Programming Language Designed to Construct Synthetic DNA
By LabMedica International staff writers
Posted on 24 Oct 2013
Chemists and biotech researchers may soon be able to use a structured series of instructions to “program” how DNA molecules interact in a test tube or cell, similar to using a programming language to write computer code.Posted on 24 Oct 2013
A team of investigators, led by the University of Washington (UW; Seattle, USA), has developed a programming language for chemistry that it hopes will streamline efforts to design a network that can guide the behavior of chemical-reaction amalgams in the same manner that embedded electronic controllers guide robots, cars, and other devices. In medicine, such networks could serve as “smart” drug deliverers or disease detectors at the cellular level.
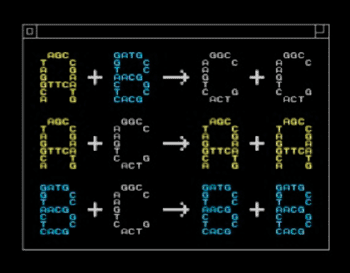
The study’s findings were published online September 29, 2013, in the journal Nature Nanotechnology. Chemists and educators teach and use chemical reaction networks, 100-year-old language of equations that describes how mixtures of chemicals behave. The UW engineers take this language an additional step and use it to write programs that direct the movement of customized molecules.
“We start from an abstract, mathematical description of a chemical system, and then use DNA to build the molecules that realize the desired dynamics,” said corresponding author Dr. Georg Seelig, a UW assistant professor of electrical engineering and of computer science and engineering. “The vision is that eventually, you can use this technology to build general-purpose tools.”
Currently, when a biologist or chemist makes a specific type of molecular network, the engineering process is complicated, cumbersome, and difficult to repurpose for constructing other systems. The UW engineers wanted to devise a framework that gives scientists more flexibility. Dr. Seelig compares this new approach to programming languages that tell a computer what to do. “I think this is appealing because it allows you to solve more than one problem,” he said. “If you want a computer to do something else, you just reprogram it. This project is very similar in that we can tell chemistry what to do.”
Humans and other organisms already have complex networks of nano-sized molecules that help to control cells and keep the body in line. Scientists now are finding ways to design synthetic systems that behave similar to biologic ones with the hope that synthetic molecules could support the body’s natural functions. To achieve this, a system is needed to create synthetic DNA molecules that vary according to their specific functions.
The new application is not yet ready to be utilized in medicine, but future uses could include using this framework to make molecules that self-assemble within cells and serve as “smart” sensors. These could be embedded in a cell, then programmed to identify abnormalities and respond as required, possibly by delivering drugs directly to those cells.
Drs. Seelig and colleague Eric Klavins, a UW associate professor of electrical engineering, recently received USD 2 million from the National Science Foundation (Washington DC, USA) as part of a national initiative to enhance research in molecular programming. The new language will be used to support that larger initiative, according to Dr. Seelig.
Related Links:
University of Washington