Rapid Filter Method Created for Isolation of CTCs from Whole Blood
By LabMedica International staff writers
Posted on 04 Nov 2019
Isolation and analysis of circulating tumor cells (CTCs) from blood samples present a potential means for basic cancer research as well as diagnosis and personalized treatment of the disease.Posted on 04 Nov 2019
CTCs are precursors of metastasis in several types of cancer and are occasionally found within the bloodstream in association with non-malignant cells such as white blood cells (WBCs). The identity and function of these CTC-associated WBCs, as well as the molecular features that define the interaction between WBCs and CTCs, has been poorly defined. The ability to study CTCs in this context is limited by current CTC isolation technologies, whihc rely on small blood volumes from a single venipuncture limiting the number of captured CTCs. This produces statistical variability and inaccurate reflection of tumor cell heterogeneity.
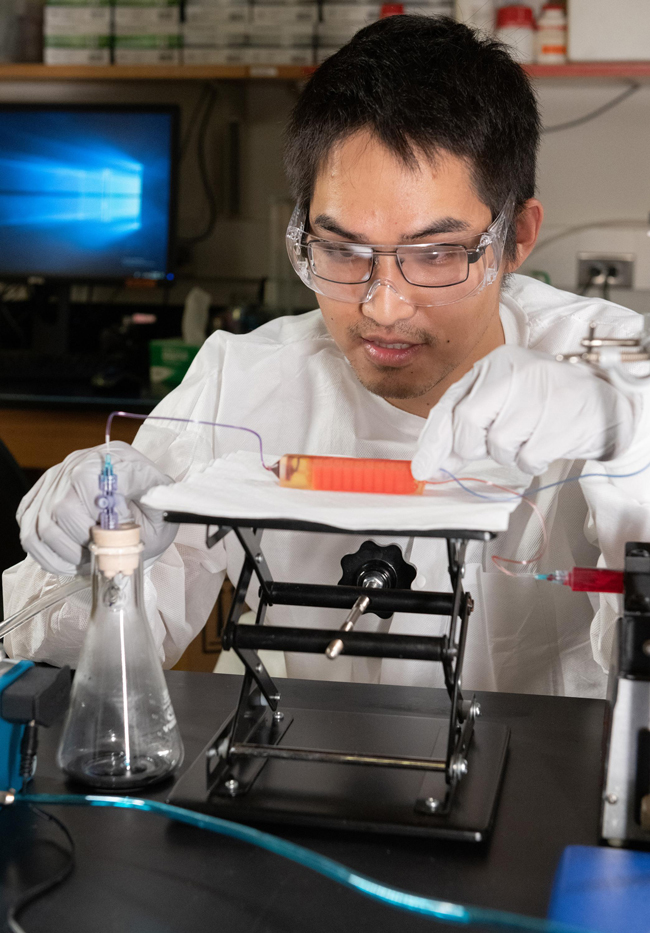

Image: A novel three-dimensionally-printed cell trap captures white blood cells and eliminates red cells from a whole blood sample while isolating circulating tumor cells (CTCs) (Photo courtesy of Allison Carter, Georgia Institute of Technology).
To improve this stituation, investigators at the Georgia Institute of Technology (Atlanta, USA) used three-dimensional printing to create a solid device that combined immunoaffinity-based microfluidic cell capture and a commercial membrane filter for negative enrichment of CTCs directly from whole blood.
The printed device comprised stacked layers of chemically functionalized microfluidic channels, which captured millions of white blood cells (WBCs) in parallel without becoming saturated. The leuko-depleted blood was post-filtered through a three micron-pore size membrane filter to eliminate red blood cells. This hybrid negative enrichment approach facilitated direct extraction of viable CTCs from the chip on a membrane filter for downstream analysis.
Evaluation of the performance of the device by immunofluorescence imaging of the enriched cell fraction revealed greater than 90% tumor cell recovery from simulated samples in which normal blood was spiked with prostate, breast, or ovarian cancer cells.
The investigators demonstrated the feasibility of the approach for processing clinical samples by isolating prostate cancer CTCs directly from a 10-ml whole blood sample.
“Isolating circulating tumor cells from whole blood samples has been a challenge because we are looking for a handful of cancer cells mixed with billions of normal red and white blood cells,” said senior author Dr. A. Fatih Sarioglu, assistant professor of electrical and computer engineering at the Georgia Institute of Technology. “With this device, we can process a clinically relevant volume of blood by capturing nearly all of the white blood cells and then filtering out the red blood cells by size. That leaves us with undamaged tumor cells that can be sequenced to determine the specific cancer type and the unique characteristics of each patient’s tumor.”
“We expect that this will really be an enabling tool for clinicians,” said Dr. Sarioglu. “In our lab, the mindset is always toward translating our research by making the device simple enough to be used in hospitals, clinics, and other facilities that will help diagnose disease in patients.”
The CTC filter method was described in the September 20, 2019, online edition of the journal Lab on a Chip.
Related Links:
Georgia Institute of Technology