Microbiome Study Links Gut Bacteria to Specific Diseases
|
By LabMedica International staff writers Posted on 01 Jul 2019 |
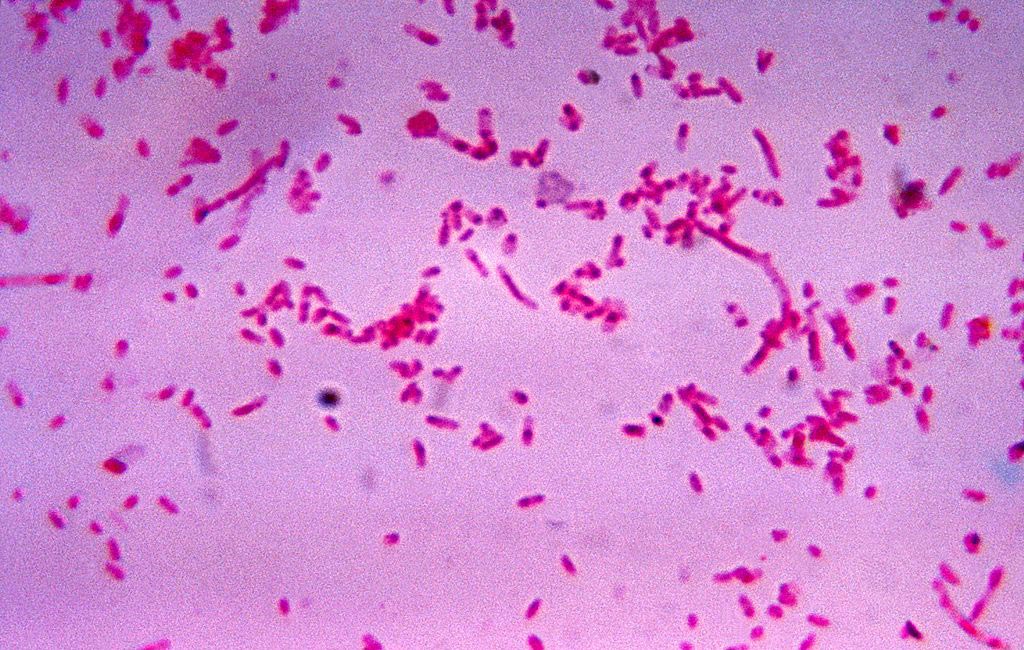
Image: Fusobacterium novum in liquid culture. Fusobacterium flourishes in colon cancer cells, and is often also associated with ulcerative colitis (Photo courtesy of the CDC).
The mix of bacteria comprising the human gut microbiome is linked to the development of specific diseases and syndromes including inflammatory bowel disease, primary sclerosing cholangitis, and chronic depression.
Inflammatory bowel disease (IBD) includes several conditions characterized by chronic inflammation of the intestinal tract. Among these conditions are ulcerative colitis and Crohn's disease. Primary sclerosing cholangitis (PSC) is a long-term progressive disease of the liver and gallbladder characterized by inflammation and scarring of the bile ducts, which normally allow bile to drain from the gallbladder. Affected individuals may have no symptoms or may experience signs and symptoms of liver disease such as yellow discoloration of the skin and eyes, itching, and abdominal pain.
Previous work by investigators at The Flanders Institute for Biotechnology (Leuven, Belgium) had characterized the populations of normal, health-associated gut microbiota. In the current study, they searched for population mixes that could be linked to specific diseases or syndromes.
For this study, the investigators revisited a disease association microbiome data set comprising 106 patients with PSC and/or IBD. They reported an increased prevalence of a low cell count Bacteroides 2 enterotype (B2 enterotype) across the pathologies studied, with microbial loads correlating inversely with intestinal and systemic inflammation markers. Quantitative analyses enabled them to differentiate between taxa associated with either intestinal inflammation severity (Fusobacterium) or cholangitis/biliary obstruction (Enterococcus) among previously suggested PSC marker genera. They identified and validated a near-exclusion pattern between the inflammation-associated Fusobacterium and Veillonella genera, with Fusobacterium detection being restricted to Crohn’s disease and patients with PSC–Crohn’s disease.
Senior author Dr. Jeroen Raes, professor of microbiology and immunology at The Flanders Institute for Biotechnology, said, "Over the years, many research groups worldwide have attempted to describe microbiota alterations associated with diseases. Especially IBD is a hot topic in microbiome research. Our study differs from these previous attempts on three fronts. First, we compared the microbiota of patients with profiles from healthy volunteers from our Flemish Gut Flora Project catalog of over 3,000 microbiomes. Second, in our analyses, we did not only look at the percentages of different bacteria present in the stool samples, but also used a new technique to quantify their abundances. Third, we corrected our results for factors such as loose stools, often symptomatic in the diseases studied, but affecting the outcome of microbiome analyses."
"There appears to be a large overlap in microbiome alterations observed across different patient groups," said Dr. Raes. "We detected the B2 enterotype in around 26% of depressed individuals. While the gut microbiota has been shown to play a role in disease development in, for example, ulcerative colitis and Crohn's disease, this is far less clear for depression. However, we will explore the association between the B2 enterotype and depression in more detail in future studies.
The study was published in the June 17, 2019, online edition of the journal Nature Microbiology.
Related Links:
The Flanders Institute for Biotechnology
Inflammatory bowel disease (IBD) includes several conditions characterized by chronic inflammation of the intestinal tract. Among these conditions are ulcerative colitis and Crohn's disease. Primary sclerosing cholangitis (PSC) is a long-term progressive disease of the liver and gallbladder characterized by inflammation and scarring of the bile ducts, which normally allow bile to drain from the gallbladder. Affected individuals may have no symptoms or may experience signs and symptoms of liver disease such as yellow discoloration of the skin and eyes, itching, and abdominal pain.
Previous work by investigators at The Flanders Institute for Biotechnology (Leuven, Belgium) had characterized the populations of normal, health-associated gut microbiota. In the current study, they searched for population mixes that could be linked to specific diseases or syndromes.
For this study, the investigators revisited a disease association microbiome data set comprising 106 patients with PSC and/or IBD. They reported an increased prevalence of a low cell count Bacteroides 2 enterotype (B2 enterotype) across the pathologies studied, with microbial loads correlating inversely with intestinal and systemic inflammation markers. Quantitative analyses enabled them to differentiate between taxa associated with either intestinal inflammation severity (Fusobacterium) or cholangitis/biliary obstruction (Enterococcus) among previously suggested PSC marker genera. They identified and validated a near-exclusion pattern between the inflammation-associated Fusobacterium and Veillonella genera, with Fusobacterium detection being restricted to Crohn’s disease and patients with PSC–Crohn’s disease.
Senior author Dr. Jeroen Raes, professor of microbiology and immunology at The Flanders Institute for Biotechnology, said, "Over the years, many research groups worldwide have attempted to describe microbiota alterations associated with diseases. Especially IBD is a hot topic in microbiome research. Our study differs from these previous attempts on three fronts. First, we compared the microbiota of patients with profiles from healthy volunteers from our Flemish Gut Flora Project catalog of over 3,000 microbiomes. Second, in our analyses, we did not only look at the percentages of different bacteria present in the stool samples, but also used a new technique to quantify their abundances. Third, we corrected our results for factors such as loose stools, often symptomatic in the diseases studied, but affecting the outcome of microbiome analyses."
"There appears to be a large overlap in microbiome alterations observed across different patient groups," said Dr. Raes. "We detected the B2 enterotype in around 26% of depressed individuals. While the gut microbiota has been shown to play a role in disease development in, for example, ulcerative colitis and Crohn's disease, this is far less clear for depression. However, we will explore the association between the B2 enterotype and depression in more detail in future studies.
The study was published in the June 17, 2019, online edition of the journal Nature Microbiology.
Related Links:
The Flanders Institute for Biotechnology
Latest Microbiology News
- Enhanced Rapid Syndromic Molecular Diagnostic Solution Detects Broad Range of Infectious Diseases
- Clinical Decision Support Software a Game-Changer in Antimicrobial Resistance Battle
- New CE-Marked Hepatitis Assays to Help Diagnose Infections Earlier
- 1 Hour, Direct-From-Blood Multiplex PCR Test Identifies 95% of Sepsis-Causing Pathogens
- Mouth Bacteria Test Could Predict Colon Cancer Progression
- Unique Metabolic Signature Could Enable Sepsis Diagnosis within One Hour of Blood Collection
- Groundbreaking Diagnostic Platform Provides AST Results With Unprecedented Speed
- Simple Blood Test Combined With Personalized Risk Model Improves Sepsis Diagnosis
- Blood Analysis Predicts Sepsis and Organ Failure in Children
- TB Blood Test Could Detect Millions of Silent Spreaders
- New Blood Test Cuts Diagnosis Time for Nontuberculous Mycobacteria Infections from Months to Hours
- New Tuberculosis Test to Expand Testing Access in Low- and Middle-Income Countries
- Rapid Test Diagnoses Tropical Disease within Hours for Faster Antibiotics Treatment
- Rapid Molecular Testing Enables Faster, More Targeted Antibiotic Treatment for Pneumonia
- Rapid AST Platform Provides Targeted Therapeutic Results Days Faster Than Current Standard of Care
- New Analysis Method Detects Pathogens in Blood Faster and More Accurately by Melting DNA
Channels
Clinical Chemistry
view channel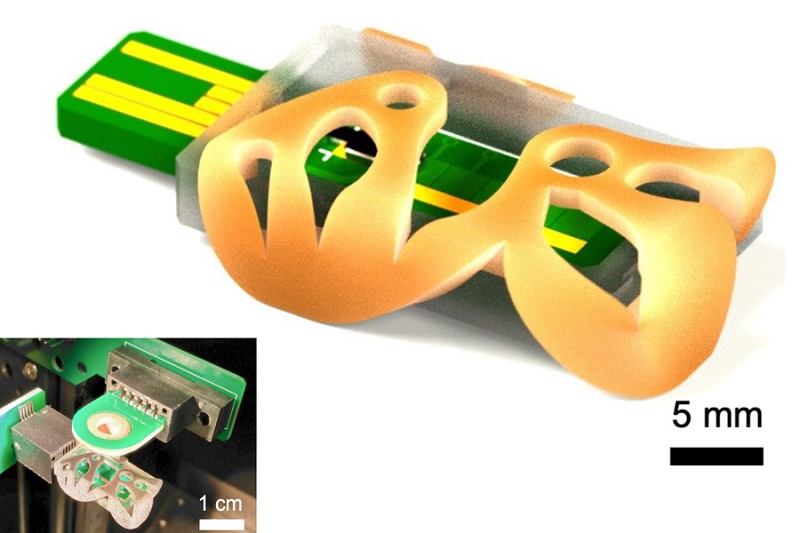
3D Printed Point-Of-Care Mass Spectrometer Outperforms State-Of-The-Art Models
Mass spectrometry is a precise technique for identifying the chemical components of a sample and has significant potential for monitoring chronic illness health states, such as measuring hormone levels... Read more.jpg)
POC Biomedical Test Spins Water Droplet Using Sound Waves for Cancer Detection
Exosomes, tiny cellular bioparticles carrying a specific set of proteins, lipids, and genetic materials, play a crucial role in cell communication and hold promise for non-invasive diagnostics.... Read more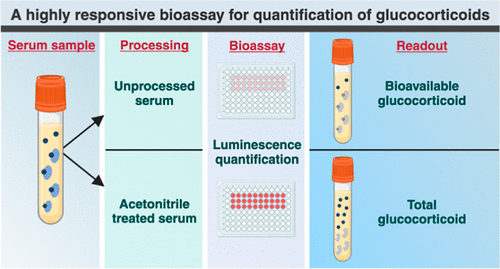
Highly Reliable Cell-Based Assay Enables Accurate Diagnosis of Endocrine Diseases
The conventional methods for measuring free cortisol, the body's stress hormone, from blood or saliva are quite demanding and require sample processing. The most common method, therefore, involves collecting... Read moreMolecular Diagnostics
view channel
Urine Test to Revolutionize Lyme Disease Testing
Lyme disease is the most common animal-to-human transmitted disease in the United States, with around 476,000 people diagnosed and treated annually, and its incidence has been increasing.... Read more
Simple Blood Test Could Enable First Quantitative Assessments for Future Cerebrovascular Disease
Cerebral small vessel disease is a common cause of stroke and cognitive decline, particularly in the elderly. Presently, assessing the risk for cerebral vascular diseases involves using a mix of diagnostic... Read more
New Genetic Testing Procedure Combined With Ultrasound Detects High Cardiovascular Risk
A key interest area in cardiovascular research today is the impact of clonal hematopoiesis on cardiovascular diseases. Clonal hematopoiesis results from mutations in hematopoietic stem cells and may lead... Read moreHematology
view channel
Next Generation Instrument Screens for Hemoglobin Disorders in Newborns
Hemoglobinopathies, the most widespread inherited conditions globally, affect about 7% of the population as carriers, with 2.7% of newborns being born with these conditions. The spectrum of clinical manifestations... Read more
First 4-in-1 Nucleic Acid Test for Arbovirus Screening to Reduce Risk of Transfusion-Transmitted Infections
Arboviruses represent an emerging global health threat, exacerbated by climate change and increased international travel that is facilitating their spread across new regions. Chikungunya, dengue, West... Read more
POC Finger-Prick Blood Test Determines Risk of Neutropenic Sepsis in Patients Undergoing Chemotherapy
Neutropenia, a decrease in neutrophils (a type of white blood cell crucial for fighting infections), is a frequent side effect of certain cancer treatments. This condition elevates the risk of infections,... Read more
First Affordable and Rapid Test for Beta Thalassemia Demonstrates 99% Diagnostic Accuracy
Hemoglobin disorders rank as some of the most prevalent monogenic diseases globally. Among various hemoglobin disorders, beta thalassemia, a hereditary blood disorder, affects about 1.5% of the world's... Read moreImmunology
view channel
Diagnostic Blood Test for Cellular Rejection after Organ Transplant Could Replace Surgical Biopsies
Transplanted organs constantly face the risk of being rejected by the recipient's immune system which differentiates self from non-self using T cells and B cells. T cells are commonly associated with acute... Read more
AI Tool Precisely Matches Cancer Drugs to Patients Using Information from Each Tumor Cell
Current strategies for matching cancer patients with specific treatments often depend on bulk sequencing of tumor DNA and RNA, which provides an average profile from all cells within a tumor sample.... Read more
Genetic Testing Combined With Personalized Drug Screening On Tumor Samples to Revolutionize Cancer Treatment
Cancer treatment typically adheres to a standard of care—established, statistically validated regimens that are effective for the majority of patients. However, the disease’s inherent variability means... Read morePathology
view channel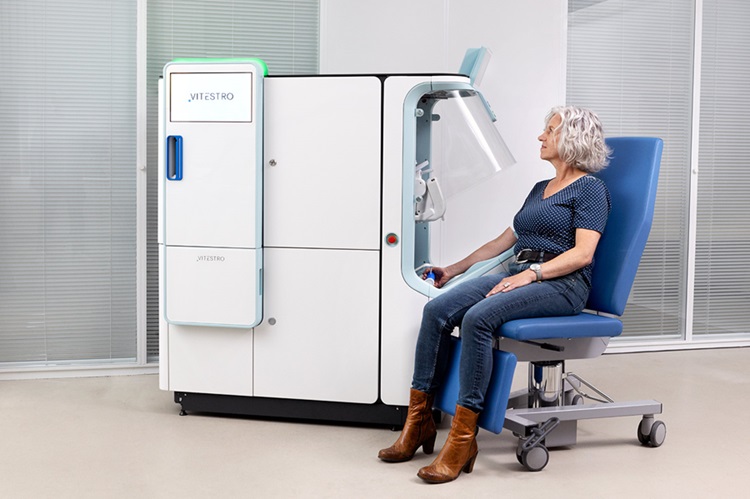
Robotic Blood Drawing Device to Revolutionize Sample Collection for Diagnostic Testing
Blood drawing is performed billions of times each year worldwide, playing a critical role in diagnostic procedures. Despite its importance, clinical laboratories are dealing with significant staff shortages,... Read more.jpg)
Use of DICOM Images for Pathology Diagnostics Marks Significant Step towards Standardization
Digital pathology is rapidly becoming a key aspect of modern healthcare, transforming the practice of pathology as laboratories worldwide adopt this advanced technology. Digital pathology systems allow... Read more
First of Its Kind Universal Tool to Revolutionize Sample Collection for Diagnostic Tests
The COVID pandemic has dramatically reshaped the perception of diagnostics. Post the pandemic, a groundbreaking device that combines sample collection and processing into a single, easy-to-use disposable... Read moreTechnology
view channel
New Diagnostic System Achieves PCR Testing Accuracy
While PCR tests are the gold standard of accuracy for virology testing, they come with limitations such as complexity, the need for skilled lab operators, and longer result times. They also require complex... Read more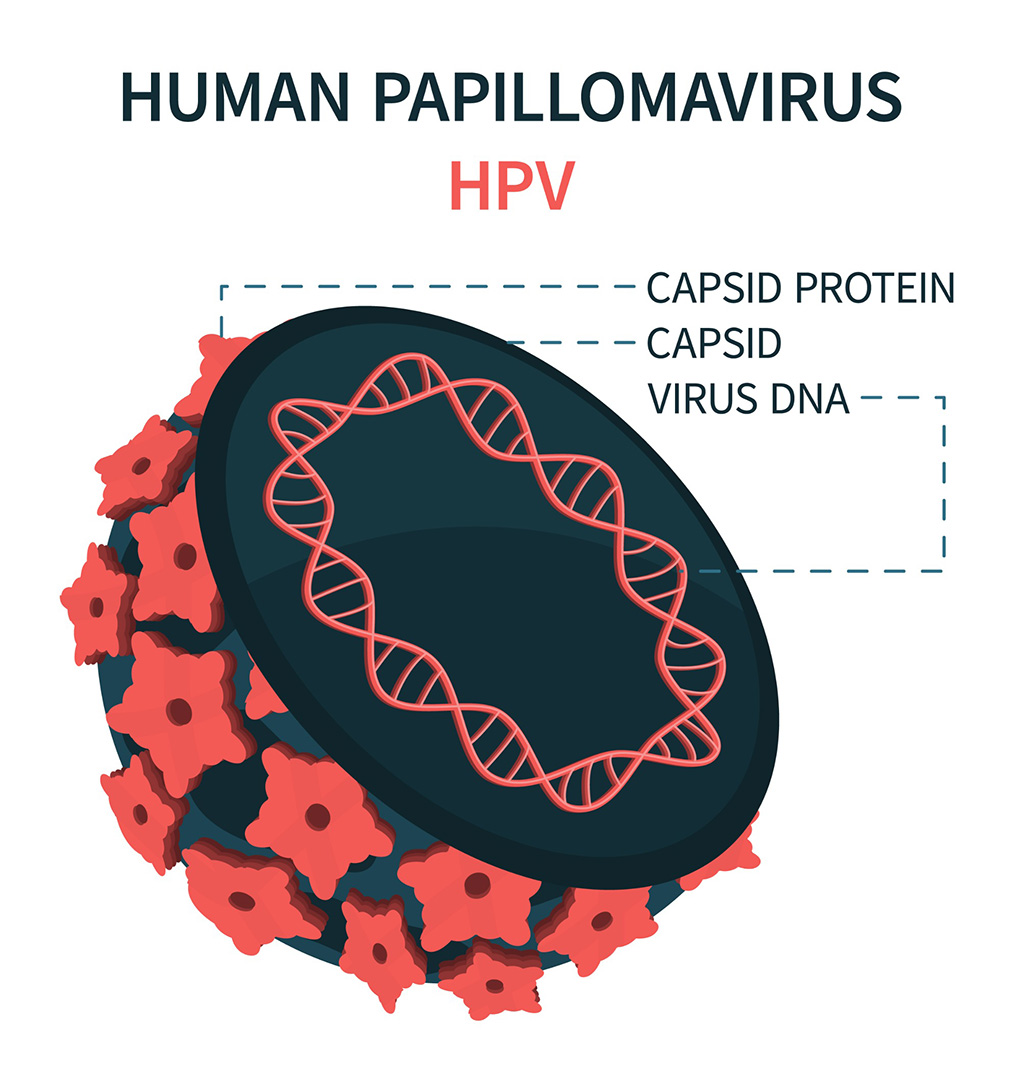
DNA Biosensor Enables Early Diagnosis of Cervical Cancer
Molybdenum disulfide (MoS2), recognized for its potential to form two-dimensional nanosheets like graphene, is a material that's increasingly catching the eye of the scientific community.... Read more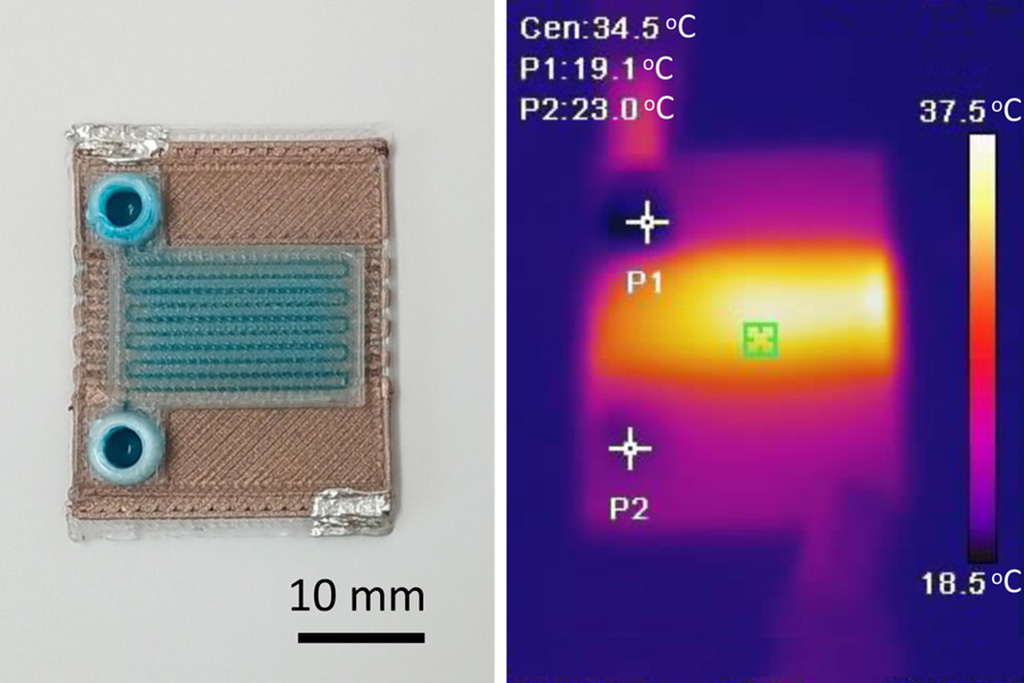
Self-Heating Microfluidic Devices Can Detect Diseases in Tiny Blood or Fluid Samples
Microfluidics, which are miniature devices that control the flow of liquids and facilitate chemical reactions, play a key role in disease detection from small samples of blood or other fluids.... Read more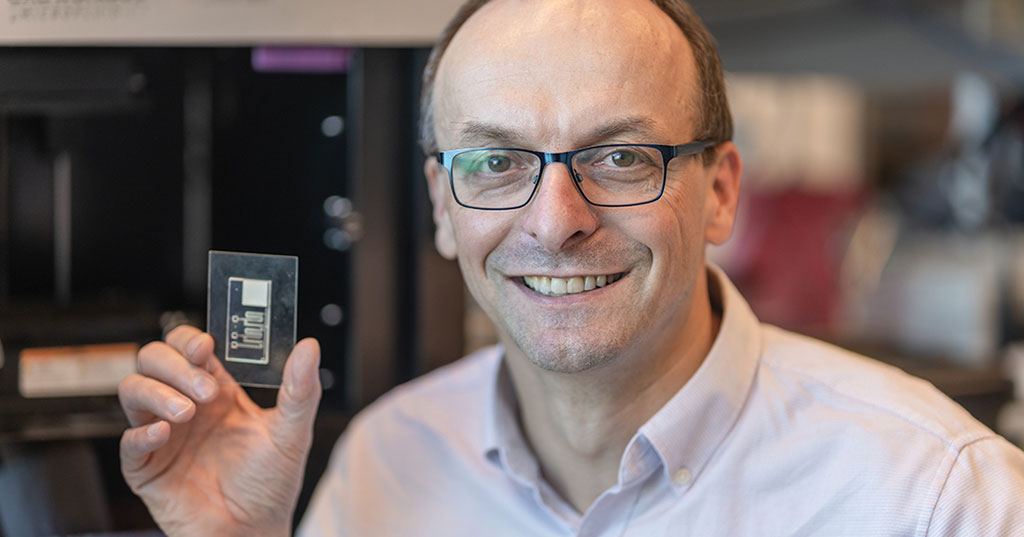
Breakthrough in Diagnostic Technology Could Make On-The-Spot Testing Widely Accessible
Home testing gained significant importance during the COVID-19 pandemic, yet the availability of rapid tests is limited, and most of them can only drive one liquid across the strip, leading to continued... Read moreIndustry
view channel_1.jpg)
Thermo Fisher and Bio-Techne Enter Into Strategic Distribution Agreement for Europe
Thermo Fisher Scientific (Waltham, MA USA) has entered into a strategic distribution agreement with Bio-Techne Corporation (Minneapolis, MN, USA), resulting in a significant collaboration between two industry... Read more
ECCMID Congress Name Changes to ESCMID Global
Over the last few years, the European Society of Clinical Microbiology and Infectious Diseases (ESCMID, Basel, Switzerland) has evolved remarkably. The society is now stronger and broader than ever before... Read more