Alzheimer's Disease Subtypes Proposed from Brain Gene Expression Profiles
|
By LabMedica International staff writers Posted on 18 Jan 2021 |
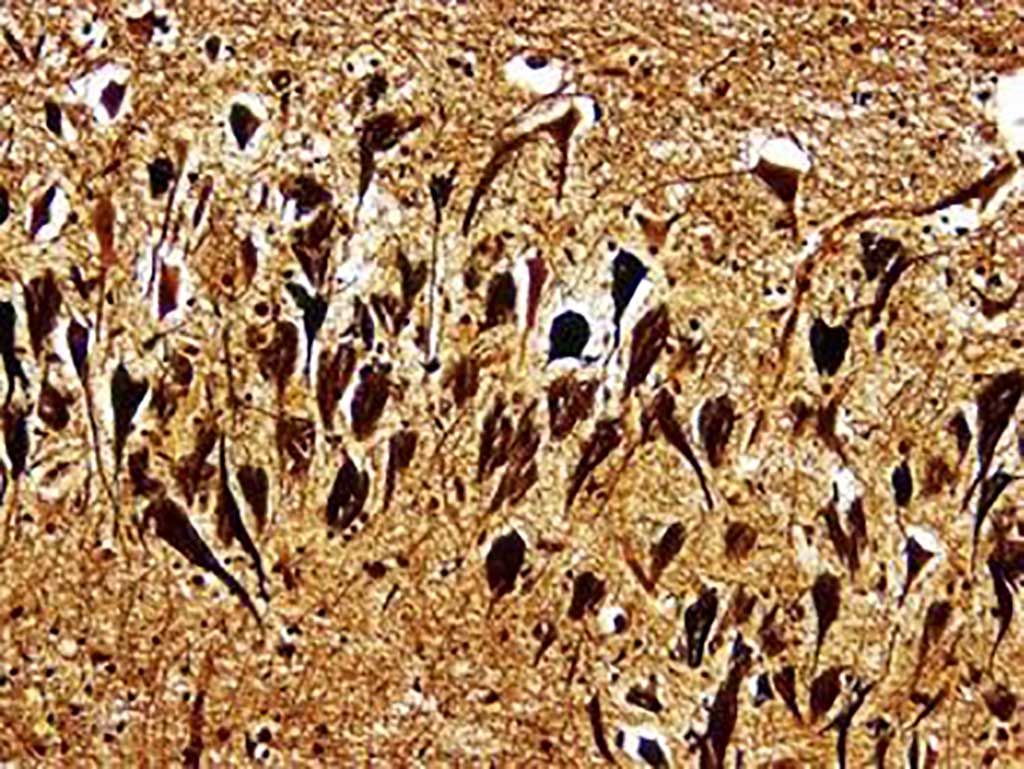
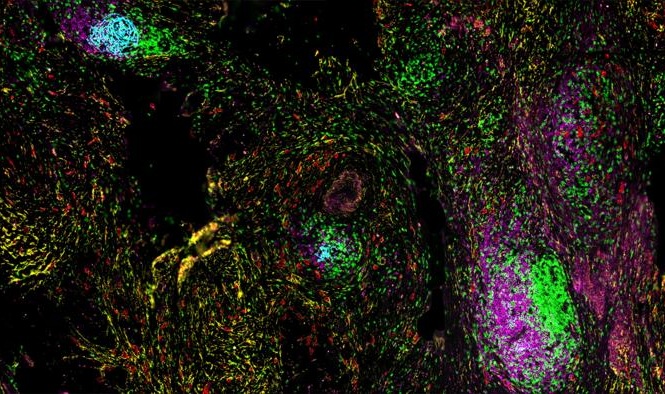
Image: Histopathology of neurofibrillary tangles in the brain of a patient with Alzheimer`s disease (Bielschowski silver stain) (Photo courtesy of Dimitri P. Agamanolis, MD).
Alzheimer’s disease (AD) is the most common form of dementia in the elderly, estimated to affect more than 5.8 million individuals in the USA and more than 50 million worldwide, with almost half of individuals aged over 75 years.
The neuropathological manifestations of AD traditionally include the accumulation of amyloid-beta (Aβ) peptide as extracellular plaques and hyperphosphorylated tau as intracellular neurofibrillary tangles (NFTs), typically identified on postmortem biopsy and used for definitive AD diagnosis.
A large team of scientists led by those at Icahn School of Medicine (New York, NY, USA) used transcriptome sequence data from more than 1,500 postmortem brain samples from individuals with or without AD to highlight several expression-based AD subtypes. They analyzed transcriptome data for more than 900 samples from the frontal pole (FP), superior temporal gyrus (STG), parahippocampal gyrus (PHG), and inferior frontal gyrus (IFG) brain regions in 364 Mount Sinai/JJ Peters VA Medical Center Brain Bank (MSBB-AD) participants with or without AD or related dementia.
The scientists focused in on differential gene expression patterns in the PHG, adjusting for AD stage and severity. Their results pointed to five PHG expression-based subtypes of AD, falling into three main clusters, along with related molecular signatures, clinical features, and potential driver genes. The team identified three major molecular subtypes of AD corresponding to different combinations of multiple dysregulated pathways, such as susceptibility to tau-mediated neurodegeneration, amyloid-β neuroinflammation, synaptic signaling, immune activity, mitochondria organization, and myelination. Multiscale network analysis reveals subtype-specific drivers such as GABRB2, LRP10, MSN, PLP1, and ATP6V1A. The team reported their results were shored up with data for postmortem brain samples from another 615 AD cases or controls in Religious Orders Study–Memory and Aging Project (ROSMAP).
Bin Zhang, PhD, a Professor of Genetics and genomic Science and senior author of the study, said, “Understanding the genetic and molecular differences between molecular subtypes of AD within these data will provide novel insights into disease pathogenesis and offer new avenues for developing effective therapeutics.” The study was published on January 6, 2021 in the journal Science Advances.
Related Links:
Icahn School of Medicine
The neuropathological manifestations of AD traditionally include the accumulation of amyloid-beta (Aβ) peptide as extracellular plaques and hyperphosphorylated tau as intracellular neurofibrillary tangles (NFTs), typically identified on postmortem biopsy and used for definitive AD diagnosis.
A large team of scientists led by those at Icahn School of Medicine (New York, NY, USA) used transcriptome sequence data from more than 1,500 postmortem brain samples from individuals with or without AD to highlight several expression-based AD subtypes. They analyzed transcriptome data for more than 900 samples from the frontal pole (FP), superior temporal gyrus (STG), parahippocampal gyrus (PHG), and inferior frontal gyrus (IFG) brain regions in 364 Mount Sinai/JJ Peters VA Medical Center Brain Bank (MSBB-AD) participants with or without AD or related dementia.
The scientists focused in on differential gene expression patterns in the PHG, adjusting for AD stage and severity. Their results pointed to five PHG expression-based subtypes of AD, falling into three main clusters, along with related molecular signatures, clinical features, and potential driver genes. The team identified three major molecular subtypes of AD corresponding to different combinations of multiple dysregulated pathways, such as susceptibility to tau-mediated neurodegeneration, amyloid-β neuroinflammation, synaptic signaling, immune activity, mitochondria organization, and myelination. Multiscale network analysis reveals subtype-specific drivers such as GABRB2, LRP10, MSN, PLP1, and ATP6V1A. The team reported their results were shored up with data for postmortem brain samples from another 615 AD cases or controls in Religious Orders Study–Memory and Aging Project (ROSMAP).
Bin Zhang, PhD, a Professor of Genetics and genomic Science and senior author of the study, said, “Understanding the genetic and molecular differences between molecular subtypes of AD within these data will provide novel insights into disease pathogenesis and offer new avenues for developing effective therapeutics.” The study was published on January 6, 2021 in the journal Science Advances.
Related Links:
Icahn School of Medicine
Latest Molecular Diagnostics News
- Plasma Protein Signature Predicts Lung Cancer Risk Up to Five Years Ahead
- Circulating Tumor DNA Testing Guides Chemotherapy, Reduces Relapse in Colon Cancer
- Researchers Uncover Distinct Chromosome Signature in Aggresive ALT Cancers
- Simple Cytogenetic Method Could Improve Classification of ALL Subtypes
- Blood-Based Assay Enables Noninvasive Monitoring of Sarcoma Immunotherapy Response
- Genomic Test Guides Chemotherapy Decisions in Early-Stage Breast Cancer
- Tumor Mutation Marker Helps Refine Lung Cancer Prognosis and Guide Therapy Selection
- Multi-Cancer Test Boosts Detection When Added to Standard Screening
- Blood-Based MRD Monitoring Supports Relapse Prevention in Leukemia
- Genomic Test Predicts Chemotherapy Benefit in Metastatic Prostate Cancer
- Blood Protein Markers Flag Multiple Sclerosis Risk Years Before Diagnosis
- Digital PCR Assays Support Surveillance of Bundibugyo Ebolavirus Outbreak
- Updated Guidance Prioritizes Stool-Based Colorectal Cancer Screening Tests
- Blood-Based Proteomic Test May Predict Treatment Response in Non-Small Cell Lung Cancer
- Position Statements Outline Evidence Standards for Multi-Cancer Detection Tests
- Ultrasensitive MRD Blood Test Detects Early Breast Cancer Recurrence
Channels
Clinical Chemistry
view channel
Saliva-Based Test Detects Biochemical Signs of Sleep Loss
Acute sleep loss impairs cognition and motor skills, raising safety risks that resemble alcohol intoxication. Clinicians currently lack an objective biochemical test to determine when someone is dangerously... Read more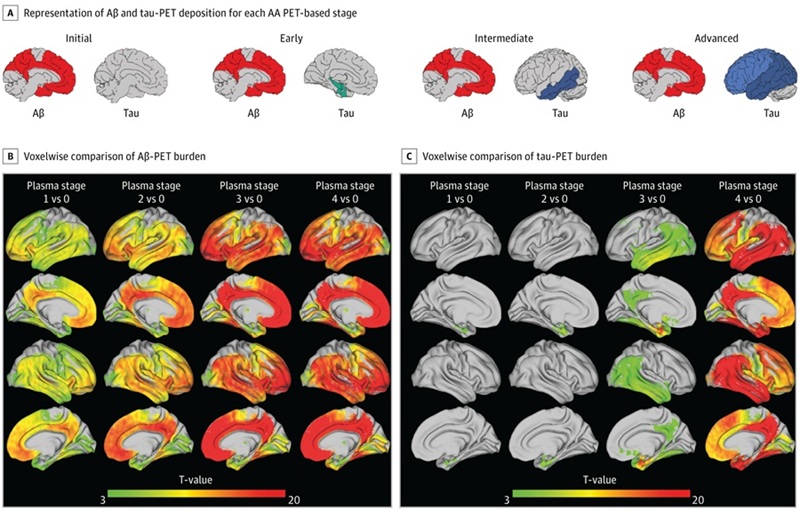
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more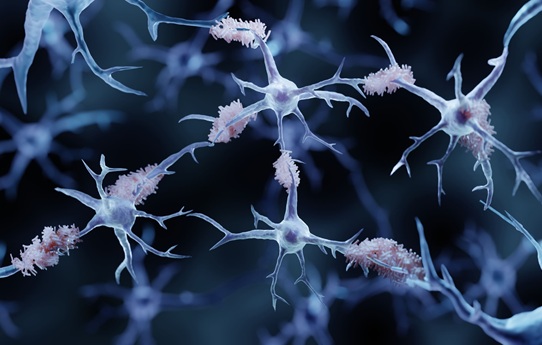
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more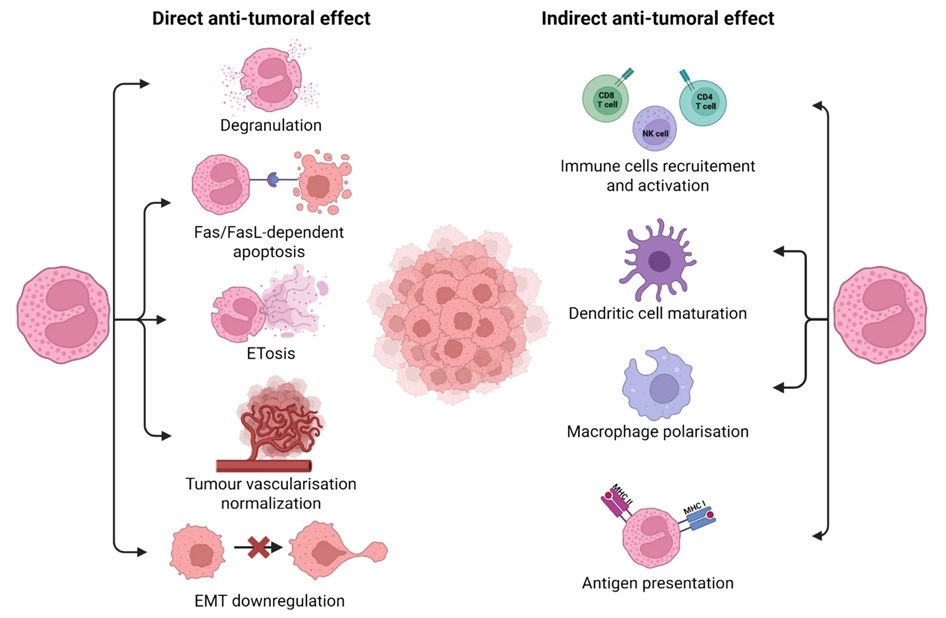
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channelAptamer-Based Biosensor Enables Mutation-Resilient SARS-CoV-2 Detection
Rapid evolution of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) can undermine existing molecular diagnostics, especially when assays target small viral components. Double-antibody sandwich... Read more
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more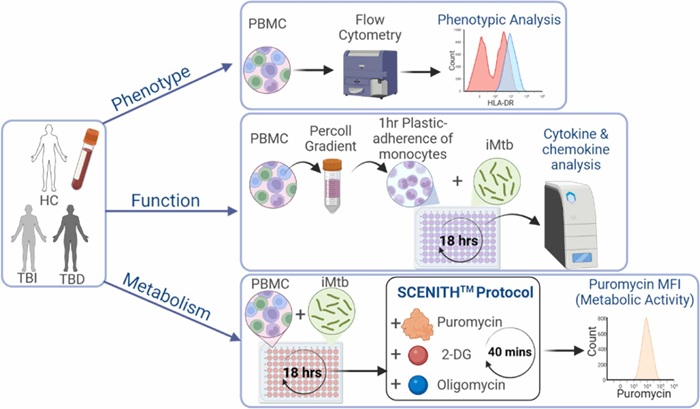
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read moreMicrobiology
view channel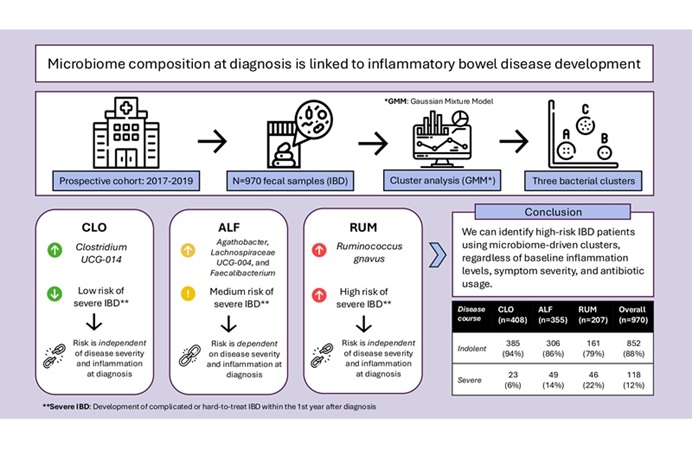
Gut Microbiome Signatures Help Identify Risk of IBD Progression
Inflammatory bowel disease (IBD), encompassing Crohn’s disease and ulcerative colitis, is a chronic relapsing inflammatory disorder of the gastrointestinal tract with highly variable outcomes.... Read more
FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
Clinical guidelines support testing based on patient presentation in suspected gastrointestinal infections, yet available technologies have often forced laboratories to choose between panels that are too... Read morePathology
view channel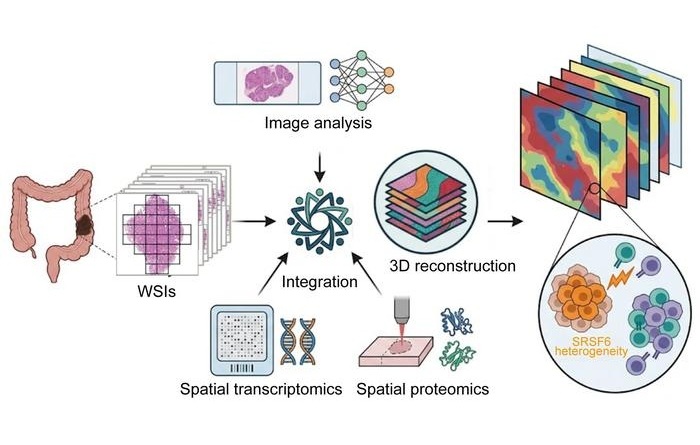
3D Spatial Multi-Omics Maps Intra-Tumor Diversity in Colorectal Cancer
Colorectal cancer remains a leading cause of cancer death, and clinical decision-making is complicated by marked intra-tumor heterogeneity. Conventional bulk sequencing averages molecular signals across... Read more
Blood-Based Method Tracks Gene Activity in the Living Brain
Real-time measurement of gene activity in the brain has been limited by assays requiring destructive tissue sampling. Tracking active genes could reveal how the body responds to environmental factors,... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel