Microfluidics Device Captures and Isolates Slow Growing Gut Bacteria
|
By LabMedica International staff writers Posted on 15 Jul 2014 |
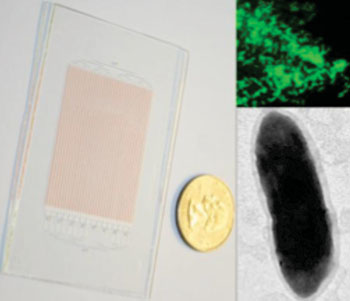
Image: Glass SlipChip for growing microbes, shown next to a US quarter dollar coin (left); fluorescent in situ hybridization image of the target organism (right, top); transmission electron microscopy image of a single cell of the target organism (right, bottom) (Photo courtesy of the California Institute of Technology).
Microbiologists have used a novel "lab-on-a-chip" approach to isolate and cultivate fastidious, slow growing bacteria from the human digestive tract.
The majority of microbes that comprises the human gut biome have not been cultured, due in part to the difficulties of both identifying proper growth conditions and characterizing and isolating each species.
Investigators at the California Institute of Technology (Pasadena, USA) developed a microfluidics-based, genetically targeted approach to overcome these problems. Their "SlipChip" device was constructed from two glass slides, each the size of a credit card, that were etched with tiny grooves that became channels when the grooved surfaces were stacked atop one another. When a sample, such as a mixed assortment of bacterial species from a colonoscopy biopsy, was applied to the device, the interconnected channels of the top chip turned the channels into individual wells, with each well ideally holding a single microbe. Once sequestered in an isolated well, each individual bacterium was able to divide and grow without having to compete for resources with other types of faster-growing microbes.
The beauty of the system was that each well could be divided into two compartments. One compartment was used for DNA sequencing and mapping studies while the other maintained a living example of the microbe for further culture and study.
The investigators validated this approach by cultivating a bacterium from a human cecal biopsy. Genetic mapping of the organism showed that it was a representative of a previously unidentified genus of the Ruminococcaceae family and that its genetic signature was listed among the high-priority group of the [US] National Institutes of Health's Human Microbiome Project’s "Most Wanted" list.
"Although a genomic sequence of the new organism is a useful tool, further studies are needed to learn how this species of microbe is involved in human health," said senior author Dr. Rustem Ismagilov, professor of chemistry and chemical engineering at the California Institute of Technology.
The study was published in the June 25, 2014, online edition of the journal Proceedings of the National Academy of Sciences of the United States of America (PNAS).
Related Links:
California Institute of Technology
The majority of microbes that comprises the human gut biome have not been cultured, due in part to the difficulties of both identifying proper growth conditions and characterizing and isolating each species.
Investigators at the California Institute of Technology (Pasadena, USA) developed a microfluidics-based, genetically targeted approach to overcome these problems. Their "SlipChip" device was constructed from two glass slides, each the size of a credit card, that were etched with tiny grooves that became channels when the grooved surfaces were stacked atop one another. When a sample, such as a mixed assortment of bacterial species from a colonoscopy biopsy, was applied to the device, the interconnected channels of the top chip turned the channels into individual wells, with each well ideally holding a single microbe. Once sequestered in an isolated well, each individual bacterium was able to divide and grow without having to compete for resources with other types of faster-growing microbes.
The beauty of the system was that each well could be divided into two compartments. One compartment was used for DNA sequencing and mapping studies while the other maintained a living example of the microbe for further culture and study.
The investigators validated this approach by cultivating a bacterium from a human cecal biopsy. Genetic mapping of the organism showed that it was a representative of a previously unidentified genus of the Ruminococcaceae family and that its genetic signature was listed among the high-priority group of the [US] National Institutes of Health's Human Microbiome Project’s "Most Wanted" list.
"Although a genomic sequence of the new organism is a useful tool, further studies are needed to learn how this species of microbe is involved in human health," said senior author Dr. Rustem Ismagilov, professor of chemistry and chemical engineering at the California Institute of Technology.
The study was published in the June 25, 2014, online edition of the journal Proceedings of the National Academy of Sciences of the United States of America (PNAS).
Related Links:
California Institute of Technology
Latest Microbiology News
- Gut Microbiome Signatures Help Identify Risk of IBD Progression
- FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
- New AMR Assay Supports Rapid Infection Control Screening in Hospitals
- Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- Rapid Color Test Stratifies Virulent and Resistant Staph Strains
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Syndromic Panel Enables Rapid Identification of Bloodstream Infections
Channels
Clinical Chemistry
view channel
Urine-Based Test Shows Promise for Autism Screening in Children
Autism spectrum disorder (ASD) is commonly diagnosed through behavioral assessments, which can involve long waits that delay intervention. Earlier identification is linked to better developmental outcomes,... Read more
Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
Childhood epilepsy remains a major neurological disorder with unmet needs for accurate, non-invasive biomarkers, as conventional tests such as electroencephalography and neuroimaging can have limited sensitivity... Read moreMolecular Diagnostics
view channel
Blood-Based MRD Monitoring Supports Relapse Prevention in Leukemia
In myelodysplastic syndromes (MDS) and acute myeloid leukemia (AML), molecular blood testing for minimal residual disease (MRD) can detect signs of impending relapse before clinical symptoms emerge.... Read more
Genomic Test Predicts Chemotherapy Benefit in Metastatic Prostate Cancer
Metastatic prostate cancer remains a major treatment challenge, particularly when deciding whether to add chemotherapy to hormonal therapy. In the United States, about 334,000 men are diagnosed with prostate... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channel
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more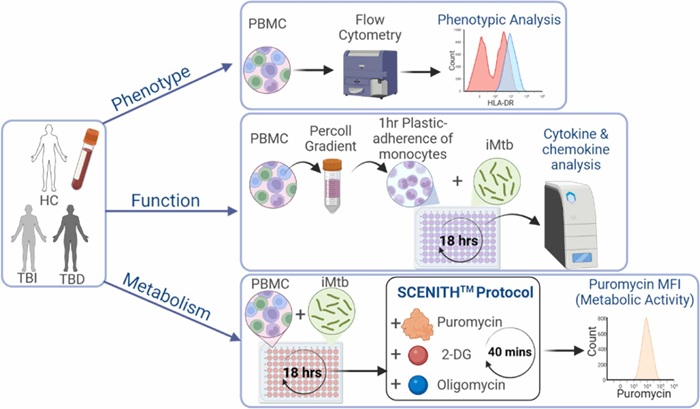
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read morePathology
view channel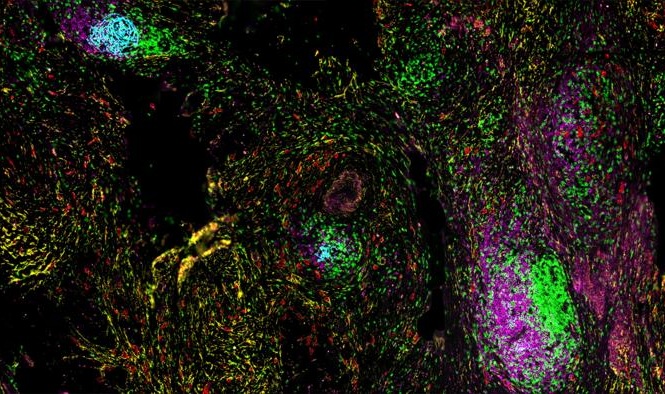
AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
Tertiary lymphoid structures are emerging as important indicators of antitumor immunity, but their heterogeneity and spatial context within tumors remain difficult to capture through routine diagnostics.... Read more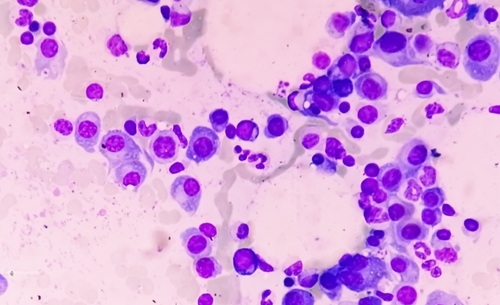
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel