Cancer-Associated Circular RNAs Catalogued Across Tumors
By LabMedica International staff writers
Posted on 18 Feb 2019
Circular RNA is a type of RNA which, unlike the better known linear RNA, forms a covalently closed continuous loop, i.e., in circular RNA the 3' and 5' ends normally present in an RNA molecule have been joined together.Posted on 18 Feb 2019
Circular ribonucleic acids (circRNAs) are an intriguing class of RNA due to their covalently closed structure, high stability, and implicated roles in gene regulation. Scientists have begun to catalogue and have started characterizing circRNAs found across cancer types, including proposed circRNA markers in the urine of prostate cancer patients.
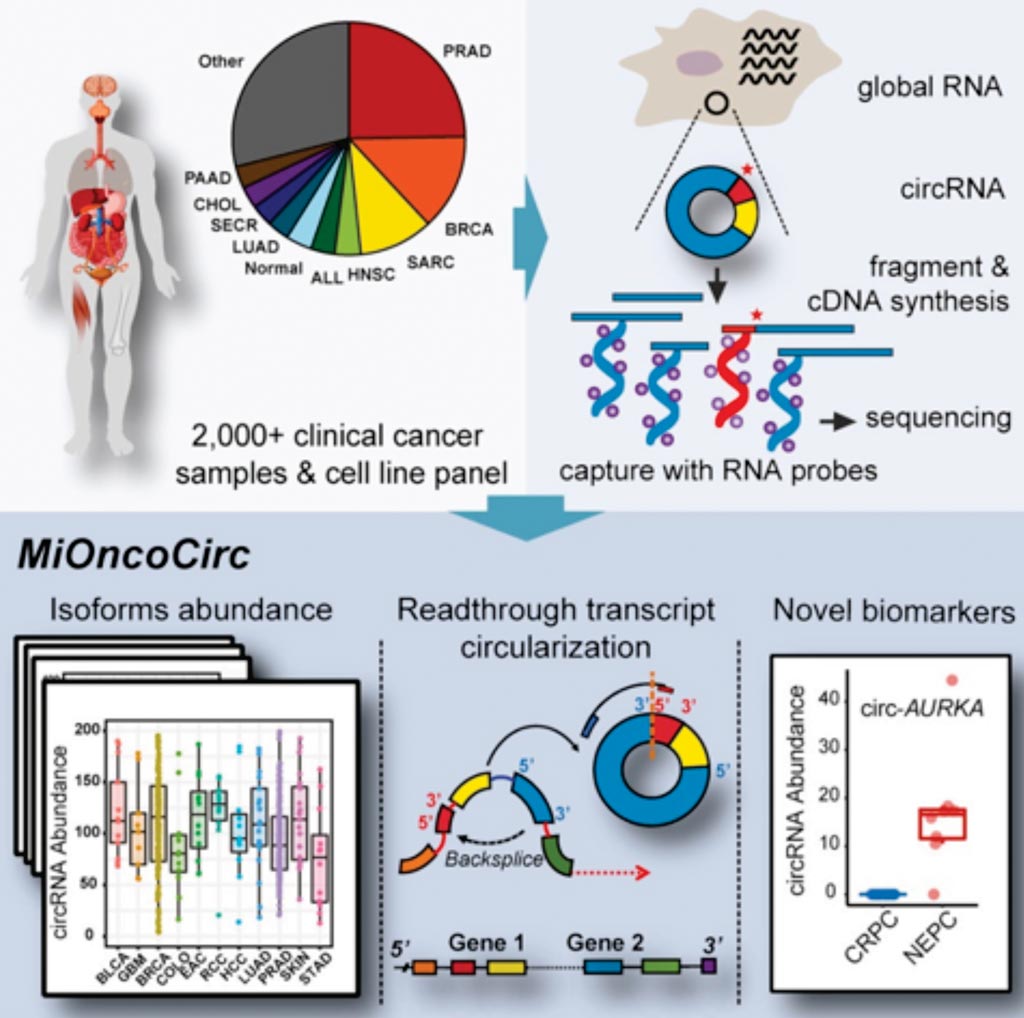
Image: A graphical interpretation of cancer-associated circular RNAs catalogued across tumor types (Photo courtesy of University of Michigan).
A team of scientists from the University of Michigan (Ann Arbor, MI, USA) used exome capture and RNA sequencing to profile circRNAs in more than 800 tumors, cell line, or pooled normal samples assessed for the Michigan Oncology Sequencing Center (MI-ONCOSEQ) program. The analyses led to nearly 129,000 circRNAs in 40 cancer types, which they compared to more than 60,000 circRNAs previously compiled in the CircBase database.
Along with circRNA expression data for more than two dozen matched tumor-normal pairs, the team's cancer-associated circRNA collection known as MiOncoCirc, helped the team focus in on circRNAs associated with metastatic castration-resistant prostate cancer or aggressive neuroendocrine prostate cancer cases, as well as potential circRNA biomarkers found in urine samples from prostate cancer patients in general.
The team used Agilent SureSelect exon capture probes to target nearly 21,000 genes and more than 334,000 exons in cDNA from 868 clinical samples, cell lines, and pooled normal tissue samples, following by sequencing with Illumina technology and identifying candidate circRNAs with an analytical pipeline called CIRCexplorer. When the team compared and validated this method against established Ribo-Zero sequencing and exoribonuclease RNase R-based enrichment approaches in a prostate cancer cell line, it found that the exome capture approach appeared to pick up some circRNAs missed by Ribo-Zero, while maintaining "circular-to-linear ratios" that cannot be gleaned with the RNase R method.
The investigators used MiOncoCirc and identified candidate circRNAs to serve as biomarkers for prostate cancer and were able to detect circRNAs in urine. They further detected a novel class of circular transcripts, termed read-through circRNAs that involved exons originating from different genes. MiOncoCirc will serve as a valuable resource for the development of circRNAs as diagnostic or therapeutic targets across cancer types. The study was published on February 7, 2019, in the journal Cell.
Related Links:
University of Michigan