Robust Method Developed for Microbiome Analysis
By LabMedica International staff writers
Posted on 16 Mar 2016
Scientists have developed a technique for genome sequence data analysis that enables to more efficiently and accurately identify differences between metagenomes for a variety of bacterial communities, which can help to study, diagnose, and treat many human diseases. In a new study, the method was successfully tested on intestinal microbiota.Posted on 16 Mar 2016
A team led by scientists from the Moscow Institute of Physics and Technology (Moscow, Russia) have proposed a new method for comparison of metagenome-coupled DNA sequences from all organisms in a biological sample. The method makes it possible to more effectively solve the task of comparing samples and can be easily embedded into a metagenome data-analysis process.
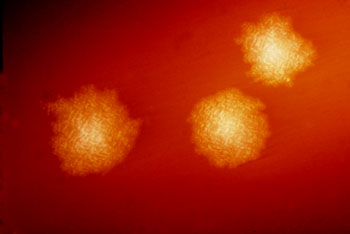
Image: Clostridium difficile colonies after 48 hours’ growth on blood agar plate. By competitive exclusion, healthy gut flora help limit growth of pathogenic yeasts and bacteria. Overgrowth of C. difficile in the gut can cause pseudomembranous colitis, and is the most frequently identified cause of antibiotic-associated diarrhea. (Image courtesy of Dr. Holdeman, Centers for Disease Control & Prevention (CDC; image ID #3647), and Wikimedia.)
Bacterial cells in the human body, most of which are located in the gut, hold a special place in metagenomics, including the "Human Microbiome Project." Microbiota composition is sensitive to processes occurring in the body. Thus, comparing samples from patients with samples from people with a healthy intestinal metagenome, will likely lead to methods that can evaluate risk of various diseases, including diabetes and inflammatory bowel disease.
The traditional approach to metagenome analysis is to compare samples on the basis of their taxonomic composition: percentages for each microbial species found. To determine sample composition, its genetic sequences are compared with a “reference set” database of known bacterial genomes. However, this approach has several disadvantages. Firstly, the reference genomes are often inaccurate, since the composition of the reference genome is a computationally complex and time-consuming task, especially for difficult-to-cultivate species; and the genomes of species isolated in the laboratory can carry genes significantly different from the same species living in a natural environment. Secondly, generally not all organisms are collected in the reference set of genomes (e.g., viruses). Therefore, the part of the sample sequence that does not match with the reference sample, is simply not taken into account during the analysis, despite the fact that it can be quite significant.
The new method is based on a comparison of “k-mer” frequencies, which does not require recourse to a reference sample or availability of any information on organisms being examined. All sequences in the sample are subjected to analysis, providing optimal results. Each genomic sequence is represented as a set with all instances of nucleotide "words" of specified length "k," called k-mers. As each genome sequence is unique, the sets of such "words" differ between individual organisms. Thus, the set of all k-mers for a metagenome can be viewed as a set of sets, namely of its constituent organisms. This enables assessment of the differences in the bacterial composition when comparing samples.
To test the effectiveness of the k-mer technique compared to traditional approaches, two sets of metagenome data were used—a set of real data and a set of artificially generated data. Artificial data (created from genomes with proportions known beforehand) is convenient for testing the method as the sequence is precisely known and the result can be assessed by comparing with an a priori correct value.
As the real-data set: intestinal metagenomes from residents of the United States and China were used. Intestinal bacterial communities differ significantly between different populations, and algorithms have claimed to allow to find exactly those indicators that show the difference in composition. Therefore, the criterion for assessing the effectiveness of the new method was the extent to which the metagenomes can be distinguished, that is how much the Chinese metagenomes differ in general from American ones.
The k-mers comparison method has shown better results in both data types than by using traditional mapping with a reference set. In addition, when using real data, a mismatch between the intestinal results for k-mer and traditional approaches allowed the researchers to detect another important component of the intestinal metagenome: namely the bacterial phage crAssphage, which had escaped the notice of researchers using the traditional method.
"Interestingly, the genes can be viewed not only as segments of DNA with proteins encoded in them, but also as information in general. It is this information distinction that has allowed us to identify new segments of DNA not described in the catalog of known genes. It [will be] interesting to see how this approach will be used by other research groups," said coauthor Dmitry Alexeev.
The study, by Dubinkina VB et al., was published January 16, 2016, in the journal BMC Bioinformatics.
Related Links:
Moscow Institute of Physics and Technology




.jpg)