Shotgun Metagenomic Technique Detects Tuberculosis Bacteria in Patient Samples Without Culture or Enrichment
|
By LabMedica International staff writers Posted on 15 Oct 2014 |
Infectious disease researchers have developed a new approach for the diagnosis of tuberculosis (TB) that relies on shotgun metagenomics, a method for direct sequencing of DNA extracted from sputum samples, which detects and characterizes the Mycobacterium that cause TB without the need for time-consuming culture or enrichment.
Metagenomics is the study of genetic material recovered directly from environmental samples. In metagenomic sequencing, DNA is recovered directly from environmental samples in an untargeted manner with the goal of obtaining an unbiased sample from all genes of all members of the community. Recent studies used shotgun Sanger sequencing or pyrosequencing to recover the sequences of the reads. Shotgun sequencing is a sequencing method designed for analysis of DNA sequences longer than 1,000 base pairs, up to and including entire chromosomes. This method requires the target DNA to be broken into random fragments. After sequencing individual fragments, the sequences can be reassembled on the basis of their overlapping regions
Investigators at Warwick Medical School (United Kingdom) explored the potential of shotgun metagenomics to detect and characterize strains from the Mycobacterium tuberculosis complex in smear-positive sputum samples. To this end, they analyzed eight samples obtained from tuberculosis patients from Gambia.
The concentration of DNA present in each extract was determined using the Qubit (Invitrogen Ltd., Paisley, United Kingdom) 2.0 fluorometer and Qubit dsDNA Assay Kits according to the manufacturer’s protocol using the HS (high-sensitivity) or BR (broad-range) kits, depending on the DNA concentration. There was no detectable DNA in the negative control samples with the HS kit, which is sensitive down to 10 picograms per microliter. DNA extracts were diluted to 0.2 nanograms per microliter and were then converted into sequencing libraries using the Illumina (Little Chesterford, United Kingdom) Nextera XT sample preparation kit. The libraries were sequenced on the Illumina MiSeq instrument at the University of Warwick.
Using this methodology, the investigators were able to detect sequences from the M. tuberculosis complex in all eight samples, with coverage of the H37Rv reference genome ranging from 0.002X to 0.7X. By analyzing the distribution of large sequence polymorphisms (deletions and the locations of the insertion element IS6110) and single nucleotide polymorphisms (SNPs), they were able to assign seven of eight metagenome-derived genomes to a species and lineage within the M. tuberculosis complex. Two metagenome-derived mycobacterial genomes were assigned to M. africanum, a species largely confined to West Africa; the others that could be assigned belonged to lineages T, H, or LAM within the clade of "modern" M. tuberculosis strains.
"Laboratory diagnosis of TB using conventional approaches is a long drawn-out process, which takes weeks or months," said senior author Dr. Mark Pallen, professor of microbial genomics at Warwick Medical School. "Plus, relying on laboratory culture means using techniques that date back to the 1880s! Metagenomics using the latest high-throughput sequencing technologies and some smart bioinformatics, allows us to detect and characterize the bacteria that cause TB in a matter of a day or two, without having to grow the bacteria, while also giving us key insights into their genome sequences and the lineages that they belong to. We have provided proof-of-principle here, but we still need to make metagenomics more sensitive and improve our workflows. But, caveats aside, let us celebrate the fact that metagenomics stands ready to document past and present infections, shedding light on the emergence, evolution, and spread of microbial pathogens."
The shotgun metagenomics study was published in the September 23, 2014, online edition of the journal PeerJ.
Related Links:
Warwick Medical School
Invitrogen Ltd.
Illumina
Metagenomics is the study of genetic material recovered directly from environmental samples. In metagenomic sequencing, DNA is recovered directly from environmental samples in an untargeted manner with the goal of obtaining an unbiased sample from all genes of all members of the community. Recent studies used shotgun Sanger sequencing or pyrosequencing to recover the sequences of the reads. Shotgun sequencing is a sequencing method designed for analysis of DNA sequences longer than 1,000 base pairs, up to and including entire chromosomes. This method requires the target DNA to be broken into random fragments. After sequencing individual fragments, the sequences can be reassembled on the basis of their overlapping regions
Investigators at Warwick Medical School (United Kingdom) explored the potential of shotgun metagenomics to detect and characterize strains from the Mycobacterium tuberculosis complex in smear-positive sputum samples. To this end, they analyzed eight samples obtained from tuberculosis patients from Gambia.
The concentration of DNA present in each extract was determined using the Qubit (Invitrogen Ltd., Paisley, United Kingdom) 2.0 fluorometer and Qubit dsDNA Assay Kits according to the manufacturer’s protocol using the HS (high-sensitivity) or BR (broad-range) kits, depending on the DNA concentration. There was no detectable DNA in the negative control samples with the HS kit, which is sensitive down to 10 picograms per microliter. DNA extracts were diluted to 0.2 nanograms per microliter and were then converted into sequencing libraries using the Illumina (Little Chesterford, United Kingdom) Nextera XT sample preparation kit. The libraries were sequenced on the Illumina MiSeq instrument at the University of Warwick.
Using this methodology, the investigators were able to detect sequences from the M. tuberculosis complex in all eight samples, with coverage of the H37Rv reference genome ranging from 0.002X to 0.7X. By analyzing the distribution of large sequence polymorphisms (deletions and the locations of the insertion element IS6110) and single nucleotide polymorphisms (SNPs), they were able to assign seven of eight metagenome-derived genomes to a species and lineage within the M. tuberculosis complex. Two metagenome-derived mycobacterial genomes were assigned to M. africanum, a species largely confined to West Africa; the others that could be assigned belonged to lineages T, H, or LAM within the clade of "modern" M. tuberculosis strains.
"Laboratory diagnosis of TB using conventional approaches is a long drawn-out process, which takes weeks or months," said senior author Dr. Mark Pallen, professor of microbial genomics at Warwick Medical School. "Plus, relying on laboratory culture means using techniques that date back to the 1880s! Metagenomics using the latest high-throughput sequencing technologies and some smart bioinformatics, allows us to detect and characterize the bacteria that cause TB in a matter of a day or two, without having to grow the bacteria, while also giving us key insights into their genome sequences and the lineages that they belong to. We have provided proof-of-principle here, but we still need to make metagenomics more sensitive and improve our workflows. But, caveats aside, let us celebrate the fact that metagenomics stands ready to document past and present infections, shedding light on the emergence, evolution, and spread of microbial pathogens."
The shotgun metagenomics study was published in the September 23, 2014, online edition of the journal PeerJ.
Related Links:
Warwick Medical School
Invitrogen Ltd.
Illumina
Read the full article by registering today, it's FREE! 

Register now for FREE to LabMedica.com and get access to news and events that shape the world of Clinical Laboratory Medicine. 
- Free digital version edition of LabMedica International sent by email on regular basis
- Free print version of LabMedica International magazine (available only outside USA and Canada).
- Free and unlimited access to back issues of LabMedica International in digital format
- Free LabMedica International Newsletter sent every week containing the latest news
- Free breaking news sent via email
- Free access to Events Calendar
- Free access to LinkXpress new product services
- REGISTRATION IS FREE AND EASY!
Sign in: Registered website members
Sign in: Registered magazine subscribers
Latest Microbiology News
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- Rapid Color Test Stratifies Virulent and Resistant Staph Strains
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Syndromic Panel Enables Rapid Identification of Bloodstream Infections
- RNA-Based Workflow Identifies Active Skin Microbes for Dermatology Research
- Cost-Effective Sampling and Sequencing Workflow Identifies ICU Infection Hotspots
- New Bacterial Target Identified for Early Detection of Noma
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
Channels
Clinical Chemistry
view channel
Urine-Based Nanosensor Tracks Lung Cancer and Fibrosis Noninvasively
Lung cancer remains difficult to monitor for early progression and treatment resistance, while pulmonary fibrosis continues to pose major challenges for early diagnosis. Clinicians need repeatable, noninvasive... Read more
Blood-Based Alzheimer’s Test Gains CE Mark for Amyloid Pathology Detection
Alzheimer’s disease is the most common cause of dementia, yet confirmatory testing remains invasive and hard to access. Diagnosis currently takes an average of 3.5 years, and about 75% of people with dementia... Read moreMolecular Diagnostics
view channel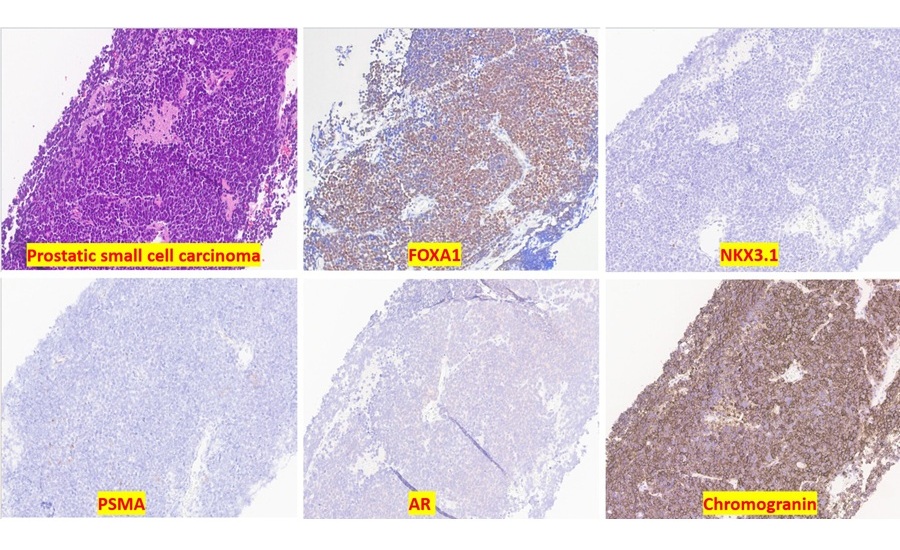
Sensitive Protein Marker Aids Diagnosis of Small Cell Prostate Cancer
Accurate identification of aggressive prostate cancer subtypes can be difficult when tumors lose expression of lineage markers used in routine pathology. Small cell carcinoma of the prostate, in particular,... Read more
Rapid Multiplex PCR Test Detects 11 Gastrointestinal Pathogens from Single Sample
Cepheid’s Xpert GI Panel has received CE marking under the In Vitro Diagnostic Medical Devices Regulation (IVDR) and is expected to begin shipping to countries that accept the CE mark in the coming weeks.... Read moreHematology
view channel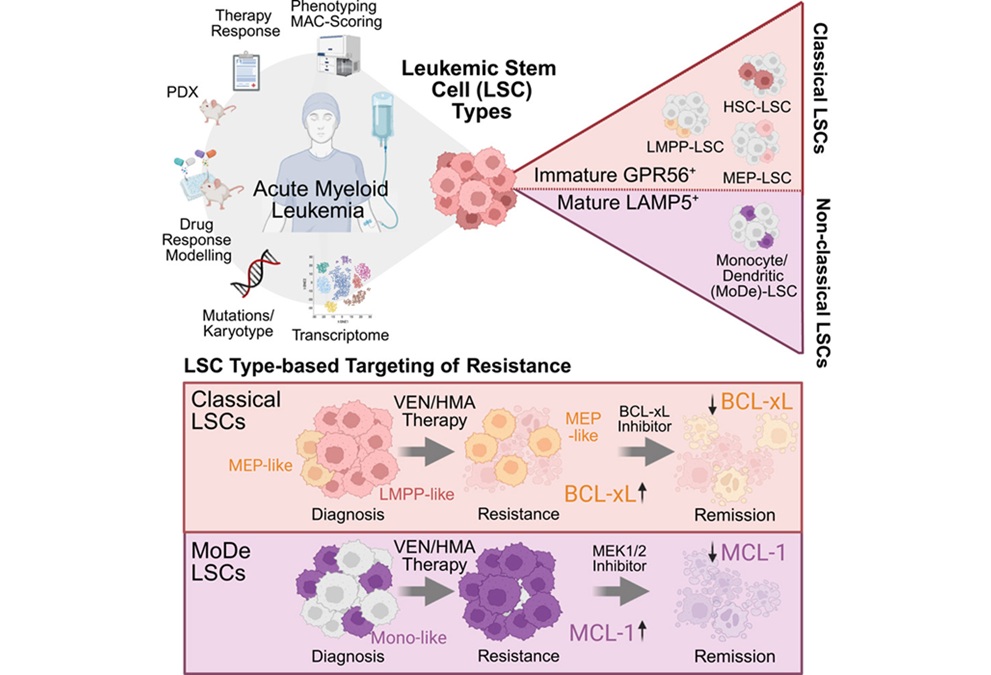
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read more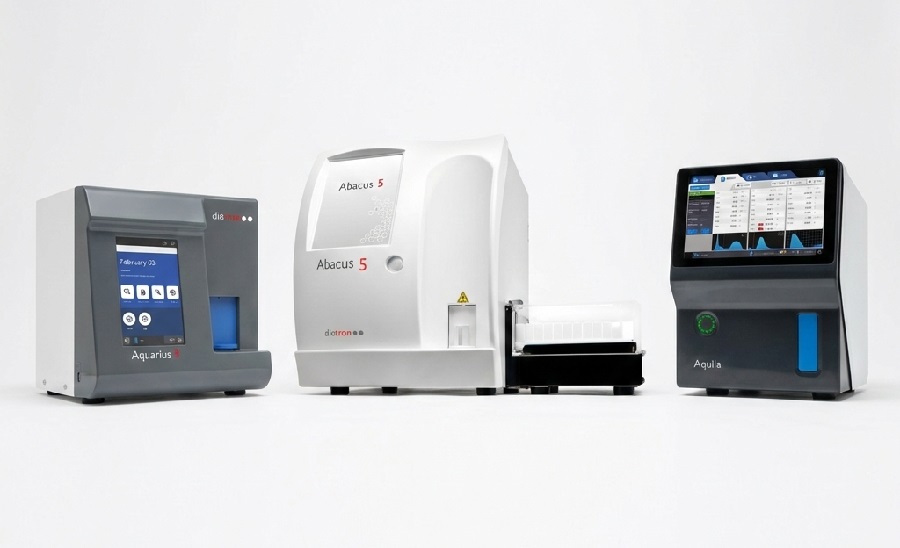
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read moreImmunology
view channel
Routine TB Screening Test May Reveal Immune Aging and Mortality Risk
Immune aging is associated with weaker responses to vaccination, greater risks of infection, and higher levels of inflammation. Leveraging routinely ordered laboratory tests to quantify that responsiveness... Read more
Biomarkers and Molecular Testing Advance Precision Allergy Care
Allergic diseases often present with similar symptoms but can be driven by distinct biological mechanisms, making standardized care inefficient for many patients. Historically, individuals with pollen... Read morePathology
view channel
FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
Risk assessment at diagnosis is central to guiding therapy for early-stage, hormone receptor-positive, human epidermal growth factor receptor 2-negative (HR+/HER2-) invasive breast cancer, where overtreatment... Read more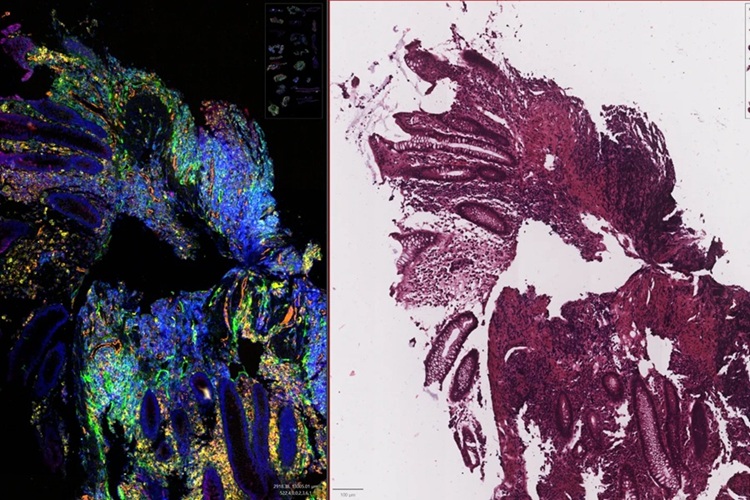
New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
Pathologists worldwide rely on hematoxylin and eosin (H&E) slides to examine tissue architecture, yet these stains do not reveal the underlying molecular activity that often drives disease.... Read moreTechnology
view channel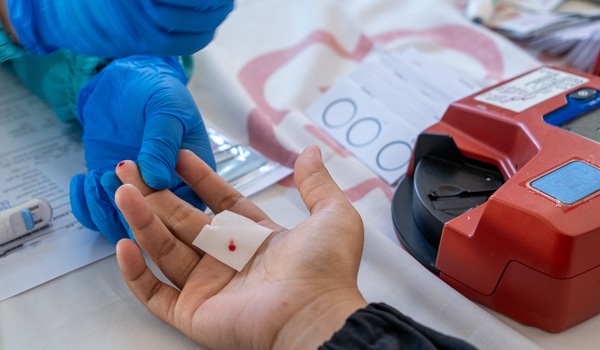
Point-of-Care Testing Enhances Health Literacy and Self-Management in Chronic Disease
Limited access to general practitioners and pathology services can delay diagnosis and monitoring for people in regional and remote communities. Rapid, on-the-spot testing can shorten turnaround times... Read more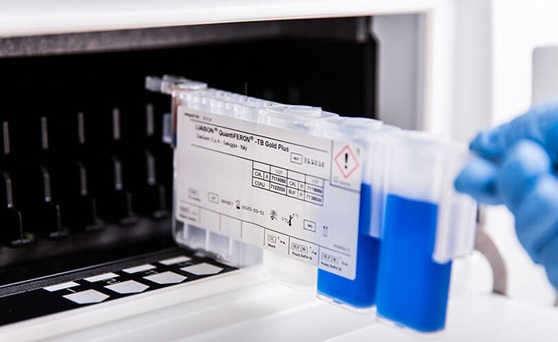
Fully Automated Sample-to-Insight Workflow Advances Latent TB Testing
Latent tuberculosis remains a substantial testing workload for clinical laboratories as screening programs expand. Despite this growth, only about 40% of testing has shifted from traditional skin tests... Read moreIndustry
view channel
AI-Powered Multi-Functional Analyzer Wins German Innovation Award
Hematology services are increasingly delivered across distributed care settings, where limited staffing and complex workflows can extend turnaround times. Advanced morphology review still often depends... Read more