Genotyping Performed by FRET-PCR Without DNA Extraction
|
By LabMedica International staff writers Posted on 07 Jul 2014 |

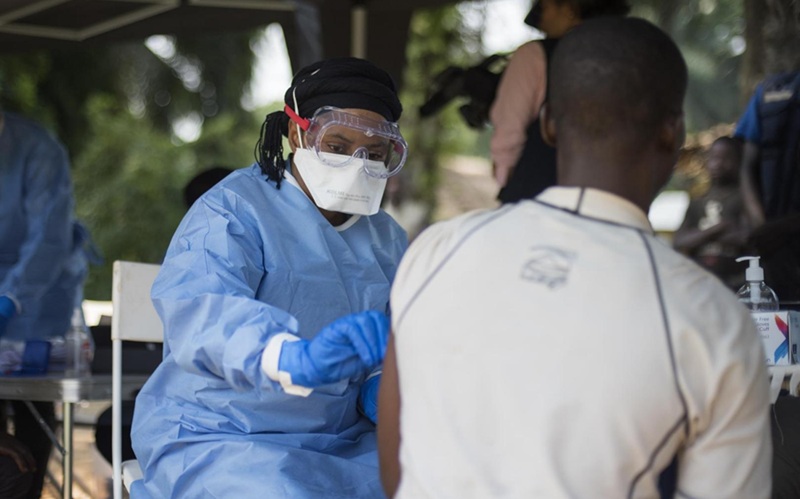
Image: The LightCycler 2.0 real-time polymerase chain reaction analyzer and centrifuge (Photo courtesy of Roche).”
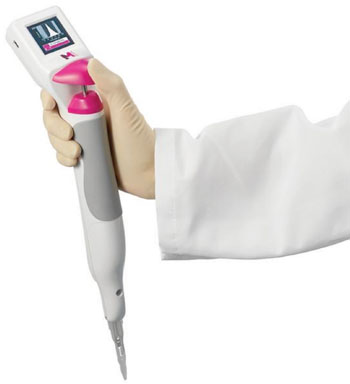
Image: The Scepter 2.0 Automated Cell Counter (Photo courtesy of Merck Millipore).
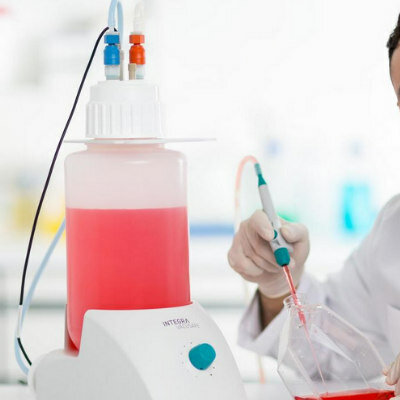
Blood samples are extensively used for the molecular diagnosis of many hematological diseases using a variety of techniques, based on the amplification of nucleic acids.
Current methods for polymerase chain reaction (PCR) use purified genomic DNA, mostly isolated from total peripheral blood cells or white blood cells (WBC), which can be improved by a real-time fluorescence resonance energy transfer-based method for genotyping directly from blood cells.
Hematologists at the Hospital Universitari Son Espases (Palma de Mallorca, Spain) studied peripheral blood from 34 patients collected into tubes containing ethylenediaminetetraacetic acid (EDTA). Among the samples, they included a mixture of mutant alleles for patients suffering from thrombosis or hereditary hemochromatosis. Red blood cells (RBCs) were lysed and white blood cells (WBCs) isolated. A real-time PCR was then performed followed by a melting curve analysis for different genes including methylenetetrahydrofolate reductase (MTHFR), hemochromatosis (HFE), coagulation factor V Leiden (F5), prothrombin factor two (F2) and coagulation factor XII (F12).
The real time PCR was performed on the LightCycler 2.0 Instrument (Roche Diagnostics Corporation, Indianapolis, IN, USA). In order to standardize the samples for the real-time PCR reaction, cells were counted in a Scepter 2.0 Automated Cell Counter (Merck Millipore, Billerica, MA, USA) and adjusted to 5×106 cells/mL. After testing 34 samples comparing the real-time crossing point (CP) values between 5×106 WBC/mL and 20 ng/µL of purified DNA, the results for F5 Leiden were as follows: CP mean value for WBC was 29.26 ± 0.57 versus purified DNA 24.79 ± 0.56. There was an observed delay of about four cycles when PCR was performed from WBC instead of DNA.
The authors concluded that their protocol obviates the DNA purification stage, thereby saving time and resources. Furthermore, since the manipulation performed on the sample is minimal, it may decrease the risk of contamination. As they reported the results from a variety of genes, they contend that their protocol will be suitable for the genotyping of almost any inherited polymorphism. The study was published on June 25, 2014, in the Journal of Blood Medicine.
Related Links:
Hospital Universitari Son Espases
Roche Diagnostics Corporation
Merck Millipore
Current methods for polymerase chain reaction (PCR) use purified genomic DNA, mostly isolated from total peripheral blood cells or white blood cells (WBC), which can be improved by a real-time fluorescence resonance energy transfer-based method for genotyping directly from blood cells.
Hematologists at the Hospital Universitari Son Espases (Palma de Mallorca, Spain) studied peripheral blood from 34 patients collected into tubes containing ethylenediaminetetraacetic acid (EDTA). Among the samples, they included a mixture of mutant alleles for patients suffering from thrombosis or hereditary hemochromatosis. Red blood cells (RBCs) were lysed and white blood cells (WBCs) isolated. A real-time PCR was then performed followed by a melting curve analysis for different genes including methylenetetrahydrofolate reductase (MTHFR), hemochromatosis (HFE), coagulation factor V Leiden (F5), prothrombin factor two (F2) and coagulation factor XII (F12).
The real time PCR was performed on the LightCycler 2.0 Instrument (Roche Diagnostics Corporation, Indianapolis, IN, USA). In order to standardize the samples for the real-time PCR reaction, cells were counted in a Scepter 2.0 Automated Cell Counter (Merck Millipore, Billerica, MA, USA) and adjusted to 5×106 cells/mL. After testing 34 samples comparing the real-time crossing point (CP) values between 5×106 WBC/mL and 20 ng/µL of purified DNA, the results for F5 Leiden were as follows: CP mean value for WBC was 29.26 ± 0.57 versus purified DNA 24.79 ± 0.56. There was an observed delay of about four cycles when PCR was performed from WBC instead of DNA.
The authors concluded that their protocol obviates the DNA purification stage, thereby saving time and resources. Furthermore, since the manipulation performed on the sample is minimal, it may decrease the risk of contamination. As they reported the results from a variety of genes, they contend that their protocol will be suitable for the genotyping of almost any inherited polymorphism. The study was published on June 25, 2014, in the Journal of Blood Medicine.
Related Links:
Hospital Universitari Son Espases
Roche Diagnostics Corporation
Merck Millipore
Latest Molecular Diagnostics News
- Digital PCR Assays Support Surveillance of Bundibugyo Ebolavirus Outbreak
- Updated Guidance Prioritizes Stool-Based Colorectal Cancer Screening Tests
- Blood-Based Proteomic Test May Predict Treatment Response in Non-Small Cell Lung Cancer
- Position Statements Outline Evidence Standards for Multi-Cancer Detection Tests
- Ultrasensitive MRD Blood Test Detects Early Breast Cancer Recurrence
- Gene Fusion Patterns May Flag High Risk Solitary Fibrous Tumors
- New RNA Origami Method Supports Faster Targeted Testing for Repeat Expansion Disorders
- FDA Approves Expanded Liquid Biopsy Panel for Advanced Cancer Profiling
- Microbial Saliva Test Could Help Triage Esophageal Cancer Risk
- Expanded DPYD Genotyping Test Supports Safer Chemotherapy Dosing
- Blood Test Detects Early Nonresponse in Metastatic Prostate Cancer
- Multi-Omics Profiling Helps Predict BCG Response and Recurrence in Bladder Cancer
- New Computational Tool Reveals Genetic Driver of Idiopathic Neuropathy
- Breast Cancer-Specific Signatures Link Genome Instability to Outcomes
- FDA-Cleared Genomic Profiling Assay Guides Treatment Selection in Solid Tumors
- ctDNA Blood Test Could Help Guide Radiotherapy in Patients with Limited Metastases
Channels
Clinical Chemistry
view channel
Urine-Based Test Shows Promise for Autism Screening in Children
Autism spectrum disorder (ASD) is commonly diagnosed through behavioral assessments, which can involve long waits that delay intervention. Earlier identification is linked to better developmental outcomes,... Read more
Liquid Biopsy Biomarkers May Improve Childhood Epilepsy Diagnosis
Childhood epilepsy remains a major neurological disorder with unmet needs for accurate, non-invasive biomarkers, as conventional tests such as electroencephalography and neuroimaging can have limited sensitivity... Read moreMolecular Diagnostics
view channel
Updated Guidance Prioritizes Stool-Based Colorectal Cancer Screening Tests
Colorectal cancer is the second-leading cause of cancer death in the United States and claimed an estimated 55,000 lives in 2026. Incidence is rising among adults younger than 50, even as overall mortality... Read more
Digital PCR Assays Support Surveillance of Bundibugyo Ebolavirus Outbreak
QIAGEN (Venlo, Netherlands) has introduced two custom-designed research-use-only (RUO) QIAcuity dPCR assays to support infectious disease research and surveillance connected to the Bundibugyo ebolavirus outbreak.... Read more
Blood-Based Proteomic Test May Predict Treatment Response in Non-Small Cell Lung Cancer
Lung cancer remains the leading cause of cancer death, with non-small cell lung cancer (NSCLC) accounting for most cases. Treatment decisions are often made without a clear indication of how a patient... Read moreImmunology
view channel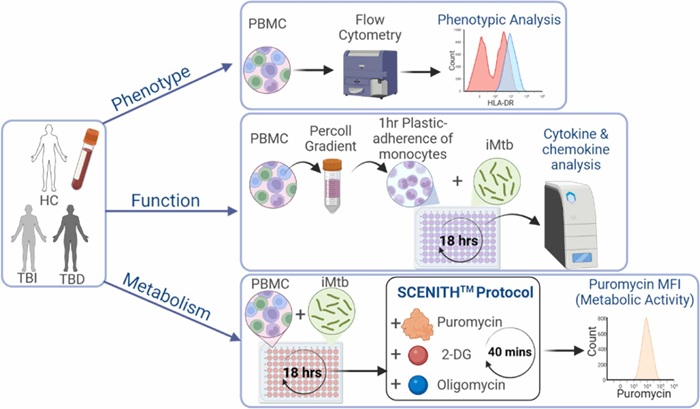
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read more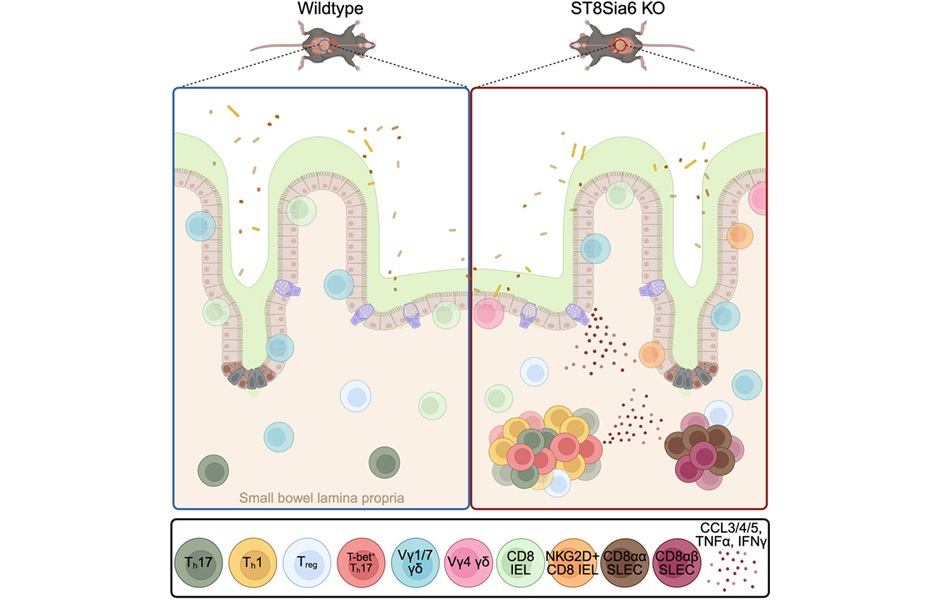
Immune Enzyme Linked to Treatment-Resistant Inflammatory Bowel Disease
Inflammatory bowel disease (IBD) affects nearly 3 million people in the United States and its prevalence continues to rise. Medications that target tumor necrosis factor (TNF)-alpha are widely used, but... Read moreMicrobiology
view channel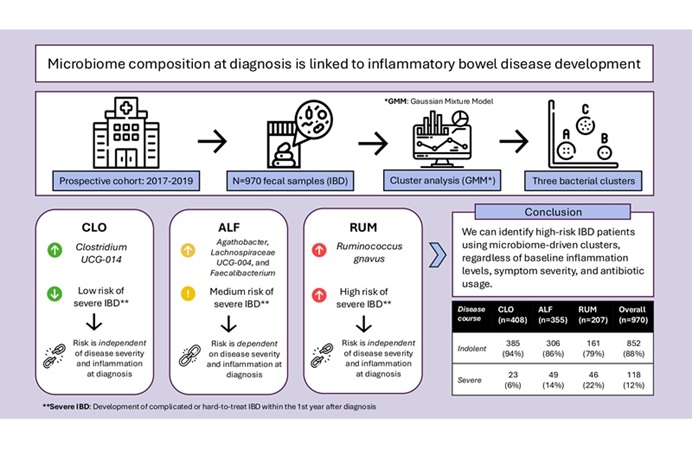
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more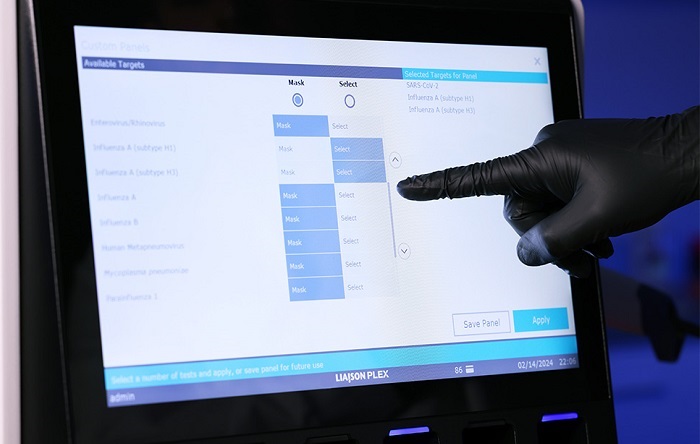
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel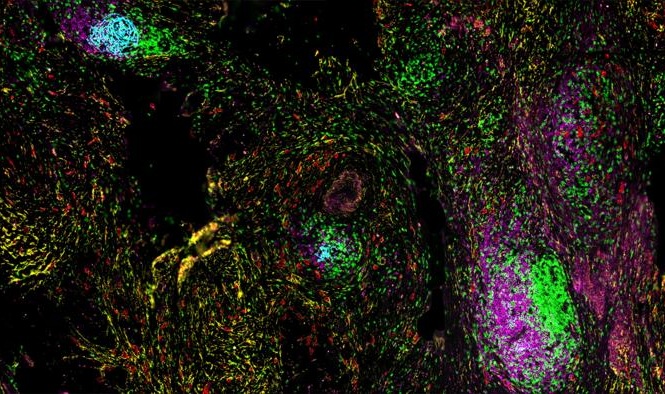
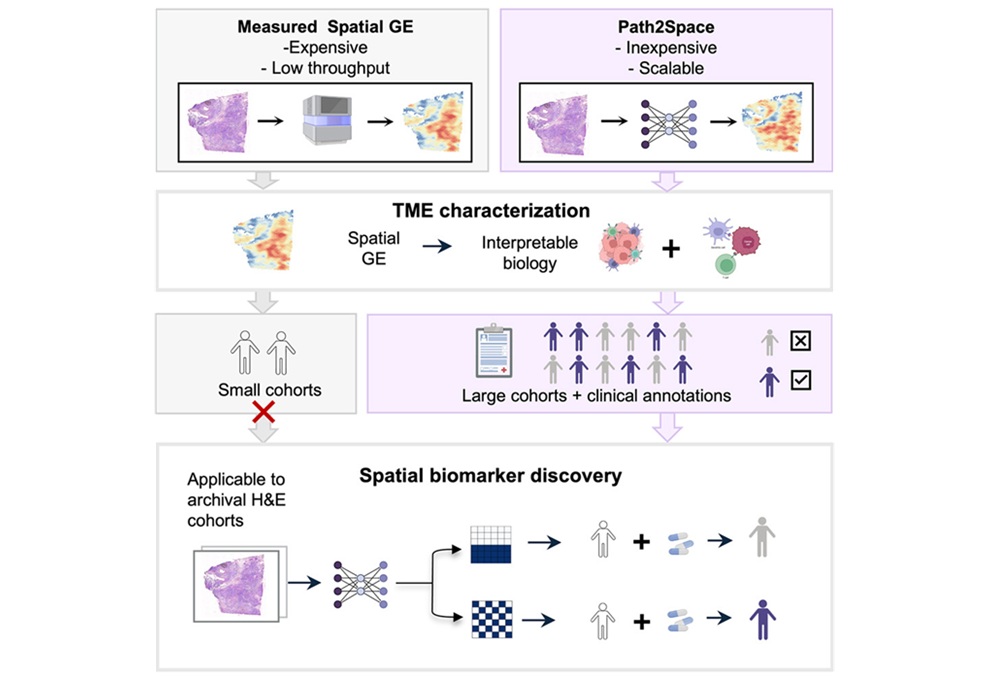
AI-Powered Atlas Maps Immune Structures Linked to Cancer Outcomes
Tertiary lymphoid structures are emerging as important indicators of antitumor immunity, but their heterogeneity and spatial context within tumors remain difficult to capture through routine diagnostics.... Read more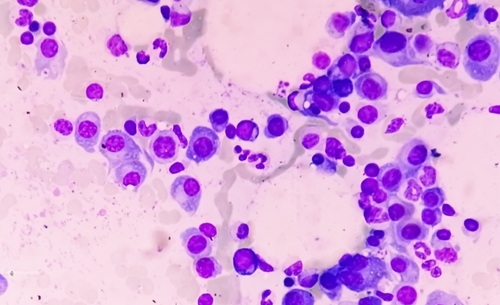
AI Tool Extracts Immune Signals from Biopsy to Inform Myeloma Therapy
Multiple myeloma is a bone marrow malignancy in which patients can respond very differently to the same treatments, making initial therapy decisions difficult. Clinicians must choose among options such... Read moreTechnology
view channel
Mailed Screening Kits Help Reduce Colorectal Cancer Screening Gaps
Colorectal cancer screening is a longstanding preventive priority, yet participation and follow-up remain uneven across patient groups. Safety‑net primary care settings often face barriers that limit screening... Read more
Algorithm Panel Aids Liver Fibrosis Assessment and Liver Cancer Surveillance
Chronic liver disease is common and often progresses silently, increasing the risk of cirrhosis and hepatocellular carcinoma when not detected early. With an estimated 1.5 billion people affected worldwide... Read moreIndustry
view channelWerfen and Oxford Nanopore Collaborate on Transplant Assay Development
Werfen (Barcelona, Spain), a global specialized diagnostics company, has announced a strategic collaboration with Oxford Nanopore Technologies (Oxford, UK), which develops nanopore-based sequencing technology,... Read more