Evolutionary Fitness for Codon Bias Selection Measured
By LabMedica International staff writers
Posted on 04 Apr 2016
Using bacteria as a model, researchers have for the first time quantified weak but effective selection forces that determine non-random codon usage in protein translation of RNA. Quantification of codon usage bias deepens our understanding of the selective forces driving long-term genome evolution and may have applications for rational design of synthetic genes and for heterologous gene expression in biotechnology.Posted on 04 Apr 2016
The genetic code is redundant, with individual amino acids being encoded by up to 6 different synonymous codons. Some codons are used more frequently than others, giving rise to codon usage bias (CUB), an important genome-shaping factor, especially among fast-growing organisms, including pathogenic bacteria and yeasts. Free-living unicellular organisms such as bacteria, which live in an environment of intense competitive selection, have an extreme CUB. Furthermore, a strong positive correlation between CUB and gene expression levels has been observed in many microorganisms. This has been attributed to selection for translational efficiency, however this putative selective advantage has never been measured in bacteria and theoretical estimates vary widely.
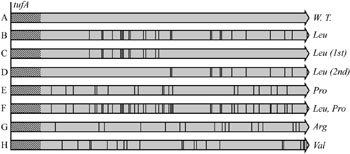
Image: Overview of the design of synonymous tuf gene alleles. The first 40 codons were left unchanged (hatched region). Black bars represent codons changed to synonymous codons. (A) Wild-type gene, tufA; (B) through (G) Mutated versions of tuf gene (Photo courtesy of Brandis Hughes, 2016, and PLoS Genetics).
In the new study, graduate student Gerrit Brandis and Prof. Diarmaid Hughes of Uppsala University (Uppsala, Sweden) used Salmonella bacteria to quantitatively address the question of how large a cost-benefit difference in fitness is required for selection to be effective.
The speed of RNA to protein translation is a determining factor of bacterial growth rate. The ancient mechanism of translation has been under selection for billions of years. The researchers investigated effects of changing various codons on producing the highly expressed EF-Tu, one of the most important Salmonella proteins. By systematically mutating codons in the gene, they found that changing even a single codon to any one of the alternative synonymous codons reduced the fitness of Salmonella. This suggested that the naturally used codons are a best fit and that any codon change reduces Salmonella’s fitness in that environment.
The researchers quantified the fitness cost of changing codons in the gene. On average, changing a single codon reduced Salmonella fitness by 0.01% per generation. So a difference of only 1/100th of 1% in fitness is sufficient for evolutionary selection that underlies the phenomenon of CUB. Over very long time scales (hundreds of millions of years), evolution selects for tiny differences in relative fitness, as small or smaller than 0.01% difference per generation for Salmonella.
The study was published March 10, 2016, in the journal PLoS Genetics.
Related Links:
Uppsala University