Gut Microbiome Data Helps Routine Screening of Cardiovascular Disease
|
By LabMedica International staff writers Posted on 22 Sep 2020 |
Image: Gut Microbiome Data Helps Routine Screening of Cardiovascular Disease (Photo courtesy of Nishant Mehta PhD).
Besides genetics and environmental factors, gut microbiota has emerged as a new factor influencing cardiovascular disease (CVD). Although cause-effect relationships are not clearly established, the reported associations between alterations in gut microbiota and CVD are prominent.
Recent studies have found a link between gut microbiota, the microorganisms in human digestive tracts, and, CVD which is the leading cause of mortality worldwide. Gut microbiota is highly variable between individuals, and differences in gut microbial compositions between people with and without CVD have been reported.
Scientists at the University of Toledo (Toledo, OH, USA) hypothesized that machine learning (ML) could be used for gut microbiome–based diagnostic screening of CVD. To test their hypothesis, fecal 16S ribosomal RNA sequencing data of 478 CVD and 473 non-CVD human subjects collected through the American Gut Project were analyzed using five supervised ML algorithms, including random forest, support vector machine, decision tree, elastic net, and neural networks.
The team identified 39 differential bacterial taxa between the CVD and non-CVD groups. ML modeling using these taxonomic features achieved a testing area under the receiver operating characteristic curve (0.0, perfect antidiscrimination; 0.5, random guessing; 1.0, perfect discrimination) of ≈0.58 (random forest and neural networks). Next, the ML models were trained with the top 500 high-variance features of operational taxonomic units, instead of bacterial taxa, and an improved testing area under the receiver operating characteristic curves of ≈0.65 (random forest) was achieved.
Further, by limiting the selection to only the top 25 highly contributing operational taxonomic unit features, the area under the receiver operating characteristic curves was further significantly enhanced to ≈0.70. Among the bacteria identified were Bacteroides, Subdoligranulum, Clostridium, Megasphaera, Eubacterium, Veillonella, Acidaminococcus and Listeria were more abundant in the CVD group. Faecalibacterium, Ruminococcus, Proteus, Lachnospira, Brevundimonas, Alistipes and Neisseria were more abundant in the non-CVD group.
Bina Joe, PhD, FAHA, Distinguished University Professor and Chairwoman of the department of Physiology and Pharmacology, said, “Despite the fact that gut microbiomes are highly variable among individuals, we were surprised by the promising level of accuracy obtained from these preliminary results, which indicate fecal microbiota composition could potentially serve as a convenient diagnostic screening method for CVD.”
The authors concluded that overall, the study was the first to identify dysbiosis of gut microbiota in CVD patients as a group and apply this knowledge to develop a gut microbiome–based ML approach for diagnostic screening of CVD. The study was published on September 10, 2020 in the journal Hypertension.
Related Links:
University of Toledo
Recent studies have found a link between gut microbiota, the microorganisms in human digestive tracts, and, CVD which is the leading cause of mortality worldwide. Gut microbiota is highly variable between individuals, and differences in gut microbial compositions between people with and without CVD have been reported.
Scientists at the University of Toledo (Toledo, OH, USA) hypothesized that machine learning (ML) could be used for gut microbiome–based diagnostic screening of CVD. To test their hypothesis, fecal 16S ribosomal RNA sequencing data of 478 CVD and 473 non-CVD human subjects collected through the American Gut Project were analyzed using five supervised ML algorithms, including random forest, support vector machine, decision tree, elastic net, and neural networks.
The team identified 39 differential bacterial taxa between the CVD and non-CVD groups. ML modeling using these taxonomic features achieved a testing area under the receiver operating characteristic curve (0.0, perfect antidiscrimination; 0.5, random guessing; 1.0, perfect discrimination) of ≈0.58 (random forest and neural networks). Next, the ML models were trained with the top 500 high-variance features of operational taxonomic units, instead of bacterial taxa, and an improved testing area under the receiver operating characteristic curves of ≈0.65 (random forest) was achieved.
Further, by limiting the selection to only the top 25 highly contributing operational taxonomic unit features, the area under the receiver operating characteristic curves was further significantly enhanced to ≈0.70. Among the bacteria identified were Bacteroides, Subdoligranulum, Clostridium, Megasphaera, Eubacterium, Veillonella, Acidaminococcus and Listeria were more abundant in the CVD group. Faecalibacterium, Ruminococcus, Proteus, Lachnospira, Brevundimonas, Alistipes and Neisseria were more abundant in the non-CVD group.
Bina Joe, PhD, FAHA, Distinguished University Professor and Chairwoman of the department of Physiology and Pharmacology, said, “Despite the fact that gut microbiomes are highly variable among individuals, we were surprised by the promising level of accuracy obtained from these preliminary results, which indicate fecal microbiota composition could potentially serve as a convenient diagnostic screening method for CVD.”
The authors concluded that overall, the study was the first to identify dysbiosis of gut microbiota in CVD patients as a group and apply this knowledge to develop a gut microbiome–based ML approach for diagnostic screening of CVD. The study was published on September 10, 2020 in the journal Hypertension.
Related Links:
University of Toledo
Latest Pathology News
- Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
- AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
- FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
- New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
- AI System Analyzes Routine Pathology Slides to Predict Cancer Outcomes
- New Tissue Mapping Approach Identifies High-Risk Form of Diabetic Kidney Disease
- Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
- Interpretable AI Reveals Hidden Cellular Features from Microscopy Images
- Tumor Immune Structure Predicts Response to Immunotherapy in Melanoma
- Plug-and-Play AI Pathology System Classifies Multiple Cancers from Few Slides
- AI-Based Assays Support Risk Stratification in Prostate and Breast Cancer
- AI Pathology Model Predicts Immunotherapy Response in Lung Cancer
- Study Reveals Moleclar Mechanism Driving Aggressive Skin Cancer
- AI Precision Tests Deliver Cancer Risk Insights from Routine H&E Slides
- Collaboration Applies AI Pathology to Predict Response to Antibody-Drug Conjugates
- Biomarker Predicts Immunotherapy Response and Prognosis in Colorectal Cancer
Channels
Clinical Chemistry
view channel
New CA19-9 Cutoff Value Helps Identify High-Risk Pancreatic Cancer Patients
Pancreatic ductal adenocarcinoma (PDAC) is frequently diagnosed at an advanced stage and remains one of the most lethal solid tumors. Clinicians commonly use serum carbohydrate antigen 19-9 (CA19-9) to... Read more
Blood-Based Biomarkers Show Promise for Psychosis Risk Prediction
Psychosis commonly emerges in adolescence or early adulthood and can severely disrupt social and occupational functioning. Hallucinations, delusions, and disorganized thinking often evolve gradually, hindering... Read moreMolecular Diagnostics
view channel
New RNA Origami Method Supports Faster Targeted Testing for Repeat Expansion Disorders
Repeat expansion disorders drive conditions such as myotonic dystrophy, Huntington’s disease, and amyotrophic lateral sclerosis (ALS), yet accurately sizing the mutated sequences remains difficult.... Read more
FDA Approves Expanded Liquid Biopsy Panel for Advanced Cancer Profiling
Timely, comprehensive tumor profiling helps clinicians make treatment selection decisions for patients with advanced cancer. Blood-based approaches can provide actionable insights from a simple draw and... Read moreHematology
view channel
Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
Iron deficiency in school-age children can affect brain development, learning, growth, and physical performance, yet early deficiency may be missed when screening focuses mainly on anemia.... Read more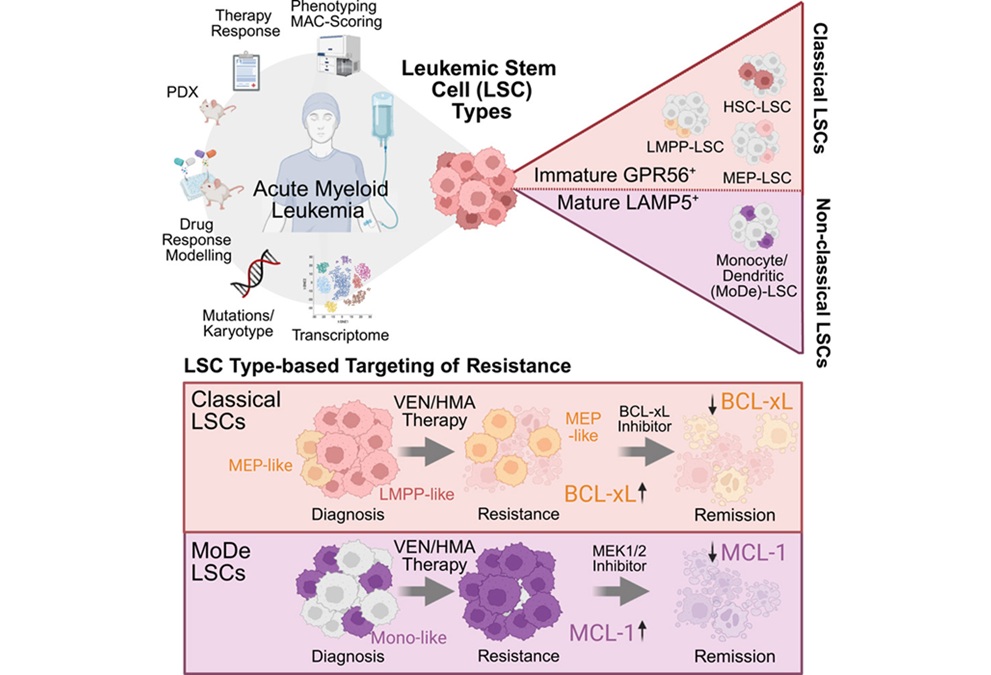
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read moreImmunology
view channel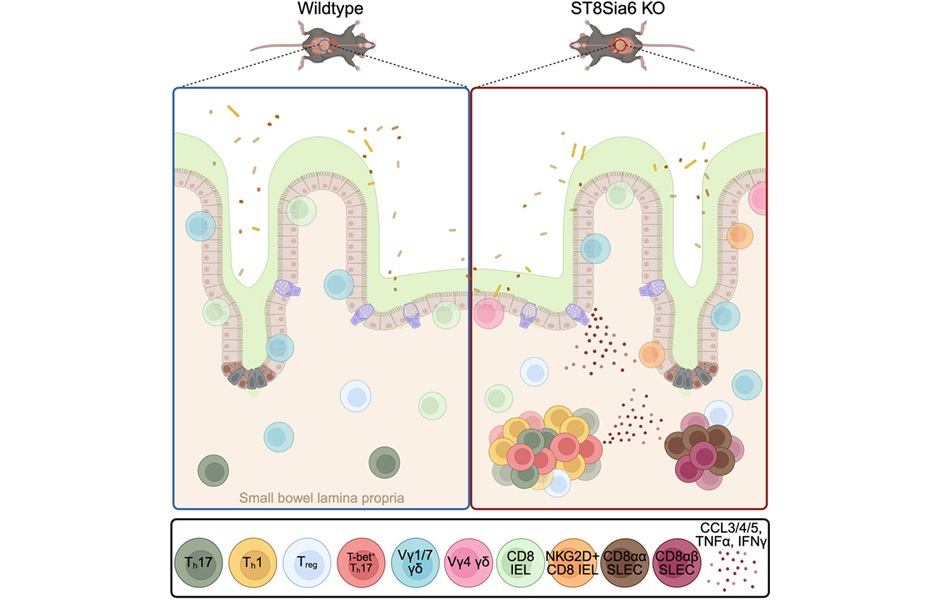
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read more
Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
Lung transplant recipients face some of the highest rates of acute cellular rejection, and routine surveillance often relies on repeated surgical biopsies. These procedures can cause complications such... Read morePathology
view channel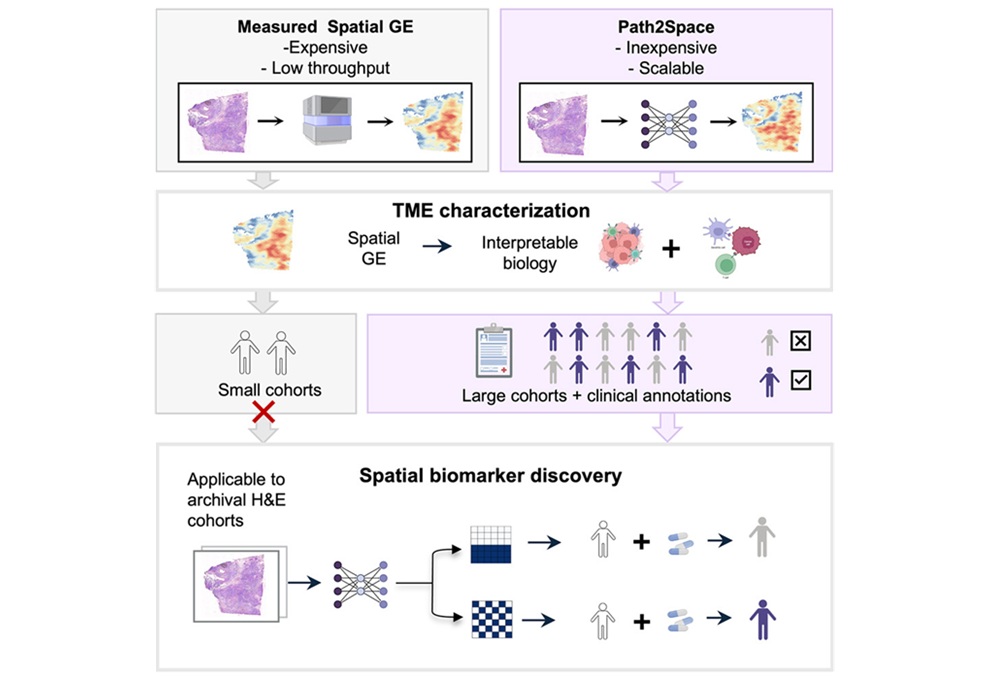
Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
Gene expression profiling can inform tumor biology and treatment selection, but spatial assays remain costly and time-consuming. Results can take weeks and cost thousands of dollars, limiting large-scale... Read more
AI Pathology Test Receives FDA Breakthrough for Bladder Cancer Risk Stratification
Non–muscle invasive bladder cancer has highly variable outcomes, complicating surveillance and treatment planning. Risk assessment typically relies on stage, grade, and tumor size, leaving uncertainty... Read moreTechnology
view channel
AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
Clinical laboratories and research groups face increasingly complex molecular workflows and expanding technical documentation spread across multiple systems. Fragmented digital tools can slow experiment... Read more
AI Tool Automates Validation of Laboratory Software Configuration Changes
Regulated laboratories face heavy documentation and requalification demands when software configurations change, slowing improvements and discouraging beneficial updates. A new capability now automates... Read moreIndustry
view channel
Strategic Collaboration Advances RNA Foundation Models for Precision Oncology
Bulk RNA sequencing is increasingly used to study tumor biology, but standard analyses often reduce results to gene-level summaries that miss important transcript variants and mutation patterns.... Read more