Spatial Tissue Profiled by Imaging-Free Molecular Tomography
|
By LabMedica International staff writers Posted on 06 May 2021 |
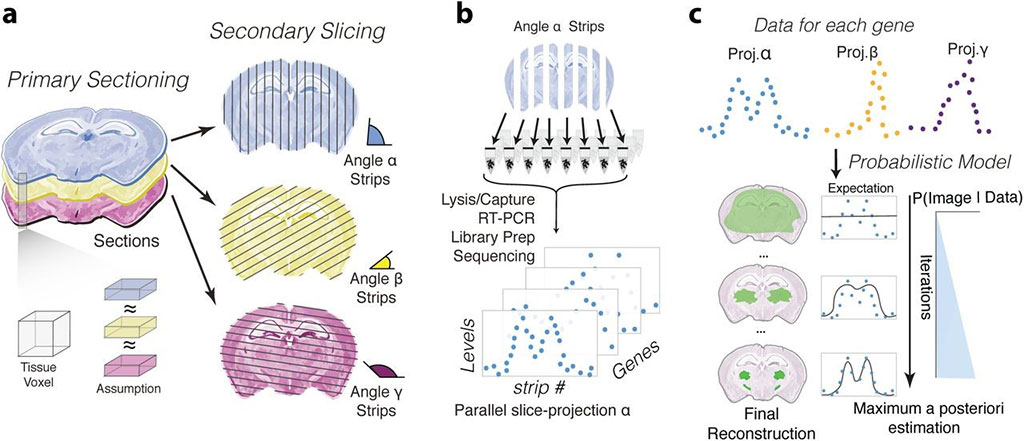
Image: Schematic representation of sampling and reconstruction approach to resolve the spatial localization of genomics data (Photo courtesy of Swiss Federal Institute of Technology)
Spatially resolved molecular atlases help scientists understand where different types of cells are located in the body and map their gene expression in specific locations in tissues and organs. However, many sequencing modalities lack spatial counterparts.
New technologies such as in situ hybridization can be used to map the expression of multiple genes on the same tissue sample and have accelerated the generation of new atlases. In situ hybridization allows for a target gene to be tagged ("hybridized") with a fluorescent marker within sections of a tissue ("in situ") and visualized under a specialized microscope. Several techniques are currently being developed for spatially resolved omics profiling, but each new method requires the setup of specific detection strategies or specialized instrumentation.
Life Scientists at the Swiss Federal Institute of Technology Lausanne (Lausanne, Switzerland) have created a computational algorithm called Tomographer that can transform gene-sequencing data into spatially resolved data such as images, without using a microscope. The framework uses a tissue sampling strategy based on multi-angle sectioning and an associated algorithm that enables the reconstruction of 2D spatial patterns.
The sampling technique involves cutting tissues into consecutive thin slices ("primary sections") that are subsequently further sliced along an orthogonal plane at predefined orientations ("secondary sections"), resulting in tissue strips spanning the entire tissue. Gene expression quantification of the sections is implemented using spatial transcriptomics by reoriented projections and sequencing (STRP-seq), a method that combines the sampling strategy presented above with a customized, low-input RNA-seq protocol based on single-cell tagged reverse transcription sequencing (STRT-seq) chemistry. The method produces parallel-slice projections for each gene by quantifying the reads that map to a transcript in each of the secondary sections.
The Tomographer framework was benchmarked for the ability to reconstruct transcriptome-wide spatial expression patterns against the Allen Adult Mouse Brain in situ hybridization atlas. First, the team measured 3,880 genes in the mouse brain. Then, they compared 923 reconstructed genes to the in situ hybridization data from the mouse brain atlas using Pearson's correlation coefficient and found that the Tomographer workflow was more than twice as accurate as iterative proportional fitting (IPF). They also compared Tomographer to the spatial reconstruction capabilities of IPF-based Tomo-seq.
The team noted that the quality of Tomographer's reconstructions depends on the balance between the number and width of the tissue strips sampled. They noted that four cutting angles provided results that are a fair compromise between the reconstruction quality and sample processing effort and cost. Also, the technique requires a distance of at least 1.15 times the secondary section width in order to discriminate between two distinct points of primary strips. The study was published on April 19, 2021 in the journal Nature Biotechnology.
Related Links:
Swiss Federal Institute of Technology
New technologies such as in situ hybridization can be used to map the expression of multiple genes on the same tissue sample and have accelerated the generation of new atlases. In situ hybridization allows for a target gene to be tagged ("hybridized") with a fluorescent marker within sections of a tissue ("in situ") and visualized under a specialized microscope. Several techniques are currently being developed for spatially resolved omics profiling, but each new method requires the setup of specific detection strategies or specialized instrumentation.
Life Scientists at the Swiss Federal Institute of Technology Lausanne (Lausanne, Switzerland) have created a computational algorithm called Tomographer that can transform gene-sequencing data into spatially resolved data such as images, without using a microscope. The framework uses a tissue sampling strategy based on multi-angle sectioning and an associated algorithm that enables the reconstruction of 2D spatial patterns.
The sampling technique involves cutting tissues into consecutive thin slices ("primary sections") that are subsequently further sliced along an orthogonal plane at predefined orientations ("secondary sections"), resulting in tissue strips spanning the entire tissue. Gene expression quantification of the sections is implemented using spatial transcriptomics by reoriented projections and sequencing (STRP-seq), a method that combines the sampling strategy presented above with a customized, low-input RNA-seq protocol based on single-cell tagged reverse transcription sequencing (STRT-seq) chemistry. The method produces parallel-slice projections for each gene by quantifying the reads that map to a transcript in each of the secondary sections.
The Tomographer framework was benchmarked for the ability to reconstruct transcriptome-wide spatial expression patterns against the Allen Adult Mouse Brain in situ hybridization atlas. First, the team measured 3,880 genes in the mouse brain. Then, they compared 923 reconstructed genes to the in situ hybridization data from the mouse brain atlas using Pearson's correlation coefficient and found that the Tomographer workflow was more than twice as accurate as iterative proportional fitting (IPF). They also compared Tomographer to the spatial reconstruction capabilities of IPF-based Tomo-seq.
The team noted that the quality of Tomographer's reconstructions depends on the balance between the number and width of the tissue strips sampled. They noted that four cutting angles provided results that are a fair compromise between the reconstruction quality and sample processing effort and cost. Also, the technique requires a distance of at least 1.15 times the secondary section width in order to discriminate between two distinct points of primary strips. The study was published on April 19, 2021 in the journal Nature Biotechnology.
Related Links:
Swiss Federal Institute of Technology
Latest Technology News
- New Diagnostic System Achieves PCR Testing Accuracy
- DNA Biosensor Enables Early Diagnosis of Cervical Cancer
- Self-Heating Microfluidic Devices Can Detect Diseases in Tiny Blood or Fluid Samples
- Breakthrough in Diagnostic Technology Could Make On-The-Spot Testing Widely Accessible
- First of Its Kind Technology Detects Glucose in Human Saliva
- Electrochemical Device Identifies People at Higher Risk for Osteoporosis Using Single Blood Drop
- Novel Noninvasive Test Detects Malaria Infection without Blood Sample
- Portable Optofluidic Sensing Devices Could Simultaneously Perform Variety of Medical Tests
- Point-of-Care Software Solution Helps Manage Disparate POCT Scenarios across Patient Testing Locations
- Electronic Biosensor Detects Biomarkers in Whole Blood Samples without Addition of Reagents
- Breakthrough Test Detects Biological Markers Related to Wider Variety of Cancers
- Rapid POC Sensing Kit to Determine Gut Health from Blood Serum and Stool Samples
- Device Converts Smartphone into Fluorescence Microscope for Just USD 50
- Wi-Fi Enabled Handheld Tube Reader Designed for Easy Portability
Channels
Clinical Chemistry
view channel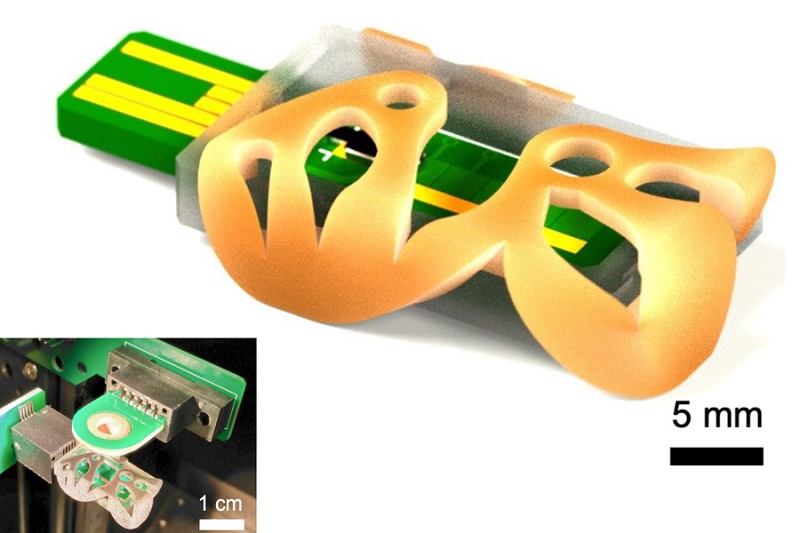
3D Printed Point-Of-Care Mass Spectrometer Outperforms State-Of-The-Art Models
Mass spectrometry is a precise technique for identifying the chemical components of a sample and has significant potential for monitoring chronic illness health states, such as measuring hormone levels... Read more.jpg)
POC Biomedical Test Spins Water Droplet Using Sound Waves for Cancer Detection
Exosomes, tiny cellular bioparticles carrying a specific set of proteins, lipids, and genetic materials, play a crucial role in cell communication and hold promise for non-invasive diagnostics.... Read more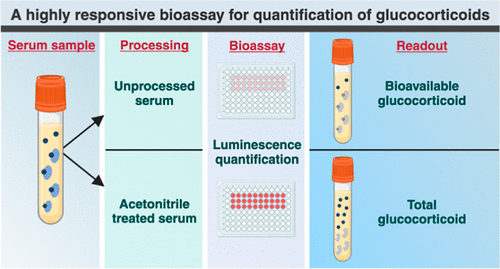
Highly Reliable Cell-Based Assay Enables Accurate Diagnosis of Endocrine Diseases
The conventional methods for measuring free cortisol, the body's stress hormone, from blood or saliva are quite demanding and require sample processing. The most common method, therefore, involves collecting... Read moreMolecular Diagnostics
view channel.jpg)
First of Its Kind NGS Assay for Precise Detection of BCR::ABL1 Fusion Gene to Enable Personalized Leukemia Treatment
The BCR::ABL1 fusion gene plays a key role in the pathogenesis of several blood cancers, particularly chronic myeloid leukemia (CML). This gene results from a chromosomal translocation that causes constitutive... Read more
Urine Test to Revolutionize Lyme Disease Testing
Lyme disease is the most common animal-to-human transmitted disease in the United States, with around 476,000 people diagnosed and treated annually, and its incidence has been increasing.... Read more
Simple Blood Test Could Enable First Quantitative Assessments for Future Cerebrovascular Disease
Cerebral small vessel disease is a common cause of stroke and cognitive decline, particularly in the elderly. Presently, assessing the risk for cerebral vascular diseases involves using a mix of diagnostic... Read more
New Genetic Testing Procedure Combined With Ultrasound Detects High Cardiovascular Risk
A key interest area in cardiovascular research today is the impact of clonal hematopoiesis on cardiovascular diseases. Clonal hematopoiesis results from mutations in hematopoietic stem cells and may lead... Read moreHematology
view channel
Next Generation Instrument Screens for Hemoglobin Disorders in Newborns
Hemoglobinopathies, the most widespread inherited conditions globally, affect about 7% of the population as carriers, with 2.7% of newborns being born with these conditions. The spectrum of clinical manifestations... Read more
First 4-in-1 Nucleic Acid Test for Arbovirus Screening to Reduce Risk of Transfusion-Transmitted Infections
Arboviruses represent an emerging global health threat, exacerbated by climate change and increased international travel that is facilitating their spread across new regions. Chikungunya, dengue, West... Read more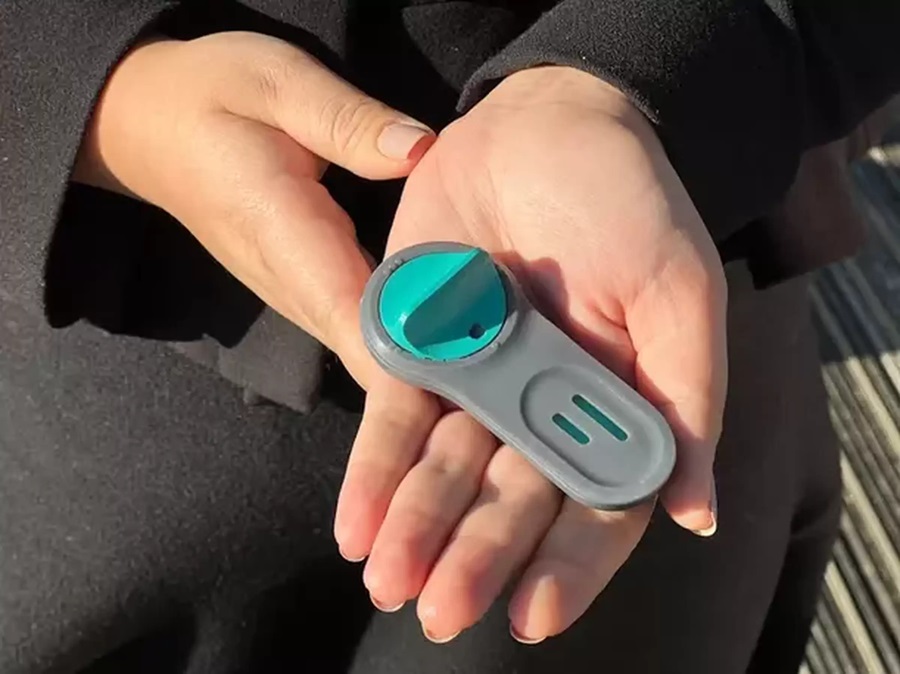
POC Finger-Prick Blood Test Determines Risk of Neutropenic Sepsis in Patients Undergoing Chemotherapy
Neutropenia, a decrease in neutrophils (a type of white blood cell crucial for fighting infections), is a frequent side effect of certain cancer treatments. This condition elevates the risk of infections,... Read more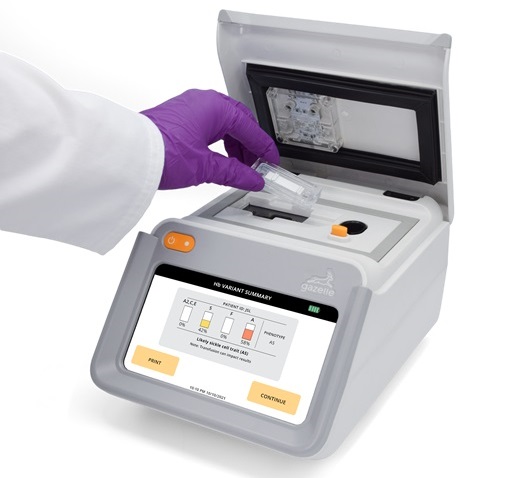
First Affordable and Rapid Test for Beta Thalassemia Demonstrates 99% Diagnostic Accuracy
Hemoglobin disorders rank as some of the most prevalent monogenic diseases globally. Among various hemoglobin disorders, beta thalassemia, a hereditary blood disorder, affects about 1.5% of the world's... Read moreImmunology
view channel
Diagnostic Blood Test for Cellular Rejection after Organ Transplant Could Replace Surgical Biopsies
Transplanted organs constantly face the risk of being rejected by the recipient's immune system which differentiates self from non-self using T cells and B cells. T cells are commonly associated with acute... Read more
AI Tool Precisely Matches Cancer Drugs to Patients Using Information from Each Tumor Cell
Current strategies for matching cancer patients with specific treatments often depend on bulk sequencing of tumor DNA and RNA, which provides an average profile from all cells within a tumor sample.... Read more
Genetic Testing Combined With Personalized Drug Screening On Tumor Samples to Revolutionize Cancer Treatment
Cancer treatment typically adheres to a standard of care—established, statistically validated regimens that are effective for the majority of patients. However, the disease’s inherent variability means... Read moreMicrobiology
view channel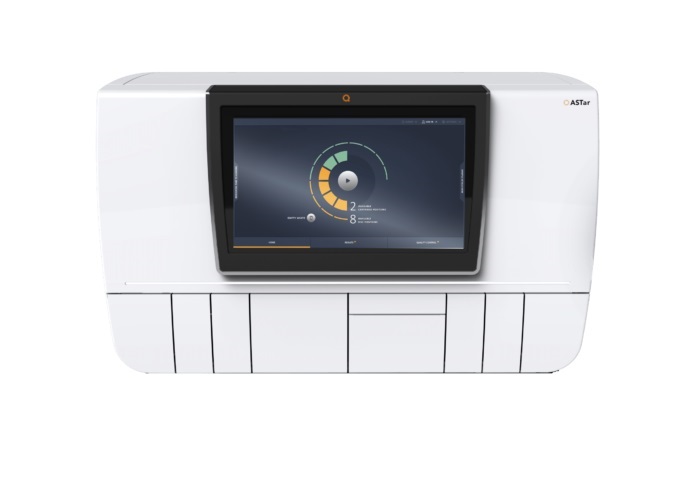
Automated Sepsis Test System Enables Rapid Diagnosis for Patients with Severe Bloodstream Infections
Sepsis affects up to 50 million people globally each year, with bacteraemia, formerly known as blood poisoning, being a major cause. In the United States alone, approximately two million individuals are... Read moreEnhanced Rapid Syndromic Molecular Diagnostic Solution Detects Broad Range of Infectious Diseases
GenMark Diagnostics (Carlsbad, CA, USA), a member of the Roche Group (Basel, Switzerland), has rebranded its ePlex® system as the cobas eplex system. This rebranding under the globally renowned cobas name... Read more
Clinical Decision Support Software a Game-Changer in Antimicrobial Resistance Battle
Antimicrobial resistance (AMR) is a serious global public health concern that claims millions of lives every year. It primarily results from the inappropriate and excessive use of antibiotics, which reduces... Read morePathology
view channel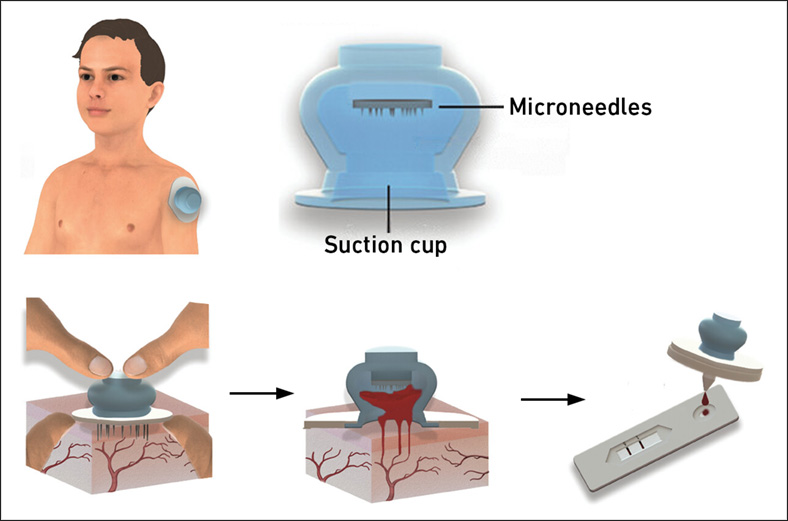
New Blood Test Device Modeled on Leeches to Help Diagnose Malaria
Many individuals have a fear of needles, making the experience of having blood drawn from their arm particularly distressing. An alternative method involves taking blood from the fingertip or earlobe,... Read more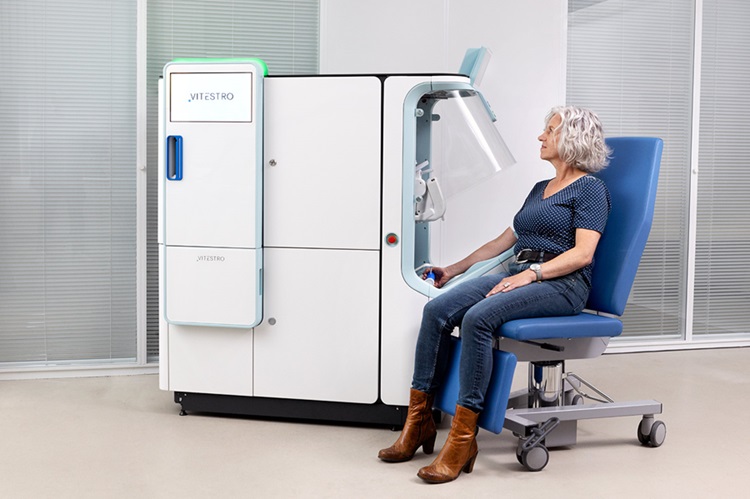
Robotic Blood Drawing Device to Revolutionize Sample Collection for Diagnostic Testing
Blood drawing is performed billions of times each year worldwide, playing a critical role in diagnostic procedures. Despite its importance, clinical laboratories are dealing with significant staff shortages,... Read more.jpg)
Use of DICOM Images for Pathology Diagnostics Marks Significant Step towards Standardization
Digital pathology is rapidly becoming a key aspect of modern healthcare, transforming the practice of pathology as laboratories worldwide adopt this advanced technology. Digital pathology systems allow... Read more
First of Its Kind Universal Tool to Revolutionize Sample Collection for Diagnostic Tests
The COVID pandemic has dramatically reshaped the perception of diagnostics. Post the pandemic, a groundbreaking device that combines sample collection and processing into a single, easy-to-use disposable... Read moreIndustry
view channel_1.jpg)
Thermo Fisher and Bio-Techne Enter Into Strategic Distribution Agreement for Europe
Thermo Fisher Scientific (Waltham, MA USA) has entered into a strategic distribution agreement with Bio-Techne Corporation (Minneapolis, MN, USA), resulting in a significant collaboration between two industry... Read more
ECCMID Congress Name Changes to ESCMID Global
Over the last few years, the European Society of Clinical Microbiology and Infectious Diseases (ESCMID, Basel, Switzerland) has evolved remarkably. The society is now stronger and broader than ever before... Read more