventional diagnostic methods, including cell culture, antigen... Read more
AI Models Could Predict Pre-Eclampsia and Anemia Earlier Using Routine Blood Tests
Pre-eclampsia and anemia are major contributors to maternal and child mortality worldwide, together accounting for more than half a million deaths each year and leaving millions with long-term health complications.... Read moreIndustry
view channelNew Collaboration Brings Automated Mass Spectrometry to Routine Laboratory Testing
Mass spectrometry is a powerful analytical technique that identifies and quantifies molecules based on their mass and electrical charge. Its high selectivity, sensitivity, and accuracy make it indispensable... Read more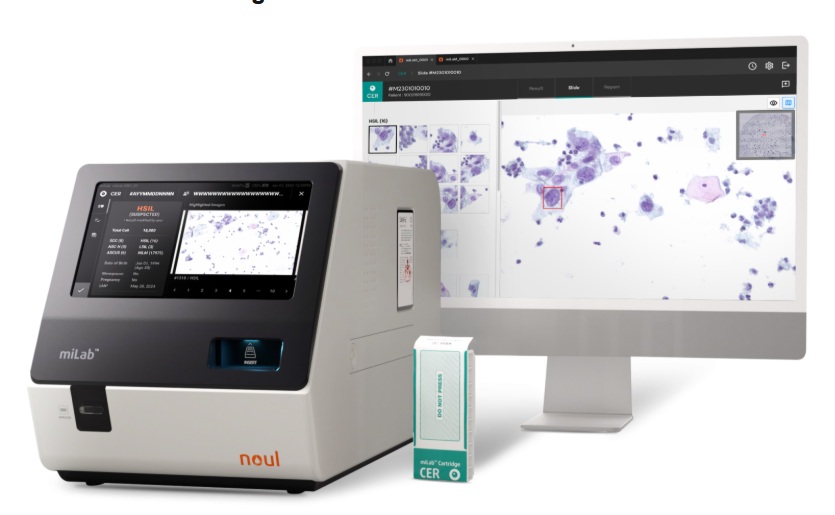
AI-Powered Cervical Cancer Test Set for Major Rollout in Latin America
Noul Co., a Korean company specializing in AI-based blood and cancer diagnostics, announced it will supply its intelligence (AI)-based miLab CER cervical cancer diagnostic solution to Mexico under a multi‑year... Read more
Diasorin and Fisher Scientific Enter into US Distribution Agreement for Molecular POC Platform
Diasorin (Saluggia, Italy) has entered into an exclusive distribution agreement with Fisher Scientific, part of Thermo Fisher Scientific (Waltham, MA, USA), for the LIAISON NES molecular point-of-care... Read more