Oral–Gut Microbiome Signatures Identify Early Gastric Cancer
Posted on 24 Apr 2026
Early detection of gastric cancer could be advanced by scalable screening strategies using minimally invasive sampling. Saliva collection is noninvasive and cost-effective, supporting wider adoption in population screening. New findings show that oral-to-gut microbial signatures measured in saliva and stool can enable early detection.
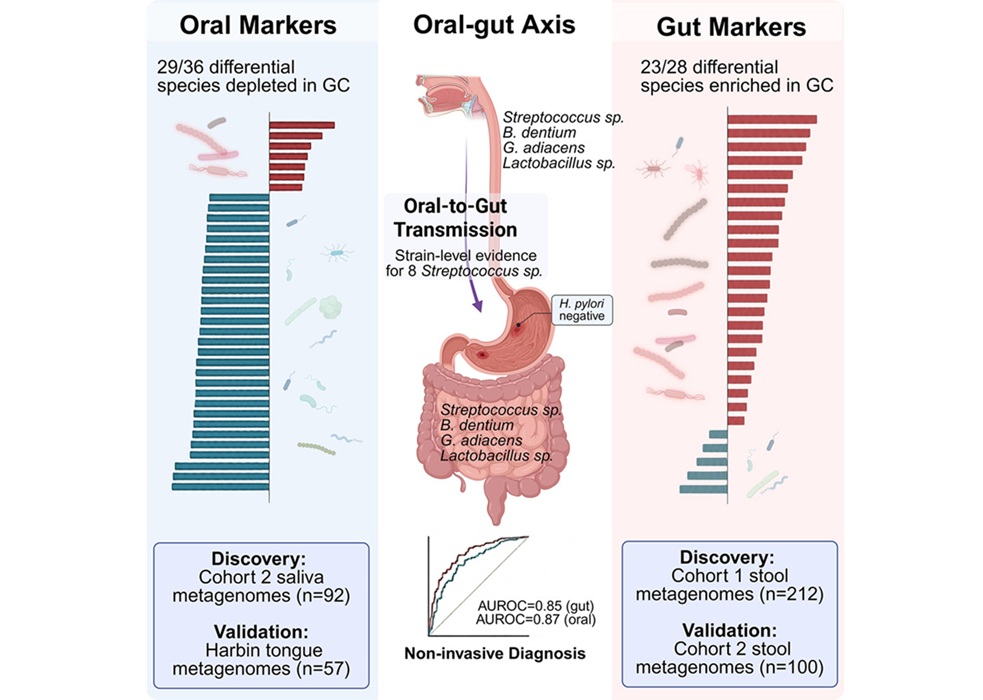
BGI Genomics, working with Renji Hospital at Shanghai Jiao Tong University School of Medicine, identified distinct microbial biomarkers in the oral cavity and gut associated with early gastric cancer. The innovation centers on saliva‑ and stool‑based microbial signatures that discriminate patients using machine learning models. The approach positions noninvasive sampling as a foundation for large‑scale screening.
Using high‑precision metagenomic sequencing of 404 samples, investigators observed a marked taxonomic shift in patients with gastric cancer, with 28 species showing differential abundance and 23 enriched in cases. Twenty enriched species were shared across sites, typically residing in the oral cavity but also present in the gut of affected individuals. Strain‑level analysis using population‑average nucleotide identity (popANI) showed greater than 99.9% genetic similarity between oral and gut strains within the same person, indicating oral‑to‑gut translocation.
Mechanistically, lactic acid–producing bacteria (LAB) that move from the mouth into the gut form interconnected communities that are more resilient and shift metabolism toward lactic acid production. This creates a more acidic tumor environment that activates enzymes involved in tissue remodeling, invasion, and new blood vessel formation, while also helping tumors evade immune responses. In an initiator–promoter model, Helicobacter pylori (Hp) acts as the initiator by triggering chronic inflammation, while oral LAB act as promoters by colonizing weakened tissue and driving disease progression—helping explain cases in Hp-negative patients and ongoing risk after Hp eradication.
Machine learning models built on these microbial markers achieved an area under the receiver operating characteristic curve (AUROC) of 0.87 for saliva‑based detection and 0.85 for stool‑based detection. The study was published in Cell Reports Medicine on April 20, with a related paper in Gut (2026) describing how Streptococcus anginosus promotes gastric cancer via methionine metabolites. The authors note that saliva collection’s noninvasive, cost‑effective nature makes it a practical option for large‑scale early screening and highlight the oral‑gut axis as a target for future diagnostic and microbiome‑based therapeutic development.