Sequencing Assay Adopted for Advanced Lung Cancer Reflex Testing
By LabMedica International staff writers
Posted on 31 Dec 2018
Guidelines are rapidly moving toward upfront profiling with next-generation sequencing (NGS) of all advanced lung cancers, but there are still practical issues with this testing method that can limit clinical testing and prevent or delay patients being started on targeted therapies.Posted on 31 Dec 2018
The growing number of genomically targeted therapies has made genomic testing an important part of the care for patients with non-small cell lung cancer. However, limited tissue availability, cost and long turnaround times can create barriers to efficient genomic testing and subsequent treatment. Effective approaches to reduce these barriers are needed.
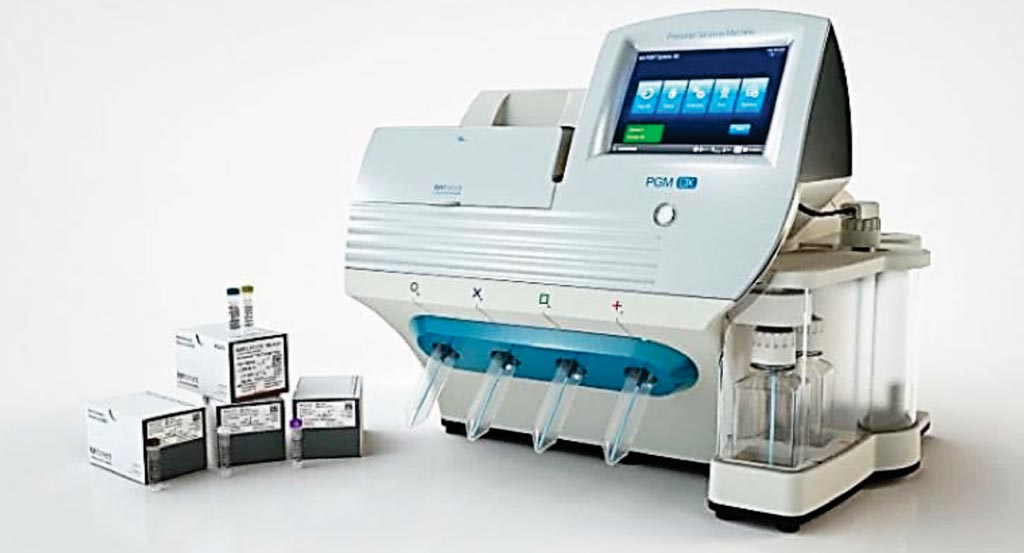

Image: The Oncomine Focus Assay is performed using the Ion Torrent technology. The test is designed to help oncologists expedite selection of a treatment plan for their patients in days instead of weeks (Photo courtesy of Thermo Fisher Scientific).
A team of scientists collaborating with the University Hospitals Cleveland Medical Center (Cleveland, OH, USA) tested 302 advanced lung adenocarcinomas from consecutive patients using a hybrid DNA/RNA NGS panel. Sample testing was reflexed from pathology for all stage III or IV tumors. Genomic alterations were tiered according to their clinical relevance and reported with guideline-recommended therapies.
The investigators used the Oncomine Focus Assay, which is an NGS oncology assay designed to simultaneously analyze hundreds of variants across 52 genes relevant to solid tumors. The assay enables concurrent analysis of DNA and RNA in a single workflow to detect hot spots, single nucleotide variants, indels, copy number variants, and gene fusions in various types of solid tumors. The clinicians validated the assay for use at the UHCMC CLIA-certified translational laboratory and expanded it to include 17 new biomarkers.
With a sample cohort consisting of 64% biopsies, 16% excisions/resections and 20% fine needle aspirations, the assay was reliable with a 95% success rate. The average turnaround time from receipt of unstained formalin-fixed paraffin embedded slides to reporting was 4.8±2.1 days, half of the recommended 10 days and similar to single-gene testing. Through testing, the clinicians found alterations associated with Food and Drug Administration-approved or the National Cancer Center Network (NCCN) guideline-recommended targeted therapies in 18% of cases, and they sent 60% of those patients for genomically guided therapies.
Joydeep Goswami, PhD, MBA, president of clinical next-generation sequencing and oncology for Thermo Fisher Scientific, said, “Perhaps the most important finding in this study is the ability to identify a much broader set of relevant targets in half the time recommended by NCCN guidelines, without increasing cost to the hospital. Single assay, multibiomarker tests with low tissue sample requirements allow more patients to access targeted therapies and lead to significantly higher test success rates, while simultaneously reducing total cost of care.” The study was published in the December 2018 issue of the Journal of Clinical Pathology.
Related Links:
University Hospitals Cleveland Medical Center