Next-Gen Sequencing Matches Blood Group Antigens for Transfusion
|
By LabMedica International staff writers Posted on 26 Sep 2019 |
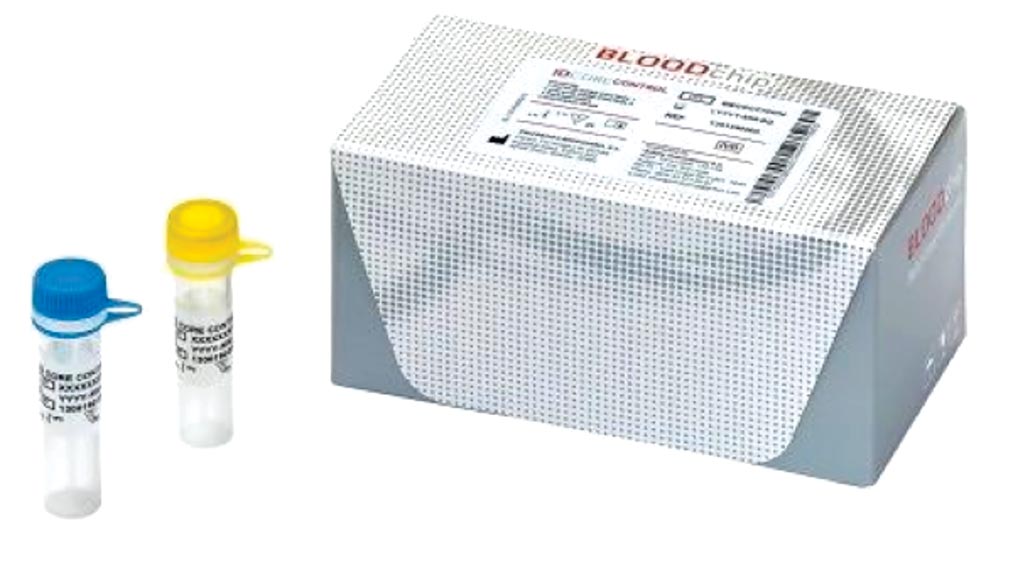

Image: The ID Core XT BLOODchip is a molecular-based assay used in blood transfusion medicine to help determine blood compatibility and could supplement the classical blood match methodology (Photo courtesy of Progenika Biopharma SA).
Transfusion is the procedure of introducing donor material with unknown blood cell antigens into the recipient’s circulatory system. The recipient’s immune system recognizes foreign antigens, produces specific antibodies and sensitization (alloimmunization) occurs.
To date, more than 300 red blood cell (RBC) and 33 human platelet antigens (HPA) have been described. Extended antigen typing is time-consuming, serological methods are costly and depend on the availability of reagents for antigen detection. The procedure is usually performed in reference laboratories, which complicates and delays the delivery of blood for transfusion.
Scientists at the Institute of Hematology and Transfusion Medicine (Warsaw, Poland) have reviewed the advances in applying next-generation sequencing (NGS) to transfusion medicine for the purpose of genotyping alleles encoding clinically important red blood cell and platelet antigens. The currently available technologies allow various levels of sequencing; either the whole genome (WGS), coding regions, exons (WES) or only selected genes or regions of interest. NGS technology significantly reduces the cost of testing. It has been successfully implemented in transplantation medicine for testing donors’ genotypes of HLA antigens in high-throughput mode. Over 9,000 HLA alleles for over 500 individuals can be identified per run.
NGS is particularly effective for finding unknown variations responsible for different phenotypes in patients with antibodies of unknown specificity because it enables screening of the whole genome, exome or particular genes and finding an unknown or rare variant. Recent studies have confirmed NGS effectiveness in resolving the molecular background of orphan antigens with an as yet unknown genetic basis. NGS is also effective in reducing the risk of post-transfusion alloimmunization since the huge capacity of one investigation enables the immediate and cost-effective determination of all RBC and platelet antigen genotypes. Study results support extended profiling of donors and patients for the best prophylactic antigen matching to prevent alloimmunization.
The application of NGS technology for blood typing contributes to the following aspects of patient care: Prevention of alloimmunization in sickle cell disease (SCD) and other transfusion-dependent patients; faster and cheaper diagnostics in the case of patients with unexplained, complex serological results; the huge capacity of the NGS investigations makes this technology an ideal tool for mass screening of blood donors for all clinically important antigens and also to detect individuals with rare blood group antigens in various ethnic groups; this facilitates access to compatible donors for alloimmunised patients.
The authors concluded that the future of NGS as a supplementary test used to provide highly compatible blood as well as to reduce the risk of patient’s alloimmunization and this is part of personalized medicine. The study was published on September 3, 2019, in the journal International Journal of Clinical Transfusion Medicine.
Related Links:
Institute of Hematology and Transfusion Medicine
To date, more than 300 red blood cell (RBC) and 33 human platelet antigens (HPA) have been described. Extended antigen typing is time-consuming, serological methods are costly and depend on the availability of reagents for antigen detection. The procedure is usually performed in reference laboratories, which complicates and delays the delivery of blood for transfusion.
Scientists at the Institute of Hematology and Transfusion Medicine (Warsaw, Poland) have reviewed the advances in applying next-generation sequencing (NGS) to transfusion medicine for the purpose of genotyping alleles encoding clinically important red blood cell and platelet antigens. The currently available technologies allow various levels of sequencing; either the whole genome (WGS), coding regions, exons (WES) or only selected genes or regions of interest. NGS technology significantly reduces the cost of testing. It has been successfully implemented in transplantation medicine for testing donors’ genotypes of HLA antigens in high-throughput mode. Over 9,000 HLA alleles for over 500 individuals can be identified per run.
NGS is particularly effective for finding unknown variations responsible for different phenotypes in patients with antibodies of unknown specificity because it enables screening of the whole genome, exome or particular genes and finding an unknown or rare variant. Recent studies have confirmed NGS effectiveness in resolving the molecular background of orphan antigens with an as yet unknown genetic basis. NGS is also effective in reducing the risk of post-transfusion alloimmunization since the huge capacity of one investigation enables the immediate and cost-effective determination of all RBC and platelet antigen genotypes. Study results support extended profiling of donors and patients for the best prophylactic antigen matching to prevent alloimmunization.
The application of NGS technology for blood typing contributes to the following aspects of patient care: Prevention of alloimmunization in sickle cell disease (SCD) and other transfusion-dependent patients; faster and cheaper diagnostics in the case of patients with unexplained, complex serological results; the huge capacity of the NGS investigations makes this technology an ideal tool for mass screening of blood donors for all clinically important antigens and also to detect individuals with rare blood group antigens in various ethnic groups; this facilitates access to compatible donors for alloimmunised patients.
The authors concluded that the future of NGS as a supplementary test used to provide highly compatible blood as well as to reduce the risk of patient’s alloimmunization and this is part of personalized medicine. The study was published on September 3, 2019, in the journal International Journal of Clinical Transfusion Medicine.
Related Links:
Institute of Hematology and Transfusion Medicine
Latest Hematology News
- Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
- Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
- Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
- Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
- Advanced CBC-Derived Indices Integrated into Hematology Platforms
- Blood Test Enables Early Detection of Multiple Myeloma Relapse
- Single Assay Enables Rapid HLA and ABO Genotyping for Transplant Matching
- Prognostic Biomarker Identified in Diffuse Large B-Cell Lymphoma
- Routine Blood Test Parameters Link Anemia to Cancer Risk and Mortality
- Prognostic Tool Guides Personalized Treatment in Rare Blood Cancer
- New Platelet Function Assay Enables Monitoring of Antiplatelet Therapy
- Open Multi-Omics Platform Identifies Prognostic Subtypes in Blood Cancers
- AI-Powered Digital Workflow Standardizes Bone Marrow Aspirate Morphology
- Rapid Cartridge-Based Test Aims to Expand Access to Hemoglobin Disorder Diagnosis
- New Guidelines Aim to Improve AL Amyloidosis Diagnosis
- Automated Hemostasis System Helps Labs of All Sizes Optimize Workflow
Channels
Clinical Chemistry
view channel
Urinary Biomarker Assay Predicts Kidney Disease Progression Beyond Standard Measures
Many patients with type 2 diabetes and chronic kidney disease continue to experience progressive renal decline, yet conventional markers such as albuminuria and estimated glomerular filtration rate (eGFR)... Read more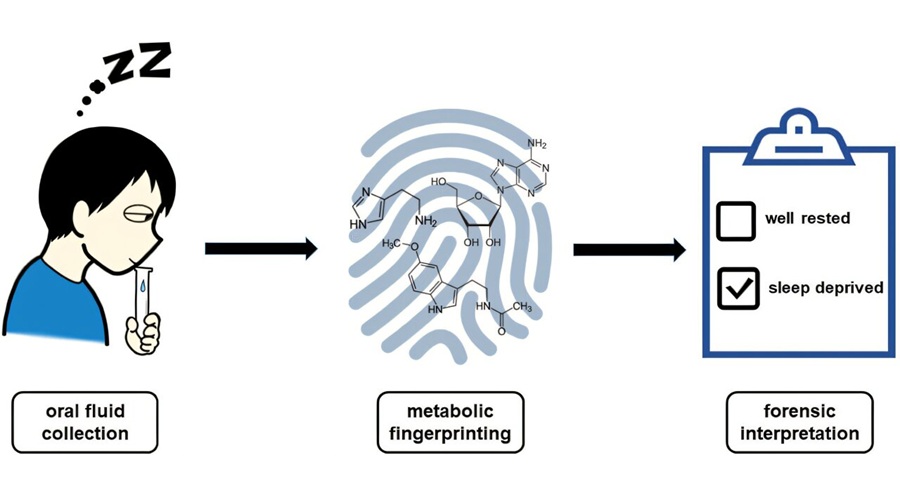
Saliva-Based Test Detects Biochemical Signs of Sleep Loss
Acute sleep loss impairs cognition and motor skills, raising safety risks that resemble alcohol intoxication. Clinicians currently lack an objective biochemical test to determine when someone is dangerously... Read more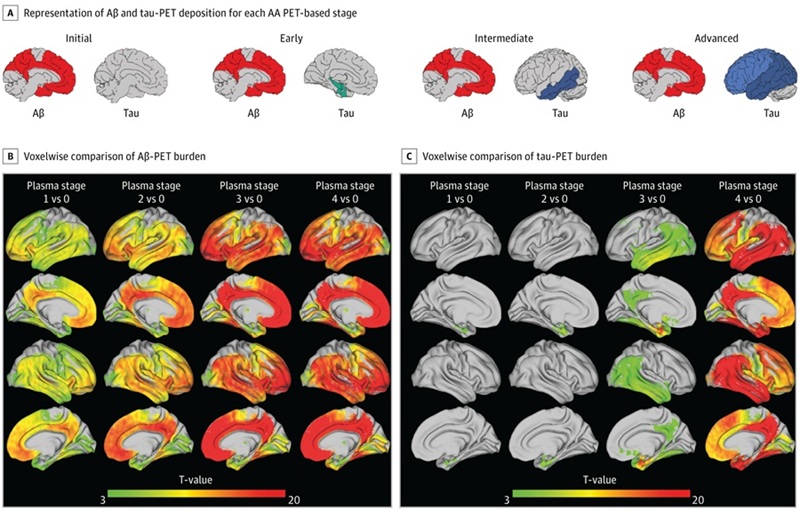
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreMolecular Diagnostics
view channel
Blood-Based RNA Test May Predict Chemotherapy Sensitivity in Lung Cancer
Lung cancer care increasingly relies on biomarker-guided patient stratification, but tissue biopsy can be impractical and treatment selection remains difficult for many patients. Blood-based assays that... Read more
Blood Test Predicts Immunotherapy Response in Head and Neck Cancer
Head and neck squamous cell carcinoma affects hundreds of thousands of people worldwide each year, yet response rates to immunotherapy remain low. Clinicians lack reliable, minimally invasive tools to... Read moreImmunology
view channelAptamer-Based Biosensor Enables Mutation-Resilient SARS-CoV-2 Detection
Rapid evolution of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) can undermine existing molecular diagnostics, especially when assays target small viral components. Double-antibody sandwich... Read more
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more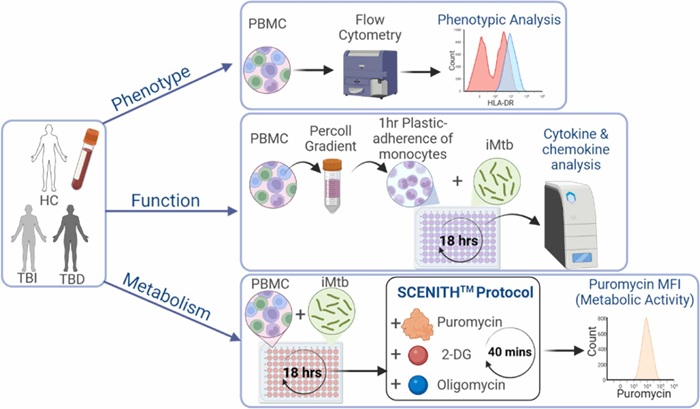
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read moreMicrobiology
view channel
New Culture Medium Speeds C. difficile Resistance Detection and Reduces Costs
Clostridioides difficile infections remain a persistent threat in hospitals and communities, affecting about 500,000 people in the United States each year. Severe cases can be fatal within 30 days of diagnosis,... Read more
Automated Blood Culture System Speeds Detection of Bloodstream Infections
Bloodstream infections and sepsis require rapid laboratory detection to guide targeted antimicrobial therapy and reduce mortality. Conventional blood culture workflows can delay actionable results by critical... Read morePathology
view channel
AI Pathology Tool Predicts Meningioma Recurrence from Routine Slides
Meningiomas are the most common primary brain tumors in adults, yet their course ranges from indolent to highly recurrent disease. Estimating an individual patient’s recurrence risk often requires advanced... Read more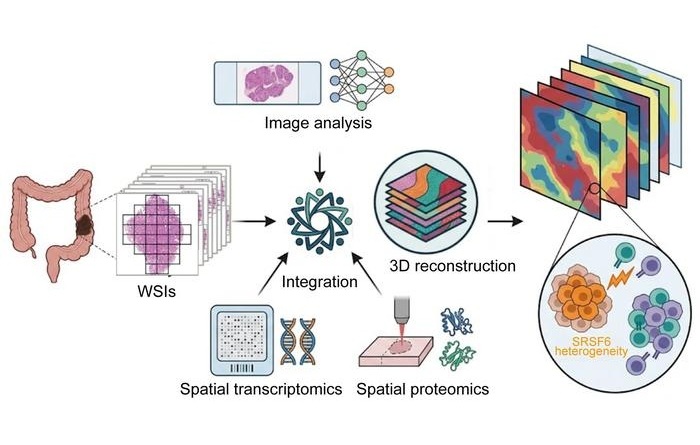
3D Spatial Multi-Omics Maps Intra-Tumor Diversity in Colorectal Cancer
Colorectal cancer remains a leading cause of cancer death, and clinical decision-making is complicated by marked intra-tumor heterogeneity. Conventional bulk sequencing averages molecular signals across... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel
Genetic Testing Program Expands Detection of Alpha-1 Antitrypsin Deficiency
Alpha-1 Antitrypsin Deficiency (AATD) is a progressive genetic condition, the leading known genetic risk factor for chronic obstructive pulmonary disease (COPD), and a cause of liver disease in both children... Read more




.jpg)