Epigenetic Markers Predict Type 2 Diabetes Patients Response to Metformin
|
By LabMedica International staff writers Posted on 29 Sep 2020 |
Image: Various types of data analysis using BeadStudio Methylation Module with Illumina`s MethylationEpic array (Photo courtesy of Phoebe Lu).
Generally, metformin is the first medication prescribed for type 2 diabetes (T2D). It works by lowering glucose production in the liver and improving the body's sensitivity to insulin so that the body uses insulin more effectively. Currently, there are no phenotypes that successfully predict glycemic response to, or tolerance of, metformin.
About 30% of patients do not respond to metformin and between 20% and 30% experience side effects that can be intolerable Gastrointestinal side effects of metformin are observed in 10% to 15% of patients, depending on the dose, and include abdominal discomfort, anorexia, bloating, and diarrhea. Because insulin secretion is unaltered, hypoglycemia is not a side effect of metformin used as monotherapy.
An international team of clinical scientists led by those at Skåne University Hospital (Malmo, Sweden) conducted multiple epigenome-wide association studies by analyzing in the blood of drug-naïve patients who were recently diagnosed with T2D. Blood samples were collected from the All New Diabetics In Scandia (ANDIS) cohort and analyzed using Illumina's MethylationEpic array (Illumina, San Diego, CA, USA). The team sought to gauge whether differences in DNA methylation prior to treatment could predict whether individuals had changes in glycated hemoglobin (HbA1c), responded to the drug treatment, or experienced intolerance to the drug following about a year and a half of metformin treatment.
The investigators identified more than 2,500 methylation sites that were significantly associated with changes in HbA1c, a marker of blood glucose levels. In the replication cohort, 132 CpGs of these sites were validated. They additionally uncovered 7,916 methylation sites that differed between individuals with T2D who responded to metformin and individuals who did not. Of those, 601 were then validated in the ANDIS replication cohort and 329 in an additional cohorts.
In all, 33 CpG sites were associated with future metformin response in all cohorts, and in a combined meta-analysis 11 sites reached genome-wide significance. At the same time, the team found 9,676 methylation sites that differed between individuals with T2D who could tolerate metformin treatment and those who could not. In the ANDIS replication cohort, 235 CpGs were validated, and in the replication cohort, 352 CpGs were. Seven CpGs were associated with metformin in all cohorts, and in a combined meta-analysis four sites reached genome-wide significance.
The scientists generated two methylation risk scores, one of metformin response and one of metformin intolerance. For the metformin response, they bundled together the 11 sites to form a weighted methylation risk score that could differentiate between responders and non-responders with an area under the curve of between 0.80 and 0.89. Meanwhile, for metformin intolerance, they combined the four sites that reached genome-wide significance into a separate risk score that could differentiate between tolerant and intolerant individuals with an area under the curve of between 0.85 and 0.94.
The authors concludes that they could discriminate between glycemic responders/non-responders and participants tolerant/intolerant to metformin at diagnosis by measuring blood-based epigenetic markers in drug-naïve patients with T2D. This epigenetics-based tool may be further developed to help patients with T2D receive optimal therapy. The study was published on September 16, 2020 in the journal Science Translational Medicine.
About 30% of patients do not respond to metformin and between 20% and 30% experience side effects that can be intolerable Gastrointestinal side effects of metformin are observed in 10% to 15% of patients, depending on the dose, and include abdominal discomfort, anorexia, bloating, and diarrhea. Because insulin secretion is unaltered, hypoglycemia is not a side effect of metformin used as monotherapy.
An international team of clinical scientists led by those at Skåne University Hospital (Malmo, Sweden) conducted multiple epigenome-wide association studies by analyzing in the blood of drug-naïve patients who were recently diagnosed with T2D. Blood samples were collected from the All New Diabetics In Scandia (ANDIS) cohort and analyzed using Illumina's MethylationEpic array (Illumina, San Diego, CA, USA). The team sought to gauge whether differences in DNA methylation prior to treatment could predict whether individuals had changes in glycated hemoglobin (HbA1c), responded to the drug treatment, or experienced intolerance to the drug following about a year and a half of metformin treatment.
The investigators identified more than 2,500 methylation sites that were significantly associated with changes in HbA1c, a marker of blood glucose levels. In the replication cohort, 132 CpGs of these sites were validated. They additionally uncovered 7,916 methylation sites that differed between individuals with T2D who responded to metformin and individuals who did not. Of those, 601 were then validated in the ANDIS replication cohort and 329 in an additional cohorts.
In all, 33 CpG sites were associated with future metformin response in all cohorts, and in a combined meta-analysis 11 sites reached genome-wide significance. At the same time, the team found 9,676 methylation sites that differed between individuals with T2D who could tolerate metformin treatment and those who could not. In the ANDIS replication cohort, 235 CpGs were validated, and in the replication cohort, 352 CpGs were. Seven CpGs were associated with metformin in all cohorts, and in a combined meta-analysis four sites reached genome-wide significance.
The scientists generated two methylation risk scores, one of metformin response and one of metformin intolerance. For the metformin response, they bundled together the 11 sites to form a weighted methylation risk score that could differentiate between responders and non-responders with an area under the curve of between 0.80 and 0.89. Meanwhile, for metformin intolerance, they combined the four sites that reached genome-wide significance into a separate risk score that could differentiate between tolerant and intolerant individuals with an area under the curve of between 0.85 and 0.94.
The authors concludes that they could discriminate between glycemic responders/non-responders and participants tolerant/intolerant to metformin at diagnosis by measuring blood-based epigenetic markers in drug-naïve patients with T2D. This epigenetics-based tool may be further developed to help patients with T2D receive optimal therapy. The study was published on September 16, 2020 in the journal Science Translational Medicine.
Latest Molecular Diagnostics News
- Blood Test Maps Tumor Microenvironment to Predict Immunotherapy Response
- Liquid Biopsy Biomarkers Distinguish Inflammatory Breast Cancer and Support Monitoring
- Multiplex Respiratory Panel Integrates Automated Extraction to Streamline High-Volume Testing
- Whole-Blood RNA Test Predicts Disease Trajectory and Treatment Response
- Blood-Based Epigenetic Test Predicts GLP-1 Response and Tracks Treatment Effects
- Tumor Genomic Testing Guides Immunotherapy Selection in Pituitary Tumors
- Liquid Biopsy Predicts Immunotherapy Response in Breast Cancer
- New Blood Test Distinguishes Pancreatic Cancer From Benign Disease
- Noninvasive Test Confirms High-Risk Prenatal Screening Results from Blood
- Machine-Learning Genetic Risk Score Improves Early Prediction of Type 1 Diabetes
- Rapid Tongue Swab Molecular Test Detects Pulmonary Tuberculosis at Point of Care
- CRISPR-Based Test Identifies Multiple Respiratory Viruses Simultaneously
- Blood Test Receives FDA Breakthrough Status to Differentiate Schizophrenia and Bipolar Disorder
- Portable Test Detects Tuberculosis from Tongue Swabs in 30 Minutes
- Multi-Omic Assay Predicts Recurrence and Radiation Benefit in Early Breast Cancer
- Genomic Risk Score Identifies Inherited Risk for Multiple Cardiovascular Conditions
Channels
Clinical Chemistry
view channel
Ultrasensitive Test Detects Key Biomarker of Frontotemporal Dementia Subtype
Dementia affects more than 57 million people worldwide and is projected to nearly double within two decades, straining health systems and families. While biomarkers now enable accurate identification of... Read more
Routine Blood Tests Years Before Pregnancy Could Identify Preeclampsia Risk
High blood pressure during pregnancy is common and can progress to pre-eclampsia, making close monitoring at antenatal visits essential. However, most risk assessment begins only after pregnancy has started.... Read moreHematology
view channel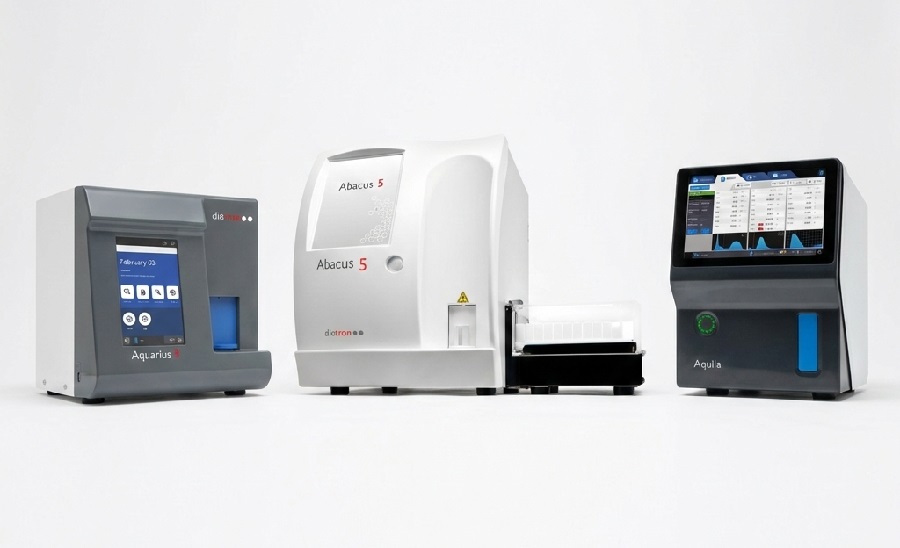
Advanced CBC-Derived Indices Integrated into Hematology Platforms
Diatron, a STRATEC brand, has introduced six advanced hematological indices on its Aquila, Aquarius 3, and Abacus 5 hematology analyzers. The new Research Use Only (RUO) indices include Neutrophil-to-Lymphocyte... Read more
Blood Test Enables Early Detection of Multiple Myeloma Relapse
Bone marrow biopsies remain central to diagnosing and monitoring multiple myeloma, yet the procedure is painful, invasive, and often repeated over time. Older patients—who represent most new cases—can... Read moreImmunology
view channel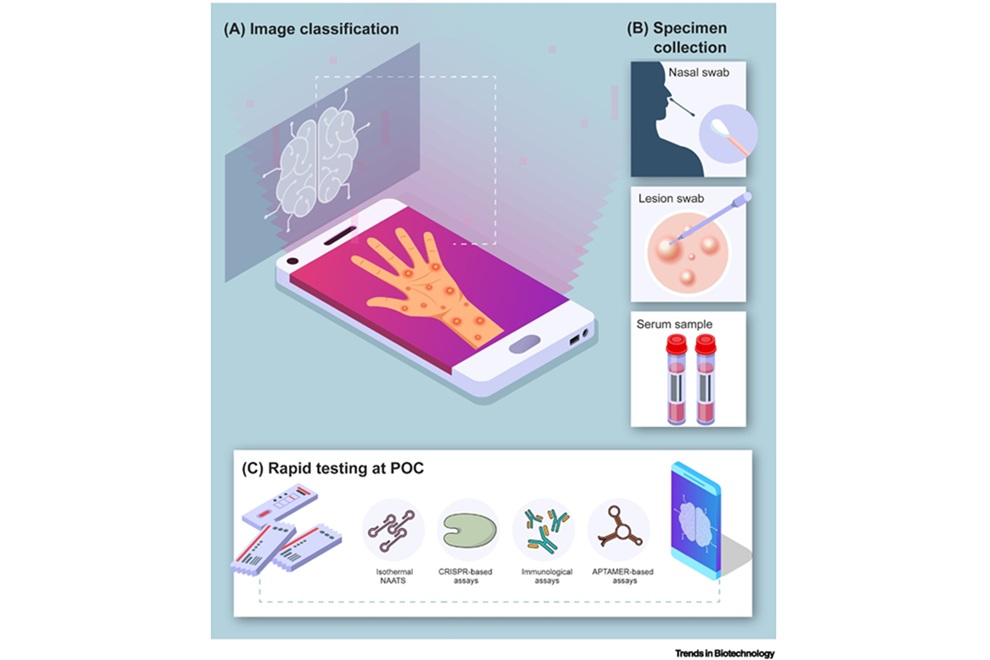
Point-of-Care Tests Could Expand Access to Mpox Diagnosis
Mpox outbreaks in non-endemic regions have underscored the need for rapid, accessible diagnostics to limit transmission. Polymerase chain reaction (PCR) remains the clinical reference, yet it depends on... Read more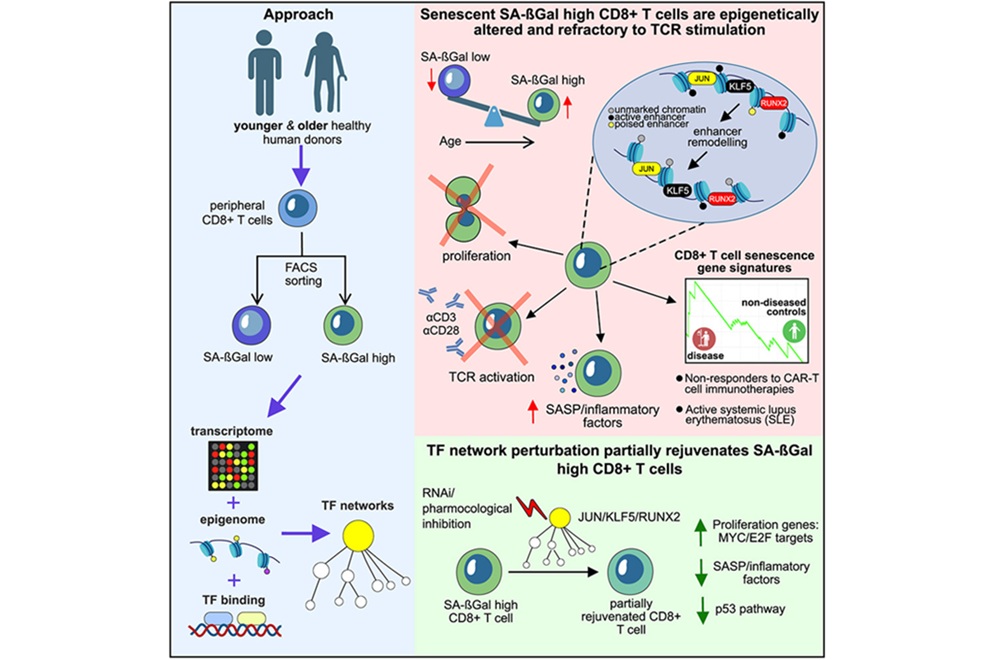
T-Cell Senescence Profiling May Predict CAR T Responses
Chimeric antigen receptor (CAR) T-cell therapy can deliver striking, durable remissions, yet many patients experience minimal or no benefit. The quality of patient-derived cytotoxic T lymphocytes used... Read moreMicrobiology
view channel
Rapid Antigen Biosensor Detects Active Tuberculosis in One Hour
Tuberculosis remains a major global health challenge and continues to drive significant morbidity and mortality. The World Health Organization’s 2024 global report cites it as the leading cause of death... Read more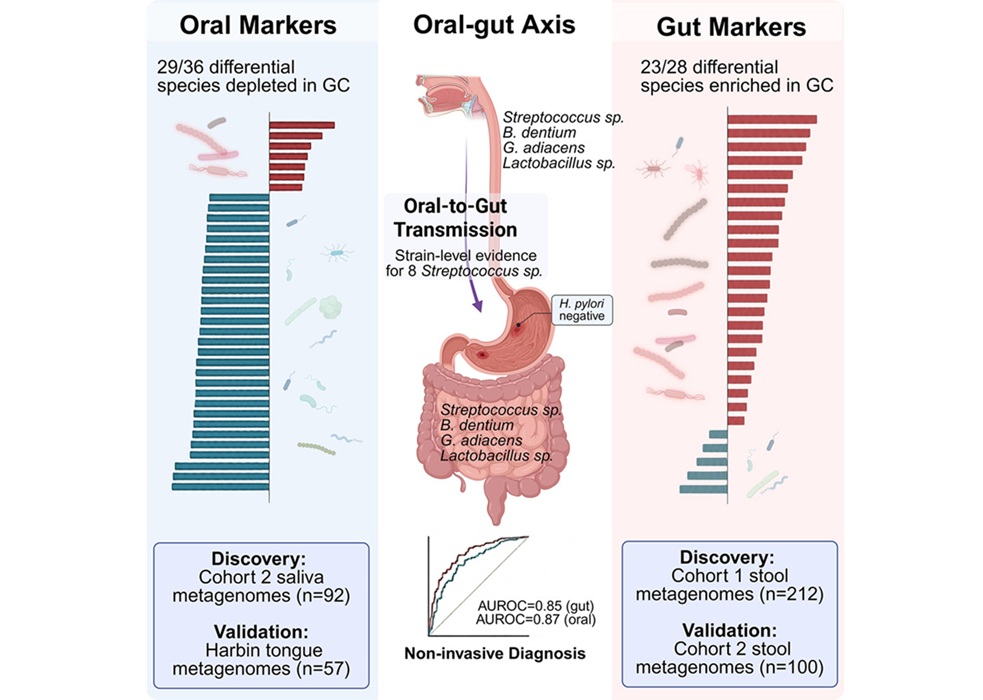
Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
Early detection of gastric cancer could be advanced by scalable screening strategies using minimally invasive sampling. Saliva collection is noninvasive and cost-effective, supporting wider adoption... Read morePathology
view channel
FDA Clears AI Digital Pathology Tool for Breast Cancer Risk Stratification
Risk assessment at diagnosis is central to guiding therapy for early-stage, hormone receptor-positive, human epidermal growth factor receptor 2-negative (HR+/HER2-) invasive breast cancer, where overtreatment... Read more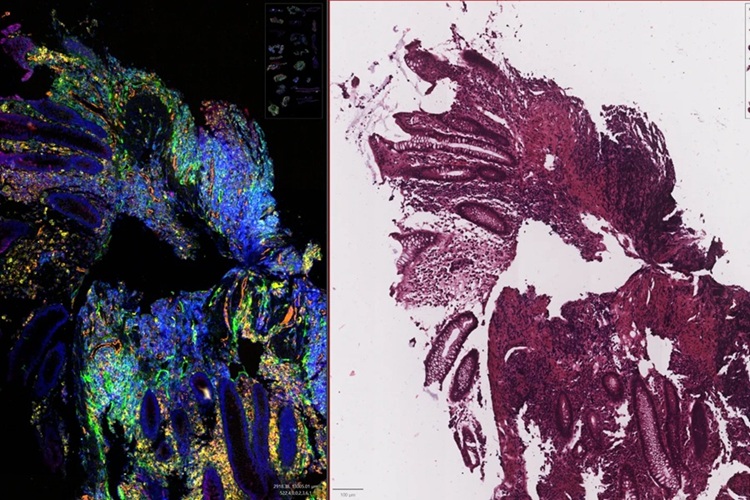
New AI Tool Reveals Hidden Genetic Signals in Routine H&E Slides
Pathologists worldwide rely on hematoxylin and eosin (H&E) slides to examine tissue architecture, yet these stains do not reveal the underlying molecular activity that often drives disease.... Read moreTechnology
view channel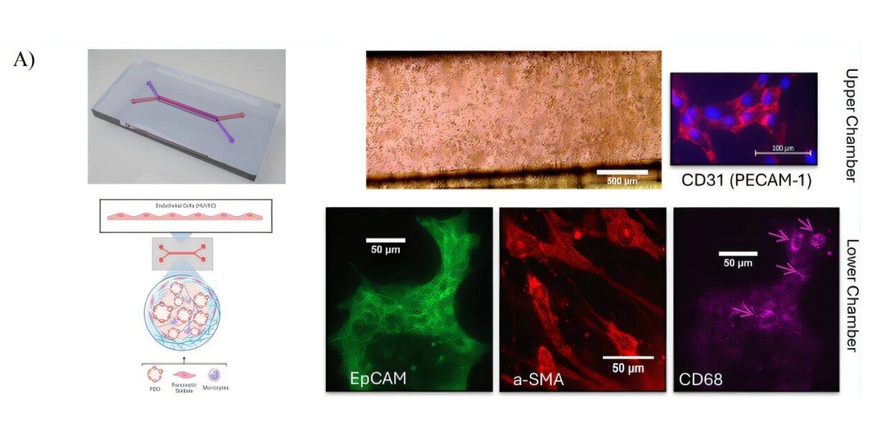
Tumor-on-a-Chip Platform Models Pancreatic Cancer Treatment Response
Pancreatic cancer remains one of the hardest malignancies to treat because tumors are embedded within a dense microenvironment that shapes growth and therapy response. Standard laboratory models often... Read more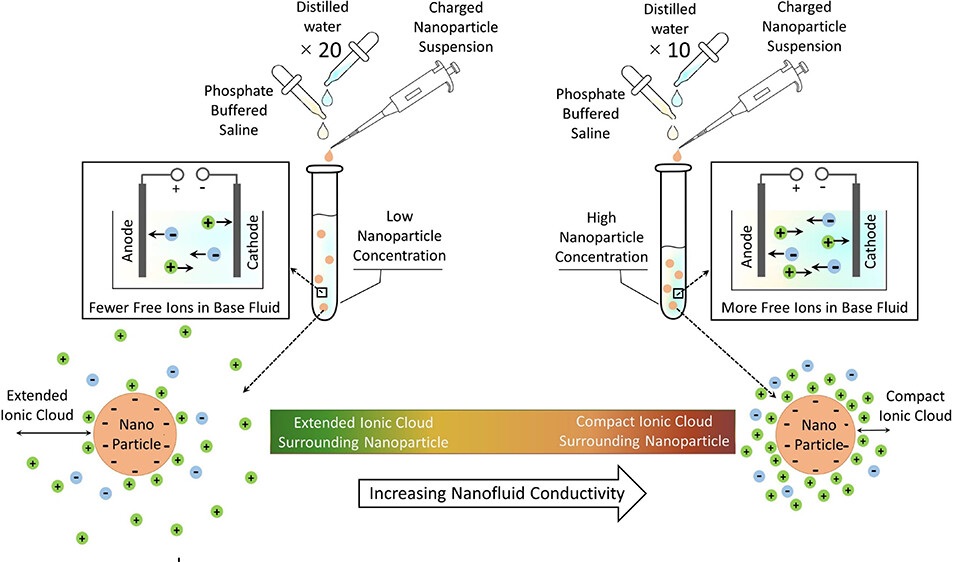
New Platform Captures Extracellular Vesicles for Early Cancer Detection
Early diagnosis remains the most effective way to reduce cancer mortality, yet many screening tools miss disease at its earliest stages. Biomarkers shed by tumors into blood and other fluids can be scarce... Read moreIndustry
view channel
Roche to Acquire PathAI for Up to $1.05 Billion to Strengthen AI Diagnostics Portfolio
Roche has entered into a definitive merger agreement to acquire PathAI, a company focused on digital pathology and artificial intelligence for pathology laboratories and the biopharma industry.... Read more