Molecular Test Developed for Nasopharyngeal Carcinoma
By LabMedica International staff writers
Posted on 12 Jun 2018
The prevalence of nasopharyngeal carcinoma (NPC) is higher in Southeast Asia than other parts of the world, around 35 cases per 100,000. Five-year survival rate of the disease is as high as 95% when it is detected early, but drops to 60% if detected at later stage. Most patients are asymptomatic, and so 80% are diagnosed with advanced disease.Posted on 12 Jun 2018
Liquid biopsies via circulating cell-free DNA (cfDNA) analysis have been shown to be of value in noninvasive monitoring of cancer treatment response and for the detection of cancer recurrence. To extend the application of circulating cell-free DNA to cancer screening, investigators have to face the challenge of developing assays that are sufficiently sensitive for detecting the expectedly low concentrations of circulating tumor DNA in early stages of cancer.
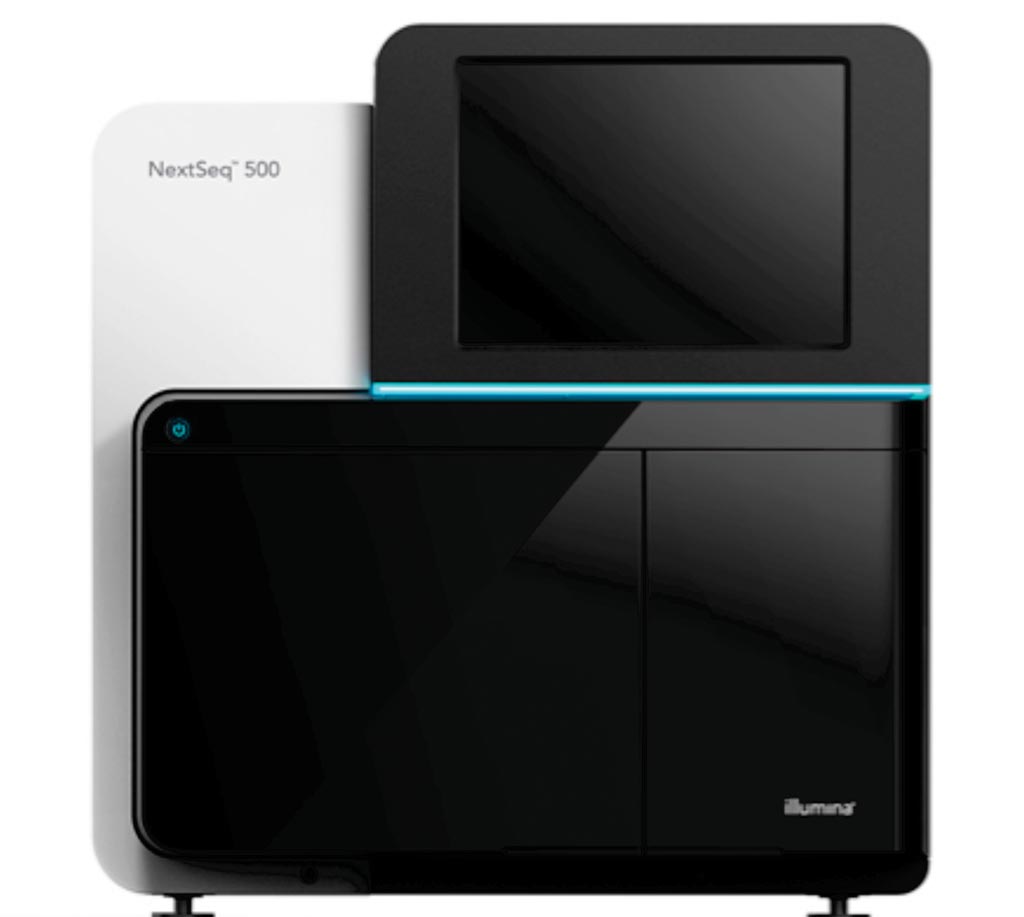

Image: The NextSeq 500 desktop sequencing platform (Photo courtesy of Illumina).
Scientists at The Chinese University of Hong Kong (Sha Tin, Hong Kong) analyzed a target population that was asymptomatic, ethnically Chinese males aged between 40 and 62 years. This group has the highest age-specific incidence of NPC. All of the participants provided a venous blood sample of 20 mL at enrollment. The team used 800 mL of plasma for Epstein - Barr virus (EBV) DNA analysis by a real-time polymerase chain reaction (RT-PCR) assay, which targeted the BamH1-W fragment of the EBV genome.
For enrichment of viral DNA molecules from the plasma DNA samples for subsequent sequencing analysis, target enrichment with EBV capture probes was performed. DNA libraries from five samples were multiplexed in one capture reaction. Equal amounts of DNA libraries for each sample were used. The team also included probes to cover human autosomal regions for reference. The captured autosomal DNA sequences were used for normalization of the viral DNA reads. The multiplexed DNA libraries were sequenced using either the NextSeq 500 or the HiSeq 2500 Sequencing platforms.
The team used a targeted sequencing approach to target the Epstein Barr virus, generating around 70 million mapped reads per sample. Sequencing confirmed that those with cancer had a higher proportion of viral DNA than those without. In addition, they found size differences between the viral DNA in individuals with and without cancer. The circulating DNA has an average size of around 160 to 170 bases because a piece of DNA wrapped around a histone core is around 140 bases, with another 20-base linker region. In the previous liver cancer study, the team demonstrated that circulating DNA (ctDNA) did not have that 20-base linker region. By contrast, viral DNA in cancer-free individuals had not integrated into the genome.
Overall, when they combined the count-based analysis to determine the concentration of viral DNA and the sized-based analysis, the targeted sequencing test had a sensitivity of 97.1%, a specificity of 99.3%, and a PPV of 19.6% an improvement over the PCR assay which had a PPV of 11% and required sampling at two time points. The PCR assay when used at a single time point had a PPV of just 3.1%. However, one advantage of a PCR-based approach is its low cost. In a previous study, for instance, the investigates reported that the cost of testing was just USD 30, as compared to USD 80 and USD 1,000 for endoscopic tests and MRI, respectively. The study was published on May 14, 2018, in the journal Proceedings of the National Academy of the Sciences.
Related Links:
The Chinese University of Hong Kong