Internet-Based Tool Simplifies Infectious Disease Diagnosis
|
By LabMedica International staff writers Posted on 07 Jun 2016 |
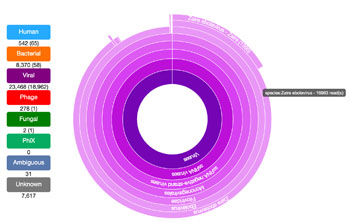
Image: The metagenomics pathogen detection tool could change how infectious diseases are diagnosed (Photo courtesy of the University of Utah).
An Internet-based tool for diagnosis of infectious diseases through the rapid interpretation of DNA sequencing data is now available to the medical community.
The metagenomics analysis software that powers the "Taxonomer" program was developed at the University of Utah (Salt Lake City, USA). Metagenomics, the genomic analysis of a population of microorganisms, makes possible the profiling of microbial communities in the environment and the human body at unprecedented depth and breadth. Its rapidly expanding use is revolutionizing the understanding of microbial diversity in natural and man-made environments and is linking microbial community profiles with health and disease.
The Taxonomer program is an ultrafast, web-tool for comprehensive metagenomics data analysis and interactive results visualization. The DNA sequence data for a clinical sample is uploaded via the Internet to Taxonomer. In less than one minute, the tool displays a thumbnail inventory of all pathogens in the sample, including viruses, bacteria, and fungi. Taxonomer is unique in providing integrated nucleotide and protein-based classification and simultaneous host messenger RNA (mRNA) transcript profiling.
Using real-world case studies, the developers showed that Taxonomer detected previously unrecognized infections and revealed antiviral host mRNA expression profiles.
“Our benchmark analyses show Taxonomer being ten to a hundred times faster than similar tools,” said senior author Dr. Robert Schlaberg, assistant professor of clinical pathology at the University of Utah. “Taxonomer provides a critical step forward, as it is extremely fast, accurate, and easy enough to use for implementation in diagnostic laboratories.”
To facilitate datasharing across geographic distances in outbreak settings, Taxonomer is publicly available through a web-based user interface at the address http://taxonomer.com.
Results of the Taxonomer study were published in the May 26, 2016, online edition of the journal Genome Biology.
Related Links:
University of Utah
Taxonomer
The metagenomics analysis software that powers the "Taxonomer" program was developed at the University of Utah (Salt Lake City, USA). Metagenomics, the genomic analysis of a population of microorganisms, makes possible the profiling of microbial communities in the environment and the human body at unprecedented depth and breadth. Its rapidly expanding use is revolutionizing the understanding of microbial diversity in natural and man-made environments and is linking microbial community profiles with health and disease.
The Taxonomer program is an ultrafast, web-tool for comprehensive metagenomics data analysis and interactive results visualization. The DNA sequence data for a clinical sample is uploaded via the Internet to Taxonomer. In less than one minute, the tool displays a thumbnail inventory of all pathogens in the sample, including viruses, bacteria, and fungi. Taxonomer is unique in providing integrated nucleotide and protein-based classification and simultaneous host messenger RNA (mRNA) transcript profiling.
Using real-world case studies, the developers showed that Taxonomer detected previously unrecognized infections and revealed antiviral host mRNA expression profiles.
“Our benchmark analyses show Taxonomer being ten to a hundred times faster than similar tools,” said senior author Dr. Robert Schlaberg, assistant professor of clinical pathology at the University of Utah. “Taxonomer provides a critical step forward, as it is extremely fast, accurate, and easy enough to use for implementation in diagnostic laboratories.”
To facilitate datasharing across geographic distances in outbreak settings, Taxonomer is publicly available through a web-based user interface at the address http://taxonomer.com.
Results of the Taxonomer study were published in the May 26, 2016, online edition of the journal Genome Biology.
Related Links:
University of Utah
Taxonomer
Latest Microbiology News
- Integrated Solution Ushers New Era of Automated Tuberculosis Testing
- Automated Sepsis Test System Enables Rapid Diagnosis for Patients with Severe Bloodstream Infections
- Enhanced Rapid Syndromic Molecular Diagnostic Solution Detects Broad Range of Infectious Diseases
- Clinical Decision Support Software a Game-Changer in Antimicrobial Resistance Battle
- New CE-Marked Hepatitis Assays to Help Diagnose Infections Earlier
- 1 Hour, Direct-From-Blood Multiplex PCR Test Identifies 95% of Sepsis-Causing Pathogens
- Mouth Bacteria Test Could Predict Colon Cancer Progression
- Unique Metabolic Signature Could Enable Sepsis Diagnosis within One Hour of Blood Collection
- Groundbreaking Diagnostic Platform Provides AST Results With Unprecedented Speed
- Simple Blood Test Combined With Personalized Risk Model Improves Sepsis Diagnosis
- Blood Analysis Predicts Sepsis and Organ Failure in Children
- TB Blood Test Could Detect Millions of Silent Spreaders
- New Blood Test Cuts Diagnosis Time for Nontuberculous Mycobacteria Infections from Months to Hours
- New Tuberculosis Test to Expand Testing Access in Low- and Middle-Income Countries
- Rapid Test Diagnoses Tropical Disease within Hours for Faster Antibiotics Treatment
- Rapid Molecular Testing Enables Faster, More Targeted Antibiotic Treatment for Pneumonia
Channels
Clinical Chemistry
view channel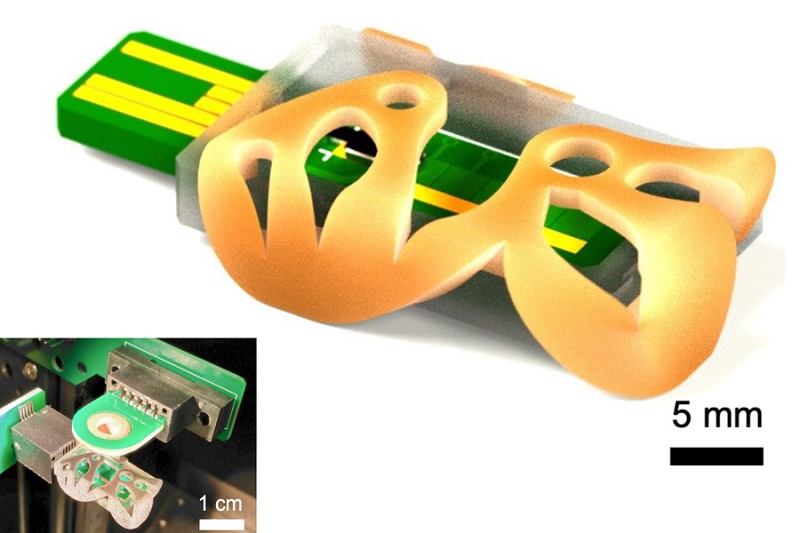
3D Printed Point-Of-Care Mass Spectrometer Outperforms State-Of-The-Art Models
Mass spectrometry is a precise technique for identifying the chemical components of a sample and has significant potential for monitoring chronic illness health states, such as measuring hormone levels... Read more.jpg)
POC Biomedical Test Spins Water Droplet Using Sound Waves for Cancer Detection
Exosomes, tiny cellular bioparticles carrying a specific set of proteins, lipids, and genetic materials, play a crucial role in cell communication and hold promise for non-invasive diagnostics.... Read more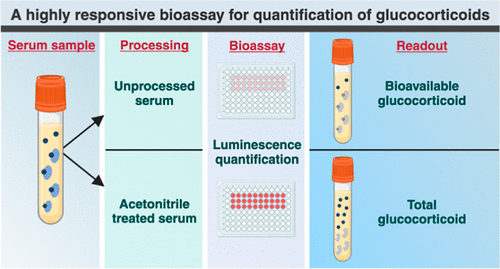
Highly Reliable Cell-Based Assay Enables Accurate Diagnosis of Endocrine Diseases
The conventional methods for measuring free cortisol, the body's stress hormone, from blood or saliva are quite demanding and require sample processing. The most common method, therefore, involves collecting... Read moreMolecular Diagnostics
view channelBlood Proteins Could Warn of Cancer Seven Years before Diagnosis
Two studies have identified proteins in the blood that could potentially alert individuals to the presence of cancer more than seven years before the disease is clinically diagnosed. Researchers found... Read moreUltrasound-Aided Blood Testing Detects Cancer Biomarkers from Cells
Ultrasound imaging serves as a noninvasive method to locate and monitor cancerous tumors effectively. However, crucial details about the cancer, such as the specific types of cells and genetic mutations... Read moreHematology
view channel
Next Generation Instrument Screens for Hemoglobin Disorders in Newborns
Hemoglobinopathies, the most widespread inherited conditions globally, affect about 7% of the population as carriers, with 2.7% of newborns being born with these conditions. The spectrum of clinical manifestations... Read more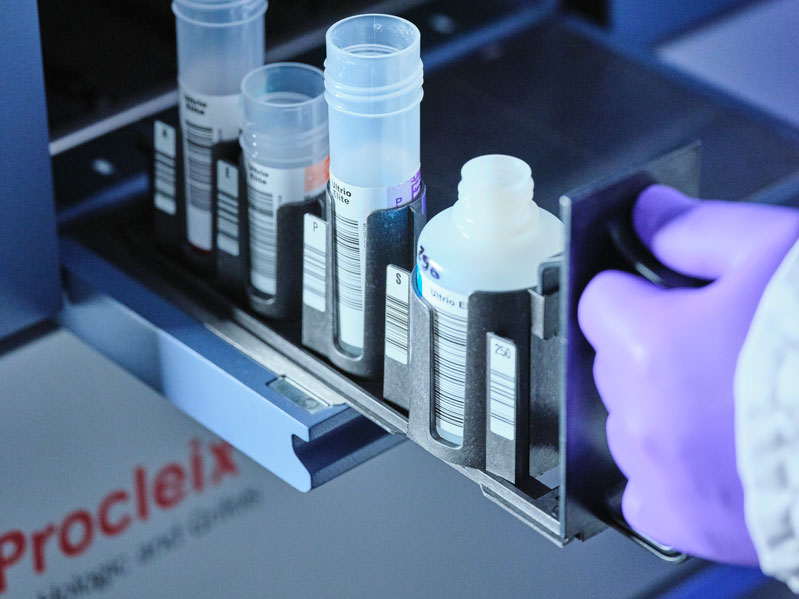
First 4-in-1 Nucleic Acid Test for Arbovirus Screening to Reduce Risk of Transfusion-Transmitted Infections
Arboviruses represent an emerging global health threat, exacerbated by climate change and increased international travel that is facilitating their spread across new regions. Chikungunya, dengue, West... Read more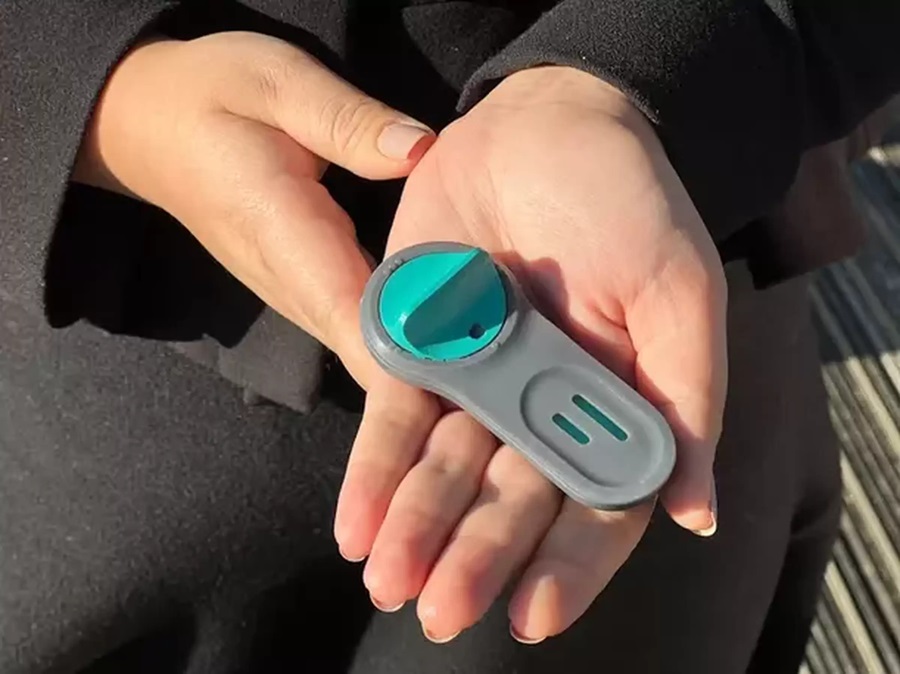
POC Finger-Prick Blood Test Determines Risk of Neutropenic Sepsis in Patients Undergoing Chemotherapy
Neutropenia, a decrease in neutrophils (a type of white blood cell crucial for fighting infections), is a frequent side effect of certain cancer treatments. This condition elevates the risk of infections,... Read more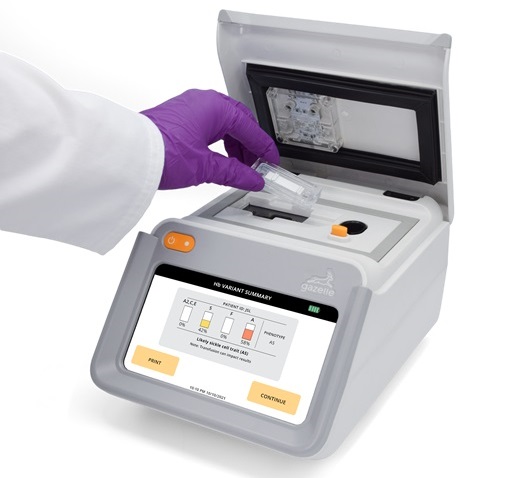
First Affordable and Rapid Test for Beta Thalassemia Demonstrates 99% Diagnostic Accuracy
Hemoglobin disorders rank as some of the most prevalent monogenic diseases globally. Among various hemoglobin disorders, beta thalassemia, a hereditary blood disorder, affects about 1.5% of the world's... Read moreImmunology
view channel.jpg)
AI Predicts Tumor-Killing Cells with High Accuracy
Cellular immunotherapy involves extracting immune cells from a patient's tumor, potentially enhancing their cancer-fighting capabilities through engineering, and then expanding and reintroducing them into the body.... Read more
Diagnostic Blood Test for Cellular Rejection after Organ Transplant Could Replace Surgical Biopsies
Transplanted organs constantly face the risk of being rejected by the recipient's immune system which differentiates self from non-self using T cells and B cells. T cells are commonly associated with acute... Read more
AI Tool Precisely Matches Cancer Drugs to Patients Using Information from Each Tumor Cell
Current strategies for matching cancer patients with specific treatments often depend on bulk sequencing of tumor DNA and RNA, which provides an average profile from all cells within a tumor sample.... Read more
Genetic Testing Combined With Personalized Drug Screening On Tumor Samples to Revolutionize Cancer Treatment
Cancer treatment typically adheres to a standard of care—established, statistically validated regimens that are effective for the majority of patients. However, the disease’s inherent variability means... Read moreMicrobiology
view channel
Integrated Solution Ushers New Era of Automated Tuberculosis Testing
Tuberculosis (TB) is responsible for 1.3 million deaths every year, positioning it as one of the top killers globally due to a single infectious agent. In 2022, around 10.6 million people were diagnosed... Read more
Automated Sepsis Test System Enables Rapid Diagnosis for Patients with Severe Bloodstream Infections
Sepsis affects up to 50 million people globally each year, with bacteraemia, formerly known as blood poisoning, being a major cause. In the United States alone, approximately two million individuals are... Read moreEnhanced Rapid Syndromic Molecular Diagnostic Solution Detects Broad Range of Infectious Diseases
GenMark Diagnostics (Carlsbad, CA, USA), a member of the Roche Group (Basel, Switzerland), has rebranded its ePlex® system as the cobas eplex system. This rebranding under the globally renowned cobas name... Read more
Clinical Decision Support Software a Game-Changer in Antimicrobial Resistance Battle
Antimicrobial resistance (AMR) is a serious global public health concern that claims millions of lives every year. It primarily results from the inappropriate and excessive use of antibiotics, which reduces... Read morePathology
view channel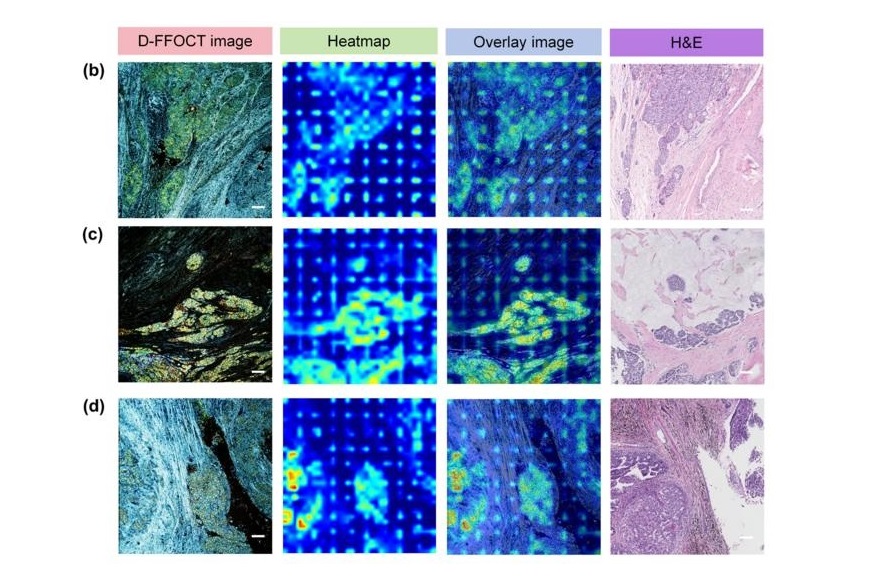
AI Integrated With Optical Imaging Technology Enables Rapid Intraoperative Diagnosis
Rapid and accurate intraoperative diagnosis is essential for tumor surgery as it guides surgical decisions with precision. Traditional intraoperative assessments, such as frozen sections based on H&E... Read more
HPV Self-Collection Solution Improves Access to Cervical Cancer Testing
Annually, over 604,000 women across the world are diagnosed with cervical cancer, and about 342,000 die from this disease, which is preventable and primarily caused by the Human Papillomavirus (HPV).... Read moreHyperspectral Dark-Field Microscopy Enables Rapid and Accurate Identification of Cancerous Tissues
Breast cancer remains a major cause of cancer-related mortality among women. Breast-conserving surgery (BCS), also known as lumpectomy, is the removal of the cancerous lump and a small margin of surrounding tissue.... Read moreIndustry
view channel
Danaher and Johns Hopkins University Collaborate to Improve Neurological Diagnosis
Unlike severe traumatic brain injury (TBI), mild TBI often does not show clear correlations with abnormalities detected through head computed tomography (CT) scans. Consequently, there is a pressing need... Read more
Beckman Coulter and MeMed Expand Host Immune Response Diagnostics Partnership
Beckman Coulter Diagnostics (Brea, CA, USA) and MeMed BV (Haifa, Israel) have expanded their host immune response diagnostics partnership. Beckman Coulter is now an authorized distributor of the MeMed... Read more_1.jpg)











_1.jpg)
.jpg)