Rapid Metagenomics Diagnoses Antibiotic Resistant Bloodstream Infections 18-42 Hours Faster Than Conventional Tests
|
By LabMedica International staff writers Posted on 20 Apr 2023 |

Bloodstream infections can quickly progress to sepsis, multiple organ failure, and even death. Timely and appropriate antibiotic therapy is crucial for managing the infection. Antimicrobial resistance (AMR) poses a significant challenge in treating bloodstream infections. Current clinical methods for identifying the causative pathogen are lengthy and labor-intensive, involving two culture and sensitivity tests that take at least 1 to 3 days to complete—first isolating and identifying the pathogen and then performing antimicrobial susceptibility testing. Now, new research presented at ECCMID 2023 demonstrates that metagenomic sequencing can offer rapid and actionable AMR predictions for treating bloodstream infections much faster than traditional laboratory tests, potentially saving lives and improving antibiotic management.
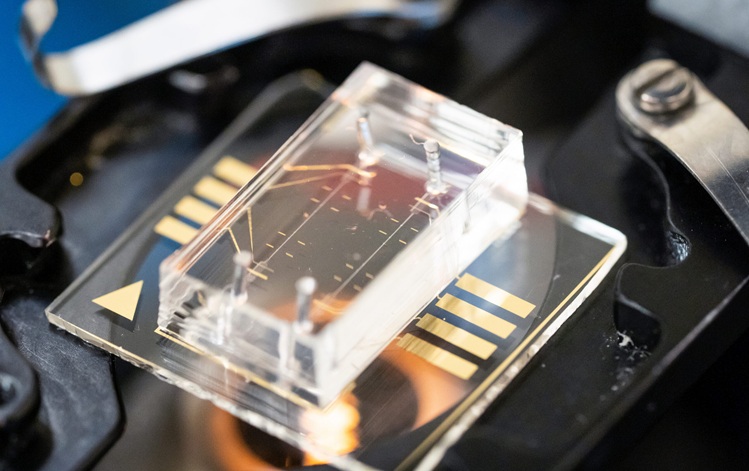
The study conducted by researchers at the University of Oxford (Oxford, UK) reveals that rapid metagenomics can provide accurate results within just six hours of detecting bacterial growth in a blood sample. Clinical metagenomics sequences all genetic material, including infectious pathogens, in a sample simultaneously, reducing the time spent on running tests, waiting for results, and conducting additional tests. For their study, the Oxford researchers randomly selected 210 positive and 61 negative blood culture specimens for metagenomic sequencing, using the Oxford Nanopore GridION platform to sequence DNA. The sequences were utilized to identify the pathogen species causing infections and to detect common species that can contaminate blood cultures.
Sequencing successfully identified 99% of infecting pathogens, including polymicrobial infections and contaminants, and yielded negative results in 100% of culture-negative samples. In some cases, sequencing detected probable infection causes that routine cultures missed, while in others, it identified uncultivable species when a result could not be determined. Sequencing could also detect antibiotic resistance in the 10 most common infection causes. A total of 741 resistant and 4047 sensitive antibiotic-pathogen combinations were examined, with traditional culture-based testing and sequencing results agreeing 92% of the time. Comparable performance could be achieved using raw reads after just two hours of sequencing, with an overall agreement of 90%. The average time from sample extraction to sequencing was four hours, with complete AMR prediction achieved two hours later, providing actionable AMR results 18-42 hours sooner than conventional laboratory methods.
“Antibiotic resistant bloodstream infections are a leading killer in hospitals, and rapidly starting the right antibiotic saves lives,” said Dr. Kumeren Govender from the John Radcliffe Hospital, University of Oxford, who led the study. “Our results suggests that metagenomics is a powerful tool for the rapid and accurate diagnosis of pathogenic organisms and antimicrobial resistance, allowing for effective treatment 18 to 42 hours earlier than would be possible using standard culture techniques.”
“This is a really exciting breakthrough that means we will be able to diagnose the cause of patients’ infections faster and more completely than has been possible before,” added David Eyre, Professor of Infectious Diseases at the University of Oxford, who co-led the study. “We are working hard to continue to overcome some of the remaining barriers to metagenomic sequencing being used more widely, which include its current high cost, further improving accuracy, and creating improved laboratory expertise in these new technologies and simpler workflows for interpreting results.”
Related Links:
University of Oxford
Latest Microbiology News
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- Rapid Color Test Stratifies Virulent and Resistant Staph Strains
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Syndromic Panel Enables Rapid Identification of Bloodstream Infections
- RNA-Based Workflow Identifies Active Skin Microbes for Dermatology Research
- Cost-Effective Sampling and Sequencing Workflow Identifies ICU Infection Hotspots
- New Bacterial Target Identified for Early Detection of Noma
- Genomic Analysis Links Emerging Streptococcal Strains to Specific Infections
- Rapid Urine Test Speeds Antibiotic Selection for UTIs
- WHO Endorses Rapid Point-of-Care Testing to Improve TB Detection
- Breath Analysis Approach Offers Rapid Detection of Bacterial Infection
Channels
Clinical Chemistry
view channel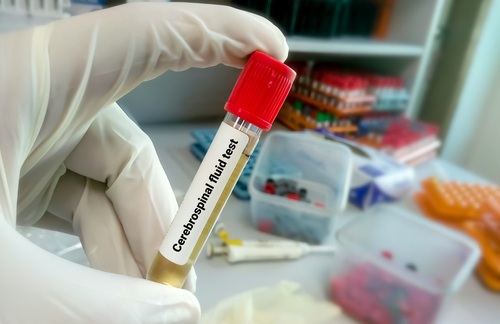
Ultrasensitive Test Detects Key Biomarker of Frontotemporal Dementia Subtype
Dementia affects more than 57 million people worldwide and is projected to nearly double within two decades, straining health systems and families. While biomarkers now enable accurate identification of... Read more
Routine Blood Tests Years Before Pregnancy Could Identify Preeclampsia Risk
High blood pressure during pregnancy is common and can progress to pre-eclampsia, making close monitoring at antenatal visits essential. However, most risk assessment begins only after pregnancy has started.... Read moreMolecular Diagnostics
view channel
Liquid Biopsy Biomarkers Distinguish Inflammatory Breast Cancer and Support Monitoring
Inflammatory breast cancer is among the most aggressive forms of breast malignancy and remains challenging to diagnose and monitor. Obtaining tumor tissue can be difficult, and standard genome and RNA... Read more
Blood Test Maps Tumor Microenvironment to Predict Immunotherapy Response
Immunotherapy has transformed cancer care, yet durable benefit remains limited to a subset of patients, and clinicians still lack reliable tools to predict response before treatment begins.... Read more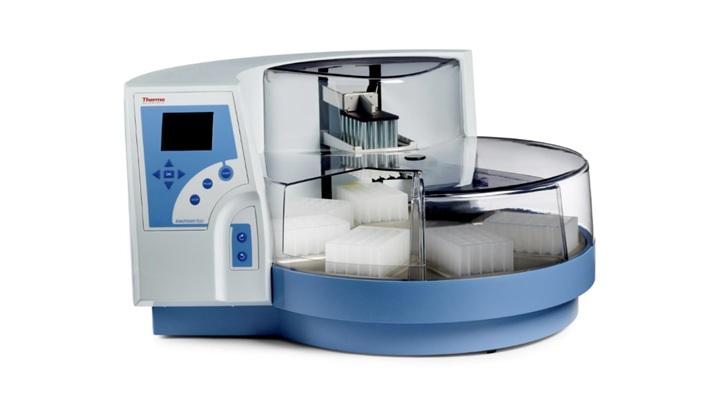
Multiplex Respiratory Panel Integrates Automated Extraction to Streamline High-Volume Testing
Respiratory infections drive heavy testing volumes in clinical laboratories, where accurate, timely results across multiple pathogens are essential. Many labs are seeking to streamline workflows and increase... Read moreHematology
view channel
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read more
Blood Test Enables Early Detection of Multiple Myeloma Relapse
Bone marrow biopsies remain central to diagnosing and monitoring multiple myeloma, yet the procedure is painful, invasive, and often repeated over time. Older patients—who represent most new cases—can... Read moreImmunology
view channel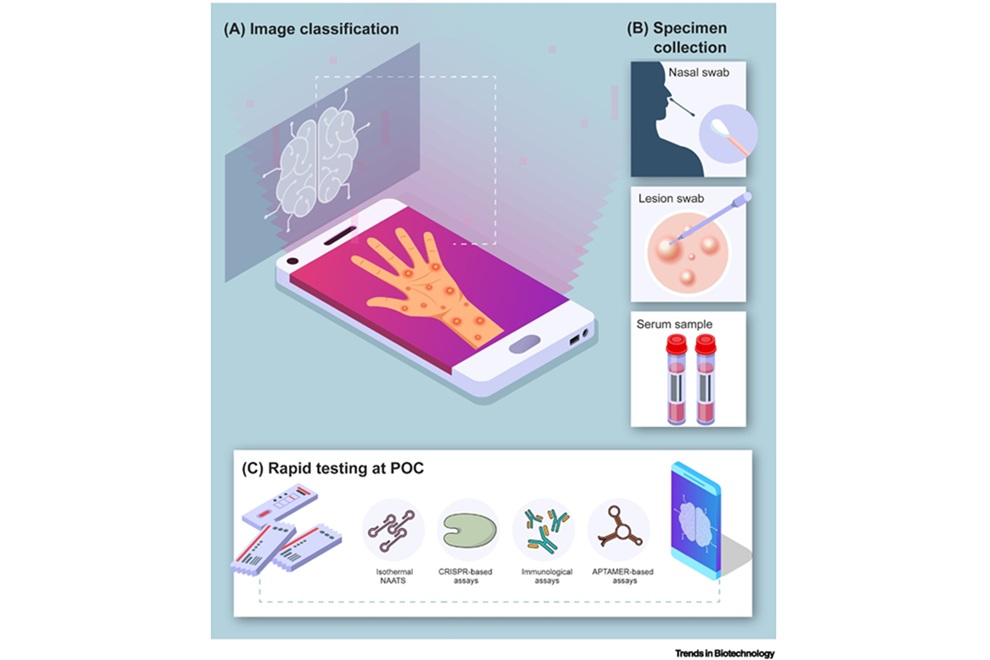
Point-of-Care Tests Could Expand Access to Mpox Diagnosis
Mpox outbreaks in non-endemic regions have underscored the need for rapid, accessible diagnostics to limit transmission. Polymerase chain reaction (PCR) remains the clinical reference, yet it depends on... Read more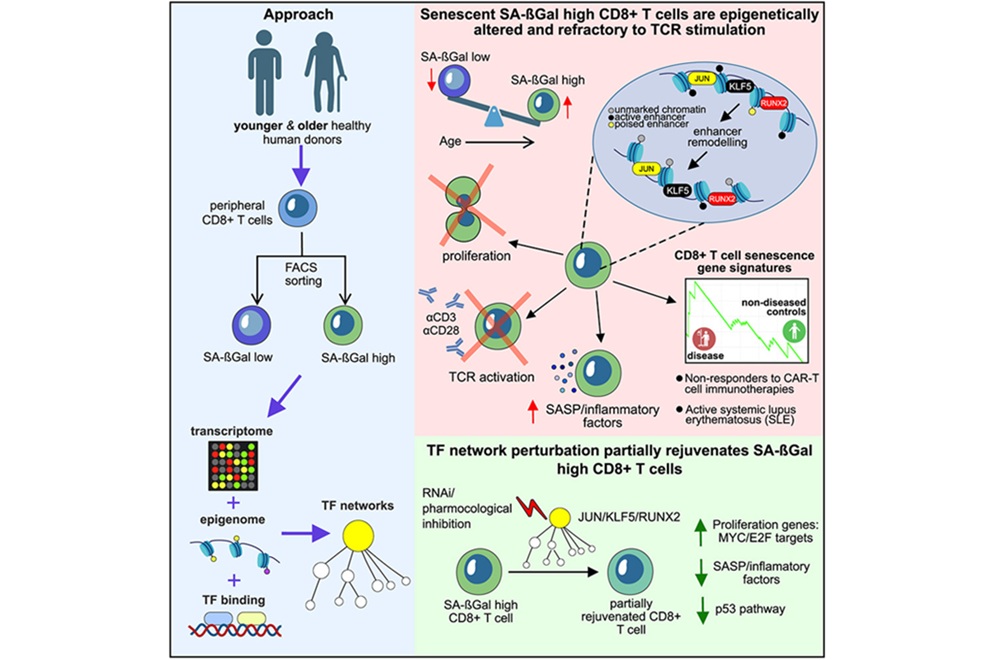
T-Cell Senescence Profiling May Predict CAR T Responses
Chimeric antigen receptor (CAR) T-cell therapy can deliver striking, durable remissions, yet many patients experience minimal or no benefit. The quality of patient-derived cytotoxic T lymphocytes used... Read morePathology
view channel
FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
Risk assessment at diagnosis is central to guiding therapy for early-stage, hormone receptor-positive, human epidermal growth factor receptor 2-negative (HR+/HER2-) invasive breast cancer, where overtreatment... Read more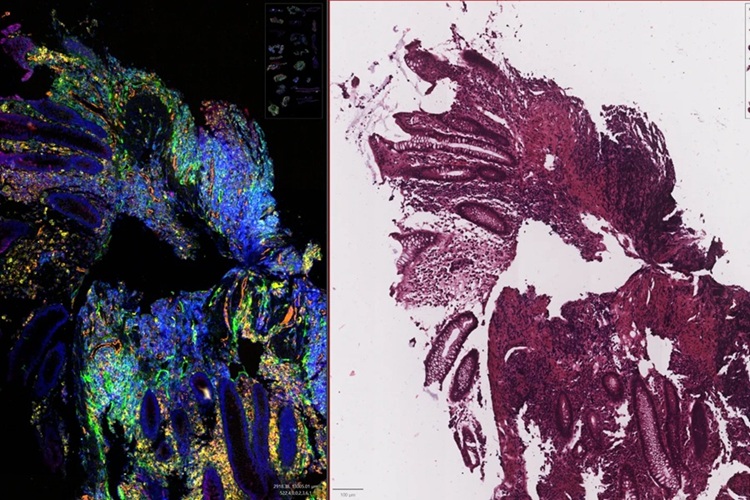
New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
Pathologists worldwide rely on hematoxylin and eosin (H&E) slides to examine tissue architecture, yet these stains do not reveal the underlying molecular activity that often drives disease.... Read moreTechnology
view channel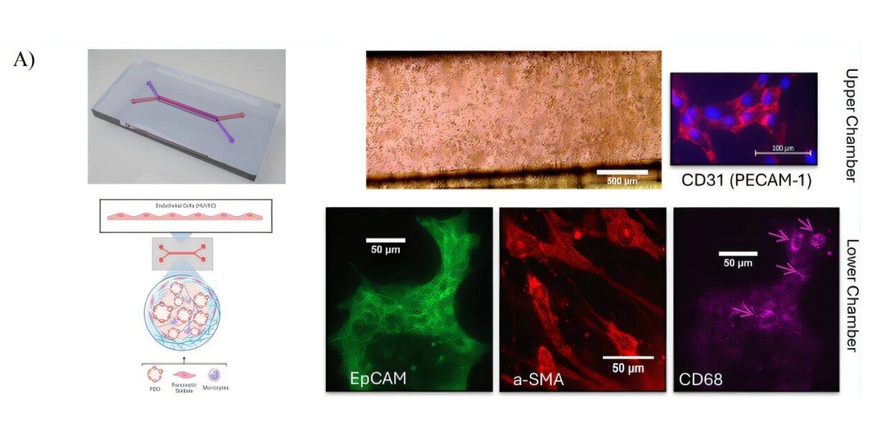
Tumor-on-a-Chip Platform Models Pancreatic Cancer Treatment Response
Pancreatic cancer remains one of the hardest malignancies to treat because tumors are embedded within a dense microenvironment that shapes growth and therapy response. Standard laboratory models often... Read more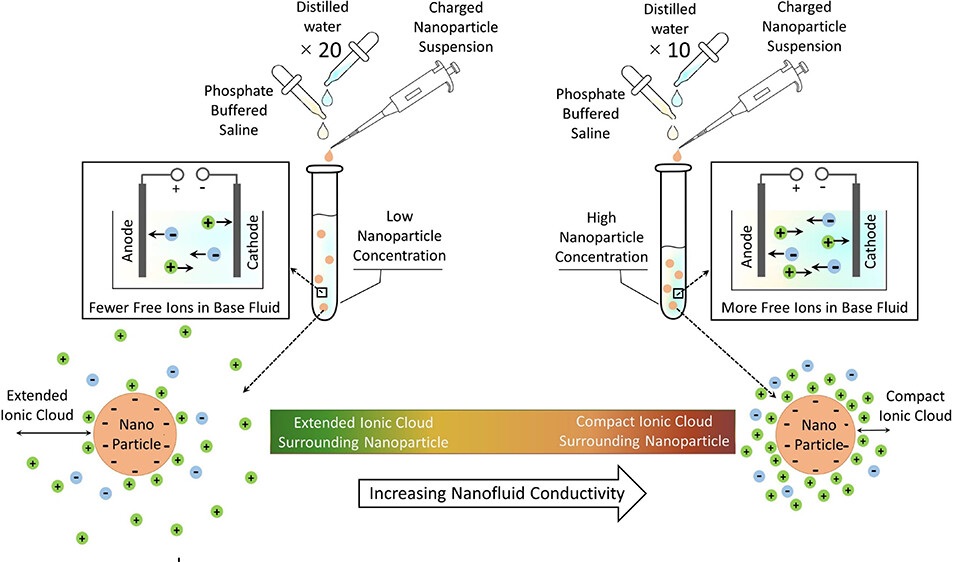
New Platform Captures Extracellular Vesicles for Early Cancer Detection
Early diagnosis remains the most effective way to reduce cancer mortality, yet many screening tools miss disease at its earliest stages. Biomarkers shed by tumors into blood and other fluids can be scarce... Read moreIndustry
view channel
Roche to Acquire PathAI for Up to $1.05 Billion to Strengthen AI Diagnostics Portfolio
Roche has entered into a definitive merger agreement to acquire PathAI, a company focused on digital pathology and artificial intelligence for pathology laboratories and the biopharma industry.... Read more