Urinary Cell-Free DNA Detects Urothelial Carcinoma
By LabMedica International staff writers
Posted on 08 Jan 2020
Most bladder cancers start in the innermost lining of the bladder, which is called the urothelium or transitional epithelium. As the cancer grows into or through the other layers in the bladder wall, it has a higher stage, becomes more advanced, and can be harder to treat.Posted on 08 Jan 2020
Urothelial carcinoma, also known as transitional cell carcinoma (TCC), is by far the most common type of bladder cancer. Current noninvasive assays for urothelial carcinoma (UC) lack clinical sensitivity and specificity. Given the utility of plasma cell-free DNA (cfDNA) bio-markers, the development of urinary cfDNA biomarkers may improve the diagnostic sensitivity.
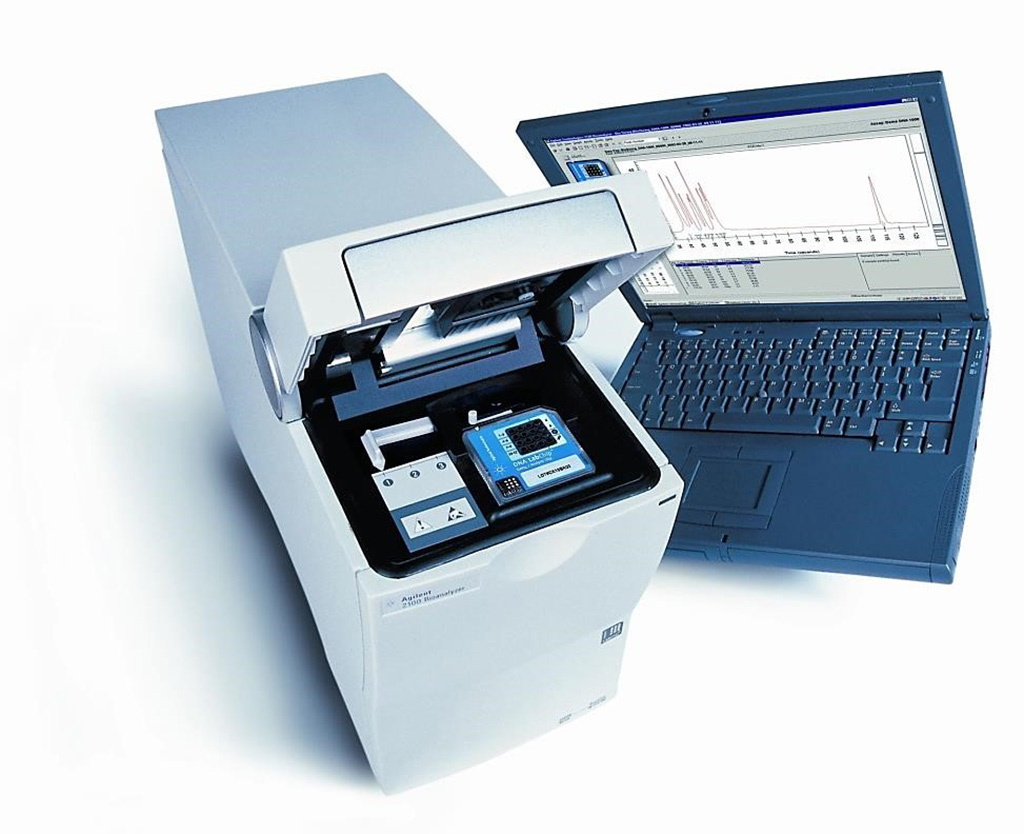

Image: The Agilent Technologies 2100 Bioanalyzer system is an established automated electrophoresis tool for the sample quality control of biomolecules (Photo courtesy of Laboratory Controls LLC)
Urologists at the Beijing Institute of Genomics (Beijing, China) and their colleagues assessed copy number alterations (CNAs) by shallow genome-wide sequencing of urinary cfDNA in 95 cancer-free individuals and 65 patients with UC, 58 with kidney cancer, and 45 with prostate cancer. They used a support vector machine to develop a diagnostic classifier based on CNA profiles to detect UC (UCdetector). A Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA) was used to profile the length distribution of cfDNA isolated from the UC patients.
The model was further validated in an independent cohort (52 patients). Genome sequencing data of tumor specimens from 90 upper tract urothelial cancers (UTUCs) and CNA data for 410 urothelial carcinomas of bladder (UCBs) from The Cancer Genome Atlas were used to validate the classifier. Genome sequencing data for urine sediment from 32 patients with UC were compared with cfDNA. To monitor the treatment efficacy, the team collected cfDNA from seven post-treatment patients.
The investigators reported that urinary cfDNA was a more sensitive alternative to urinary sediment. The UCdetector could detect UC at a median clinical sensitivity of 86.5% and specificity of 94.7%. UCdetector performed well in an independent validation data set. Notably, the CNA features selected by UCdetector were specific markers for both UTUC and UCB. Moreover, CNA changes in cfDNA were consistent with the treatment effects. Meanwhile, the same strategy could localize genitourinary cancers to tissue of origin in 70.1% of patients.
The authors concluded that their findings underscore the potential utility of urinary cfDNA CNA profiles as a basis for non-invasive UC detection and surveillance. The study was published in the December 2019 issue of the journal Clinical Chemistry.
Related Links:
Beijing Institute of Genomics
Agilent Technologies