Genomics Technique Accelerate Detection of Foodborne Bacterial Outbreaks
|
By LabMedica International staff writers Posted on 15 Dec 2016 |
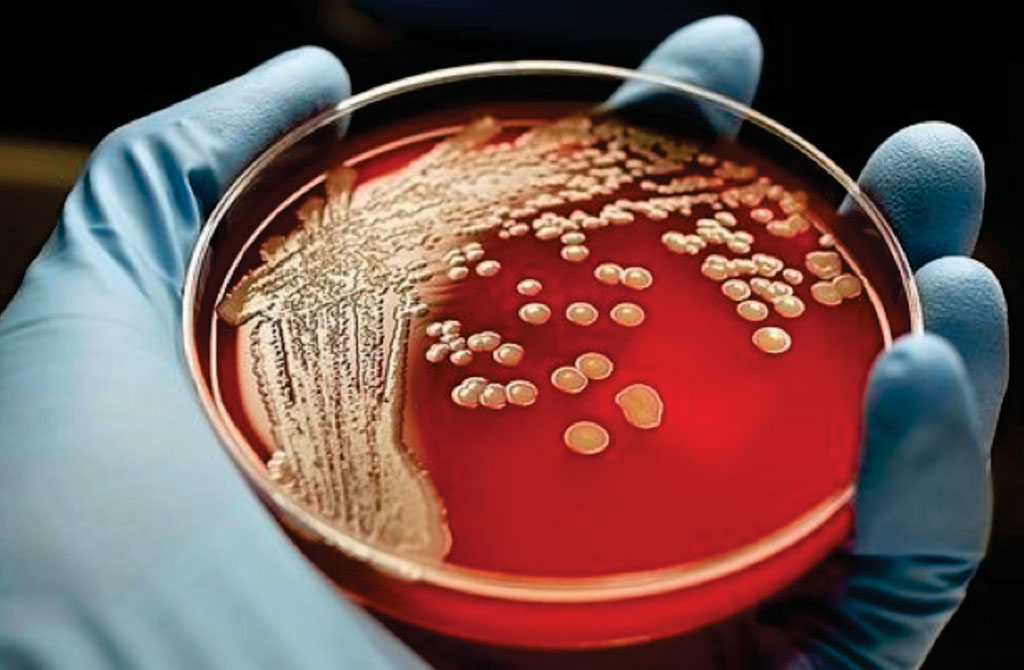
Image: Bacterial colonies of Staphylococcus aureus growing on horse blood agar (Photo courtesy of OMICS International).
Diagnostic testing for foodborne pathogens relies on culture-based techniques that are not rapid enough for real-time disease surveillance and do not give a quantitative picture of pathogen abundance or the response of the natural microbiome.
Metagenomics identifies the microbes present by sequencing the entire DNA present in a sample and comparing the genomic data to a database of known microbes. In addition to identifying the bacteria present in the samples, the methodology can also measure the relative abundance of each microbial species and their virulence potential, among other things.
A collaboration of scientists from the Centers for Disease Control and Prevention (Atlanta, GA USA) and the Georgia Institute of Technology (Atlanta, GA, USA) applied shotgun metagenomics to stool samples collected from two geographically isolated foodborne outbreaks in Alabama and Colorado, where the etiologic agents were identified as distinct strains of Salmonella enterica serovar Heidelberg by culture-dependent methods. The metagenomics data provided specific information about the bacterial phenotype involved and identified a secondary Staphylococcus aureus pathogen present in two of the samples tested. Knowing the specific phenotype can help in pinpointing the origins of an outbreak, while information about the secondary infection may help explain related factors such as the severity of the infection.
The scientists were also able to rule out one species, Escherichia coli (or E. coli), because the variant present was not of a virulent type. Variants of these bacteria are present naturally in the gut microbiome (called "commensal E. coli") while other variants are notorious enteric pathogens. Metagenomics showed the abundant E. coli population in the outbreak samples was probably commensal, and its growth may have been accelerated when conditions became more favorable during the Salmonella infection. In the two cases evaluated, scientists were able to determine that although the symptoms were similar, the outbreaks were caused by different variants of Salmonella and therefore were probably not connected.
Andrew D. Huang, PhD, a microbiologist/ bioinformatician and lead author of the study said, “Currently, the most advanced DNA fingerprinting method, whole genome sequencing, requires first pulling out, or isolating in a pure culture, the bacteria that made a person sick to generate a fingerprint. Metagenomics differs from whole genome sequencing because it could allow us to sequence the entire DNA in a patient's sample. It could allow us to skip the isolation steps and go directly from a stool sample to a highly detailed DNA fingerprint of the bacteria that made you sick. This method saves time and provides more detail that could be helpful for diagnosing a patient and identifying an outbreak.” The study was published on November 23, 2016, in the journal Applied and Environmental Microbiology.
Related Links:
Centers for Disease Control and Prevention
Georgia Institute of Technology
Metagenomics identifies the microbes present by sequencing the entire DNA present in a sample and comparing the genomic data to a database of known microbes. In addition to identifying the bacteria present in the samples, the methodology can also measure the relative abundance of each microbial species and their virulence potential, among other things.
A collaboration of scientists from the Centers for Disease Control and Prevention (Atlanta, GA USA) and the Georgia Institute of Technology (Atlanta, GA, USA) applied shotgun metagenomics to stool samples collected from two geographically isolated foodborne outbreaks in Alabama and Colorado, where the etiologic agents were identified as distinct strains of Salmonella enterica serovar Heidelberg by culture-dependent methods. The metagenomics data provided specific information about the bacterial phenotype involved and identified a secondary Staphylococcus aureus pathogen present in two of the samples tested. Knowing the specific phenotype can help in pinpointing the origins of an outbreak, while information about the secondary infection may help explain related factors such as the severity of the infection.
The scientists were also able to rule out one species, Escherichia coli (or E. coli), because the variant present was not of a virulent type. Variants of these bacteria are present naturally in the gut microbiome (called "commensal E. coli") while other variants are notorious enteric pathogens. Metagenomics showed the abundant E. coli population in the outbreak samples was probably commensal, and its growth may have been accelerated when conditions became more favorable during the Salmonella infection. In the two cases evaluated, scientists were able to determine that although the symptoms were similar, the outbreaks were caused by different variants of Salmonella and therefore were probably not connected.
Andrew D. Huang, PhD, a microbiologist/ bioinformatician and lead author of the study said, “Currently, the most advanced DNA fingerprinting method, whole genome sequencing, requires first pulling out, or isolating in a pure culture, the bacteria that made a person sick to generate a fingerprint. Metagenomics differs from whole genome sequencing because it could allow us to sequence the entire DNA in a patient's sample. It could allow us to skip the isolation steps and go directly from a stool sample to a highly detailed DNA fingerprint of the bacteria that made you sick. This method saves time and provides more detail that could be helpful for diagnosing a patient and identifying an outbreak.” The study was published on November 23, 2016, in the journal Applied and Environmental Microbiology.
Related Links:
Centers for Disease Control and Prevention
Georgia Institute of Technology
Latest Microbiology News
- Gut Microbiome Signatures Help Identify Risk of IBD Progression
- FDA-Cleared Gastrointestinal Panel Detects 24 Pathogen Targets
- New AMR Assay Supports Rapid Infection Control Screening in Hospitals
- Diagnostic Gaps Complicate Bundibugyo Ebola Outbreak Response in Congo
- Study Finds Hidden Mpox Infections May Drive Ongoing Spread
- Large-Scale Genomic Surveillance Tracks Resistant Bacteria Across European Hospitals
- Molecular Urine and Stool Tests Do Not Improve Early TB Treatment in Hospitalized HIV Patients
- Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
- Label-Free Microscopy Method Enables Faster, Quantitative Detection of Malaria
- Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
- Gut Microbiome Test Predicts Melanoma Recurrence After Surgery
- Rapid Blood-Culture Susceptibility Panel Expands Coverage for Gram-Negative Infections
- Antibiotic Resistance Genes Found in Newborns Within Hours of Birth
- Rapid Color Test Stratifies Virulent and Resistant Staph Strains
- mNGS CSF Test Identifies CNS Pathogens Missed by Standard Panels
- Syndromic Panel Enables Rapid Identification of Bloodstream Infections
Channels
Clinical Chemistry
view channel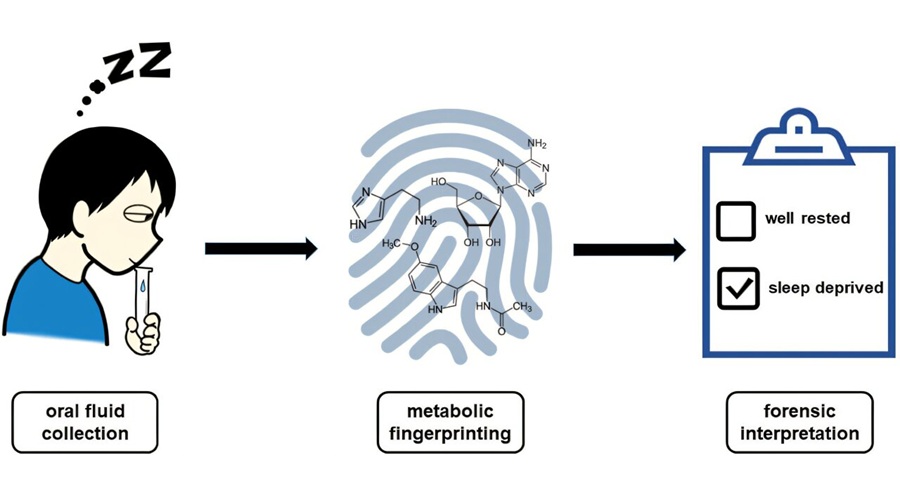
Saliva-Based Test Detects Biochemical Signs of Sleep Loss
Acute sleep loss impairs cognition and motor skills, raising safety risks that resemble alcohol intoxication. Clinicians currently lack an objective biochemical test to determine when someone is dangerously... Read more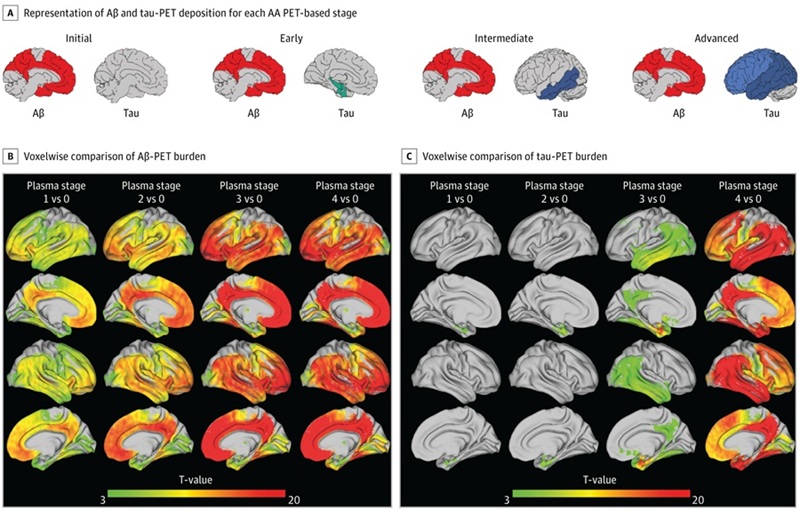
Simple Dual-Tau Blood Test Detects and Stages Alzheimer’s Disease
Alzheimer’s disease is typically confirmed and staged with positron emission tomography scans and cerebrospinal fluid testing, procedures that are costly and invasive. Broader access to minimally invasive... Read more
Alzheimer’s Blood Biomarkers Linked to Early Cognitive Differences Before Dementia
Blood-based screening for Alzheimer’s disease offers a noninvasive, lower-cost alternative to brain imaging or spinal fluid testing, yet its ability to flag the earliest cognitive changes has been unclear.... Read moreMolecular Diagnostics
view channel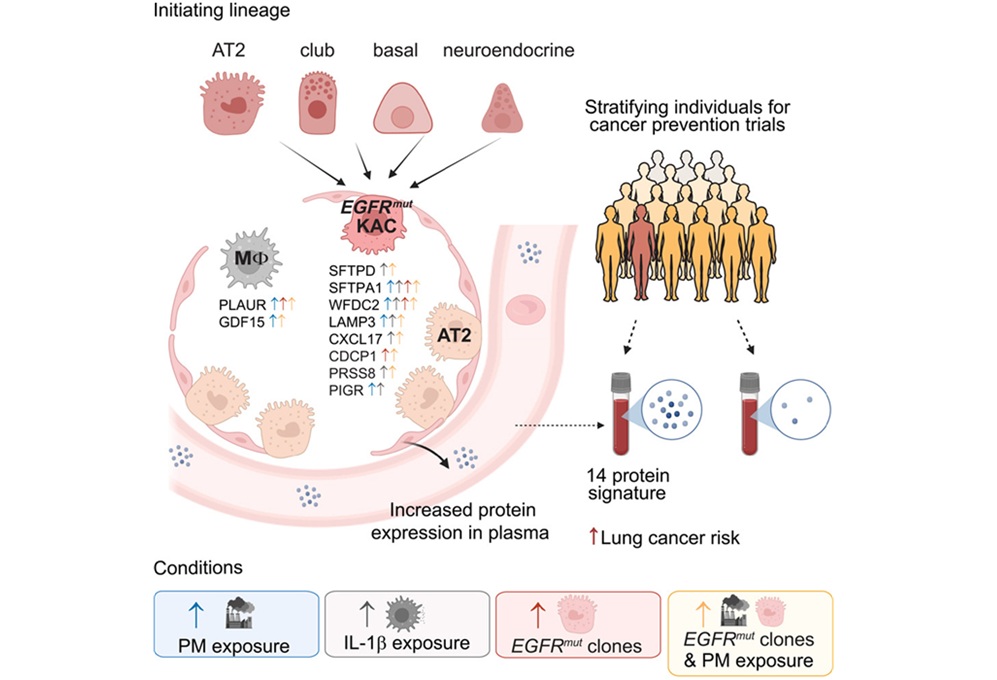
Plasma Protein Signature Predicts Lung Cancer Risk Up to Five Years Ahead
Lung cancer remains a leading cause of cancer death, and many cases are detected only after symptoms appear. Current screening programs largely target people with a history of smoking, leaving other at-risk... Read more
Circulating Tumor DNA Testing Guides Chemotherapy, Reduces Relapse in Colon Cancer
Adjuvant therapy decisions after curative surgery for colon cancer remain difficult, as conventional clinicopathologic factors often fail to capture residual disease risk. Liquid biopsy approaches that... Read moreHematology
view channel
Next-Generation Hematology Platform Streamlines High-Complexity Lab Workflows
Sysmex America (Chicago, IL, USA) has introduced the next generation XR-Series, centered on the XR-10 Automated Hematology Module for high-complexity laboratories. The platform builds on the widely used... Read more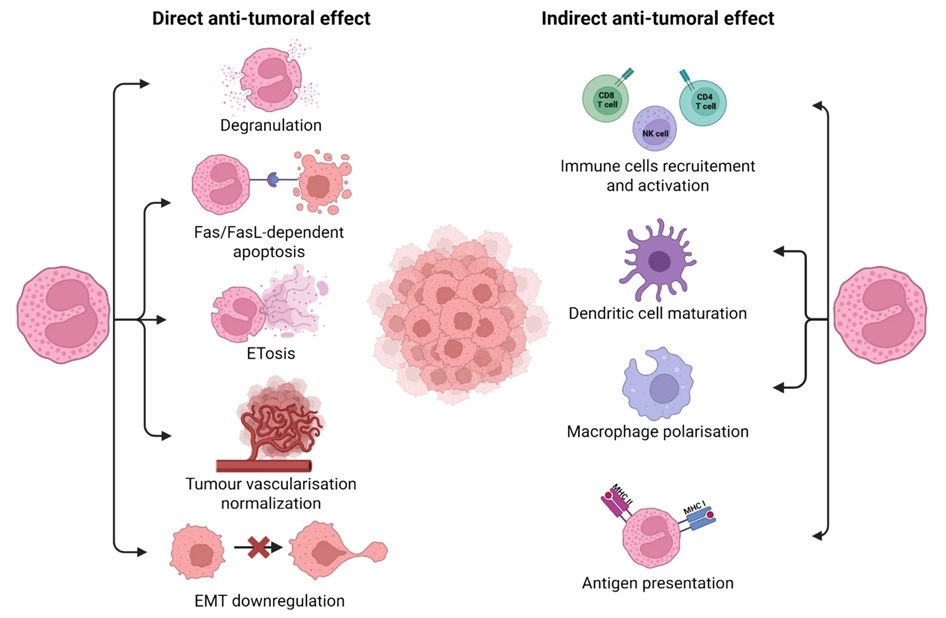
Blood Eosinophil Count May Predict Cancer Immunotherapy Response and Toxicity
Immune checkpoint inhibitors have improved outcomes across many cancers, yet only a subset of patients derive durable benefit and biomarkers to guide treatment remain limited. Eosinophils, best known for... Read moreImmunology
view channelAptamer-Based Biosensor Enables Mutation-Resilient SARS-CoV-2 Detection
Rapid evolution of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) can undermine existing molecular diagnostics, especially when assays target small viral components. Double-antibody sandwich... Read more
Study Points to Autoimmune Pathway Behind Long COVID Symptoms
Long COVID leaves many SARS-CoV-2 survivors with persistent fatigue, cognitive issues, palpitations, and musculoskeletal pain for months or years. Estimates cited in new research suggest 4%–20% of infected... Read more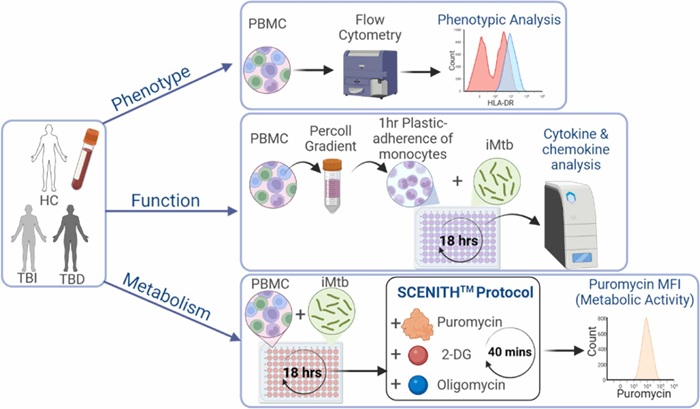
Metabolic Biomarker Distinguishes Latent from Active Tuberculosis and Tracks Treatment Response
Tuberculosis (TB) remains the world’s leading infectious killer, with 10.8 million cases and 1.25 million deaths recorded globally in 2023. Yet many infected individuals never develop active disease, underscoring... Read morePathology
view channel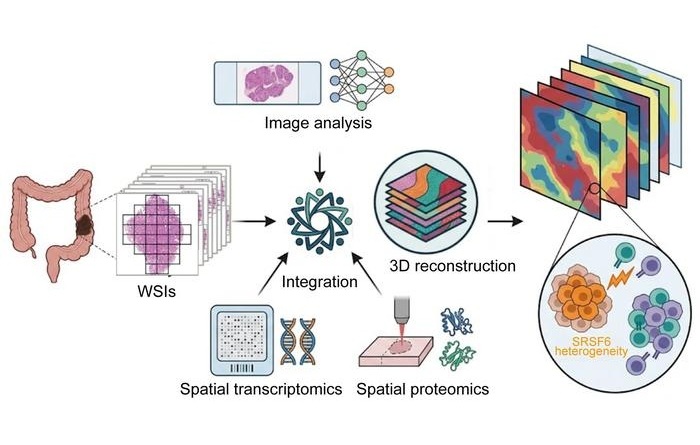
3D Spatial Multi-Omics Maps Intra-Tumor Diversity in Colorectal Cancer
Colorectal cancer remains a leading cause of cancer death, and clinical decision-making is complicated by marked intra-tumor heterogeneity. Conventional bulk sequencing averages molecular signals across... Read more
Blood-Based Method Tracks Gene Activity in the Living Brain
Real-time measurement of gene activity in the brain has been limited by assays requiring destructive tissue sampling. Tracking active genes could reveal how the body responds to environmental factors,... Read moreTechnology
view channel
AI Platform Links Biomarker Results to Cancer Clinical Trials and Guidelines
Oncology teams must manage growing volumes of genomic data, rapidly evolving clinical trial options, and frequently updated care guidelines, all within tight clinic schedules. Translating complex tumor... Read more
Agentic AI Platform Supports Genomic Decision-Making in Oncology
Oncology care teams increasingly face the challenge of managing complex molecular diagnostics, evolving treatment options, and extensive electronic health record documentation. Translating multimodal data... Read moreIndustry
view channel