Image Recognition Software Increases Accuracy of Malaria Diagnosis
|
By LabMedica International staff writers Posted on 31 Aug 2014 |
Image: The parasite detection method is based on computer vision algorithms similar to those used in facial recognition systems combined with visualization of only the diagnostically most relevant areas. Tablet computers can be utilized in viewing the images (Photo courtesy of the Institute for Molecular Medicine).
A facial recognition software program has been adapted to assist in the identification of the malaria parasite by microscopic examination of blood smears.
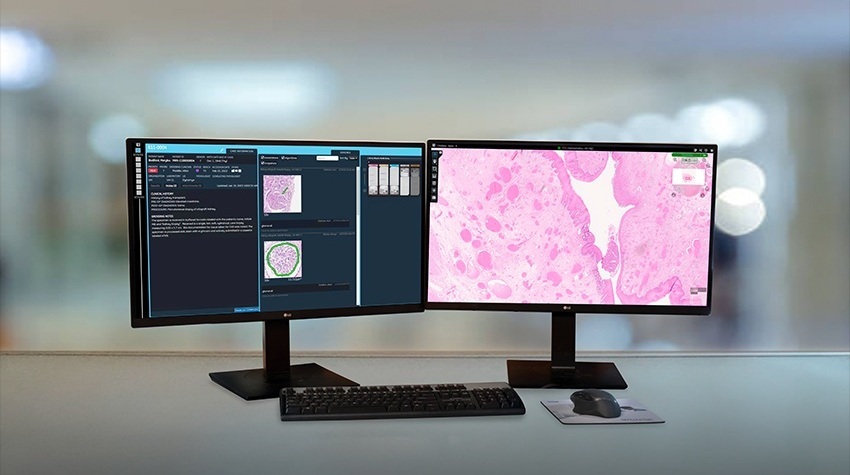
To develop a simpler, more effective visual method to diagnose malaria, a team of Scandinavian researchers coopted computer vision algorithms similar to those used in facial recognition systems. The program operates on a digitalized image of a thin layer of blood that had been smeared on a microscope slide. The algorithm analyzes more than 50,000 red blood cells per sample and ranks them according to likelihood of the cell being infected. The program then generates a panel of images of about a hundred cells most likely to contain the parasite. This panel is then evaluated by an experience microscopist who makes the final diagnosis.
To verify the technique Giemsa-stained thin blood films with Plasmodium falciparum ring-stage trophozoites (n = 27) and uninfected controls (n = 20) were digitally scanned with an oil immersion objective to capture approximately 50,000 erythrocytes per sample. Parasite candidate regions were identified based on color and object size, followed by extraction of image features (local binary patterns, local contrast, and Scale-invariant feature transform descriptors) used as input to a support vector machine classifier. The classifier was trained on digital slides from ten patients and validated on six samples.
From each digitized area of a blood smear, a panel with the 128 most probable parasite candidate regions was generated. Two expert microscopists were asked to visually inspect the panel on a tablet computer and to judge whether the patient was infected with P. falciparum. The method achieved a diagnostic sensitivity and specificity of 95% and 100% as well as 90% and 100% for the two readers respectively using the diagnostic tool. Parasitemia was separately calculated by an automated system and the correlation coefficient between manual and automated parasitemia counts was 0.97.
"We are not suggesting that the whole malaria diagnostic process could or should be automated. Rather, our aim is to develop methods that are significantly less labor intensive than the traditional ones and have a potential to considerably increase the throughput in malaria diagnostics," said senior author Dr. Johan Lundin, research director at the Institute for Molecular Medicine (Helsinki, Finland).
The study with complete description of the new diagnostic approach was published in the August 21, 2014, online edition of the journal PLOS One.
Related Links:
Institute for Molecular Medicine
To develop a simpler, more effective visual method to diagnose malaria, a team of Scandinavian researchers coopted computer vision algorithms similar to those used in facial recognition systems. The program operates on a digitalized image of a thin layer of blood that had been smeared on a microscope slide. The algorithm analyzes more than 50,000 red blood cells per sample and ranks them according to likelihood of the cell being infected. The program then generates a panel of images of about a hundred cells most likely to contain the parasite. This panel is then evaluated by an experience microscopist who makes the final diagnosis.
To verify the technique Giemsa-stained thin blood films with Plasmodium falciparum ring-stage trophozoites (n = 27) and uninfected controls (n = 20) were digitally scanned with an oil immersion objective to capture approximately 50,000 erythrocytes per sample. Parasite candidate regions were identified based on color and object size, followed by extraction of image features (local binary patterns, local contrast, and Scale-invariant feature transform descriptors) used as input to a support vector machine classifier. The classifier was trained on digital slides from ten patients and validated on six samples.
From each digitized area of a blood smear, a panel with the 128 most probable parasite candidate regions was generated. Two expert microscopists were asked to visually inspect the panel on a tablet computer and to judge whether the patient was infected with P. falciparum. The method achieved a diagnostic sensitivity and specificity of 95% and 100% as well as 90% and 100% for the two readers respectively using the diagnostic tool. Parasitemia was separately calculated by an automated system and the correlation coefficient between manual and automated parasitemia counts was 0.97.
"We are not suggesting that the whole malaria diagnostic process could or should be automated. Rather, our aim is to develop methods that are significantly less labor intensive than the traditional ones and have a potential to considerably increase the throughput in malaria diagnostics," said senior author Dr. Johan Lundin, research director at the Institute for Molecular Medicine (Helsinki, Finland).
The study with complete description of the new diagnostic approach was published in the August 21, 2014, online edition of the journal PLOS One.
Related Links:
Institute for Molecular Medicine
Latest Microbiology News
- FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
- New AMR Assay Supports Rapid Infection Control Screening in Hospitals
- Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- Rapid Color Test Stratifies Virulent and Resistant Staph Strains
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Syndromic Panel Enables Rapid Identification of Bloodstream Infections
- RNA-Based Workflow Identifies Active Skin Microbes for Dermatology Research
Channels
Clinical Chemistry
view channel
Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
Childhood epilepsy remains a major neurological disorder with unmet needs for accurate, non-invasive biomarkers, as conventional tests such as electroencephalography and neuroimaging can have limited sensitivity... Read more
Blood-Based Sensor Detects Early Signs of Alzheimer’s and Parkinson’s
Alzheimer’s disease and Parkinson’s disease are increasing as populations age, yet diagnosis remains largely symptom-driven and often occurs after irreversible brain damage has begun. Earlier detection,... Read moreMolecular Diagnostics
view channel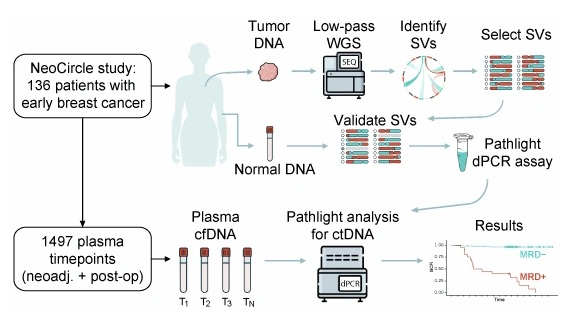
Ultrasensitive MRD Blood Test Detects Early Breast Cancer Recurrence
SAGA Diagnostics (Morrisville, NC, USA), a company specializing in tumor-informed, blood-based cancer detection and precision medicine, announced the publication of a new study evaluating its Pathlight... Read more
Position Statements Outline Evidence Standards for Multi-Cancer Detection Tests
Cancer screening is intended to reduce mortality, but policy decisions often depend on early indicators that may not fully reflect true survival benefit. The emergence of blood-based tests capable of detecting... Read moreHematology
view channel
Higher Ferritin Threshold May Improve Iron Deficiency Detection in Children
Iron deficiency in school-age children can affect brain development, learning, growth, and physical performance, yet early deficiency may be missed when screening focuses mainly on anemia.... Read more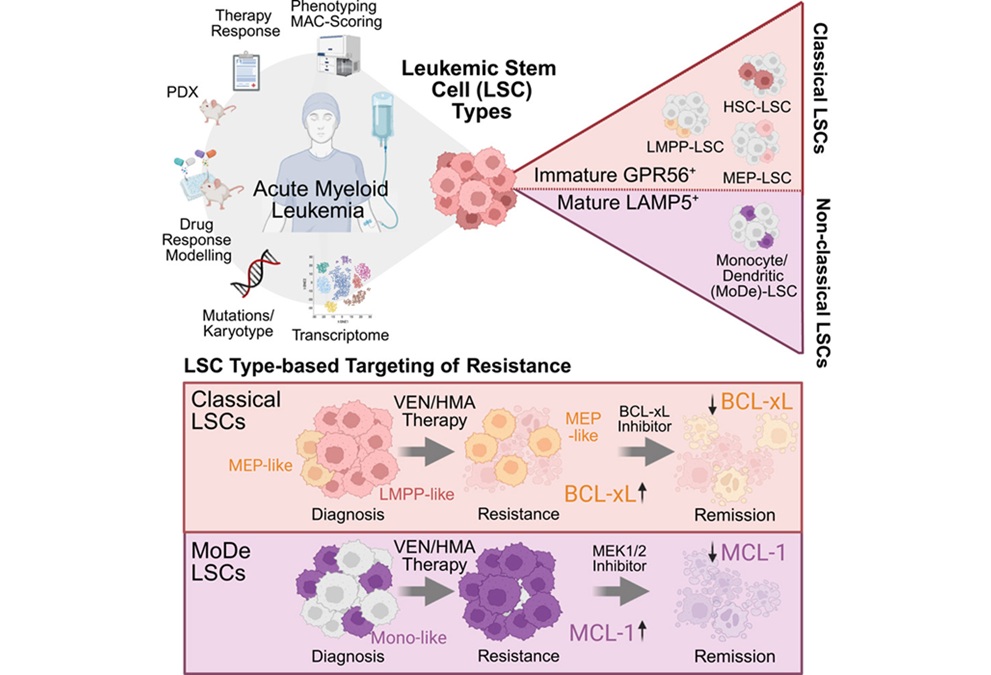
Stem Cell Biomarkers May Guide Precision Treatment in Acute Myeloid Leukemia
Acute myeloid leukemia (AML) is an aggressive blood cancer that most often affects older adults and still carries a poor prognosis despite therapeutic advances. Venetoclax-based regimens have improved... Read moreImmunology
view channel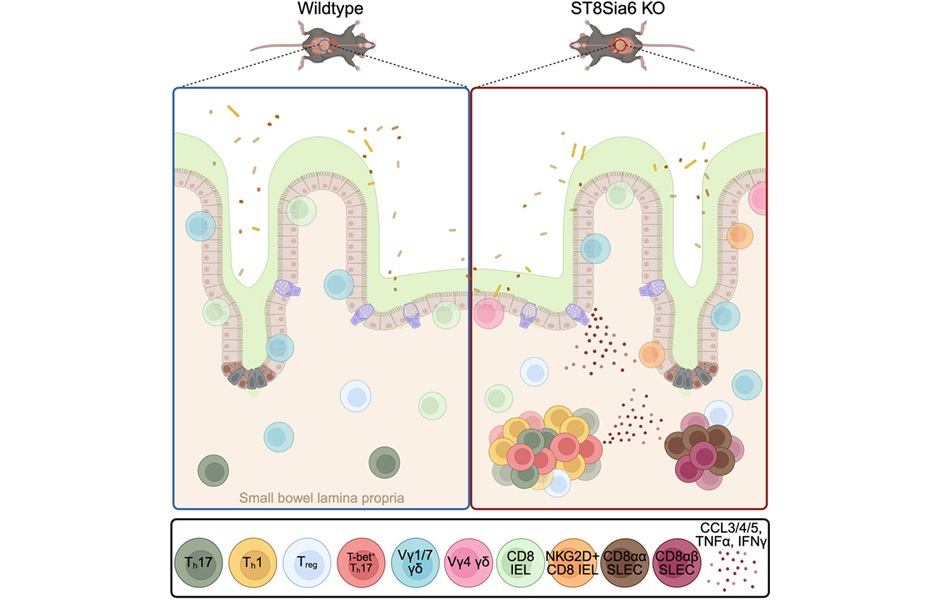
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read more
Simple Blood Test Could Replace Biopsies for Lung Transplant Rejection Monitoring
Lung transplant recipients face some of the highest rates of acute cellular rejection, and routine surveillance often relies on repeated surgical biopsies. These procedures can cause complications such... Read morePathology
view channel
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read more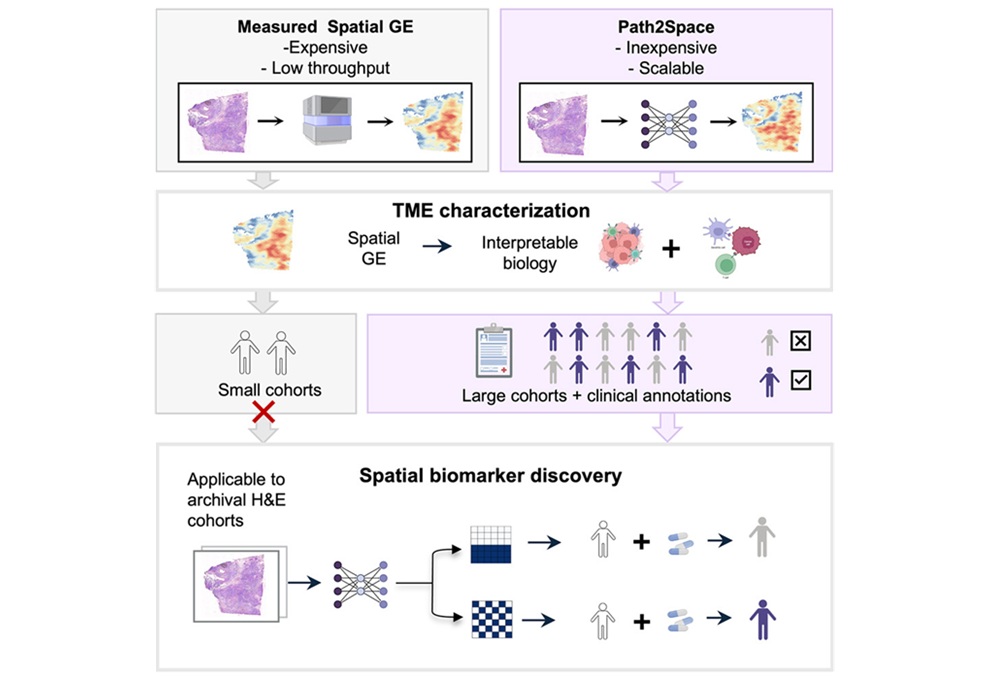
Rapid AI Tool Predicts Cancer Spatial Gene Expression from Pathology Images
Gene expression profiling can inform tumor biology and treatment selection, but spatial assays remain costly and time-consuming. Results can take weeks and cost thousands of dollars, limiting large-scale... Read moreTechnology
view channel
AI-Enabled Assistant Unifies Molecular Workflow Planning and Support
Clinical laboratories and research groups face increasingly complex molecular workflows and expanding technical documentation spread across multiple systems. Fragmented digital tools can slow experiment... Read more
AI Tool Automates Validation of Laboratory Software Configuration Changes
Regulated laboratories face heavy documentation and requalification demands when software configurations change, slowing improvements and discouraging beneficial updates. A new capability now automates... Read moreIndustry
view channel
New Distribution Agreement Expands Access to CE-Marked Precision Oncology Assays
Eurobio Scientific (Les Ulis, France) has signed a distribution agreement with Canhelp Genomics (Hangzhou, China) to broaden availability of the Canhelp‑UCa and Canhelp‑Origin assays. The agreement extends... Read more