Proteomics Approach Used for Analyses of Breast Cancer
By LabMedica International staff writers
Posted on 31 Jul 2019
Accurate classification of breast tumors is vital for patient management decisions and enables more precise cancer treatment. Despite the progress achieved in early cancer diagnosis and therapy, many patients develop the deadly disease.Posted on 31 Jul 2019
The search for better tumor classifiers significantly concentrates on the application of omics approaches, which are able to analyze thousands of gene sequences, gene transcripts, or proteins in a single study. The biochemical effector molecules in cells are proteins, and their direct measurement is, therefore, in principle preferable over the inference of protein quantities from transcript measurements.
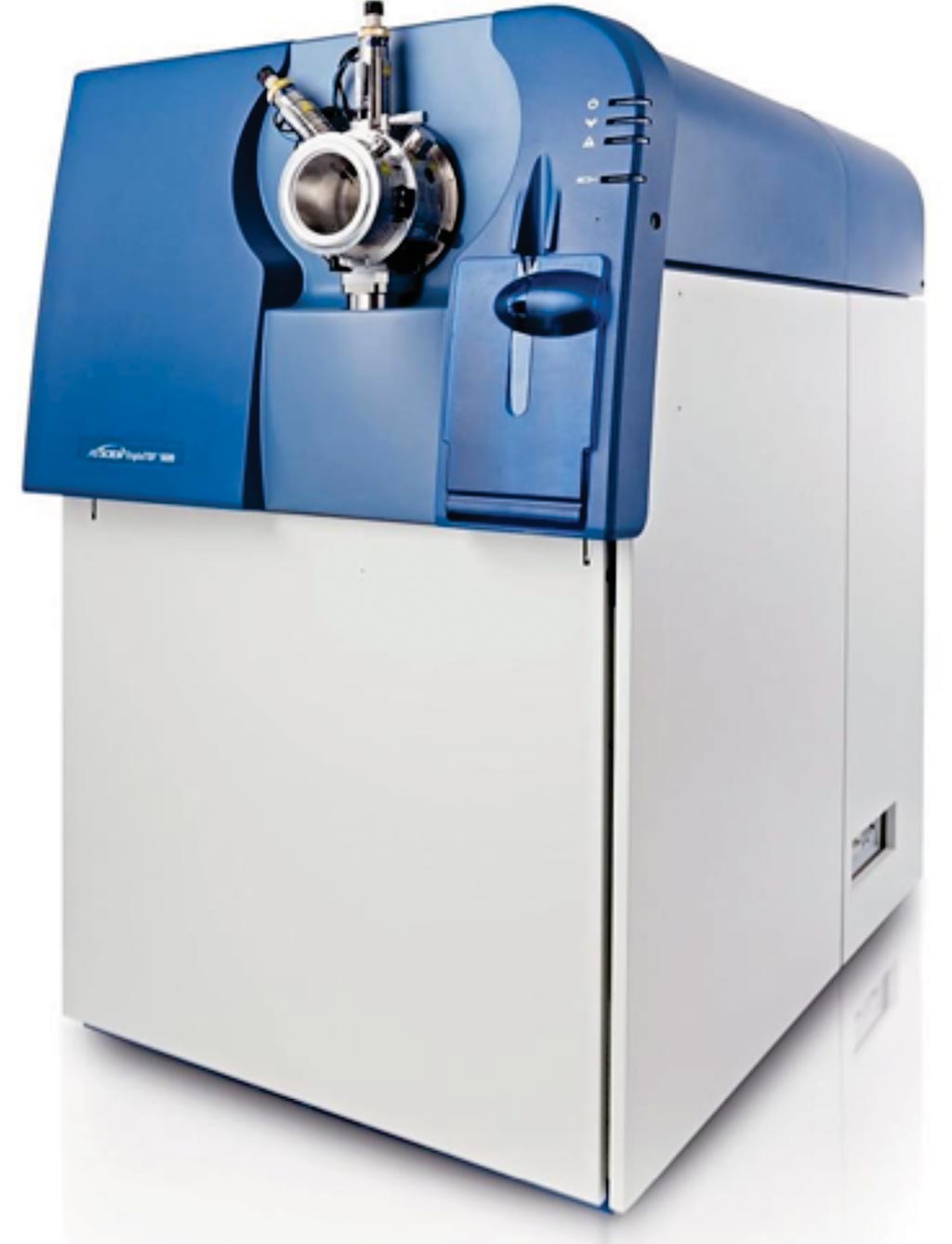

Image: The Triple TOF 5600+ mass spectrometer (Photo courtesy of SCIEX).
A team of scientist collaborating with Masaryk University (Brno, Czech Republic) compared the classification of breast cancer tissues based on proteotypes obtained using a novel next-generation proteomics approach, sequential windowed acquisition of all theoretical fragment ion spectra mass spectrometry (SWATH-MS), with clinically used subtypes classified by immunohistological markers and grade.
Breast cancer tissue samples were frozen in liquid nitrogen within 20 minutes of surgical removal and stored at −180 °C in the tissue bank. A set of 96 preoperatively untreated women’s breast carcinomas of 11-20 mm maximum diameter (pT1c) were selected. The team developed a spectral library based on samples from five classical breast cancer subtypes: luminal A, luminal B HER2-, luminal B HER2+, HER2 enriched, and triple-negative. They then ran the 96 samples using the SWATH DIA approach, collecting quantitative data in every sample for a set of 2,842 proteins.
MS/MS datasets for spectral library generation were acquired on a TripleTOF 5600+ mass spectrometer interfaced to an Eksigent Ekspert nanoLC 400 system. Total cellular RNA was extracted and TP53 mRNA from tumor tissue was amplified using the SuperScript III One Step RT-PCR System with Platinum Taq High Fidelity and sequenced using the ABI PRISM BigDye Terminator v 3.1 Cycle Sequencing Kit on an ABI 3130 genetic analyzer.
The team found their proteomic profiles largely recapitulated the five traditional subtypes, with the highest intra-group correlations seen within the luminal A samples and the luminal B HER2- and luminal B HER2+ subtypes. The proteomic profiles also identified a high correlation between some members of the luminal B HER2+ and HER2-enriched subtypes as well as higher levels of tumor heterogeneity at the protein level within the triple-negative subtype. They also identified from their data a set of three key proteins: type II inositol 3,4-bisphosphate 4-phosphatase (INPP4B); cyclin-dependent kinase 1 (CDK1); and receptor tyrosine-protein kinase erbB-2 (ERBB2), that was able to correctly assign 84% of the 96 tumor samples to their conventional subtype.
Ruedi Aebersold, PhD, a professor and a senior author of the study said, “Essentially what we tried to show is that with a technique that is very fast, relatively cheap, and quite simple, we can recapitulate conventional classification information in a way that has really not been possible in proteomics before, calling it a step towards the "commoditizing or democratization of proteomics.” The study was published on July 16, 2019, in the journal Cell Reports.
Related Links:
Masaryk University