Nanopore Method Developed for Genetic Screening
By LabMedica International staff writers
Posted on 13 Apr 2018
Rapid sequencing of short DNA reads may be useful for a wide range of clinical applications including targeted mutation analysis, cancer-panel testing, and aneuploidy screening. Fast, point-of-care preimplantation genetic screening could improve the success of in vitro fertilization procedures by not requiring embryos to be frozen.Posted on 13 Apr 2018
However, the time and skill required for library preparation and sequencing using existing DNA sequencing methods limits their widespread clinical use. Nanopore sequencing technology is one of the fastest growing third generation next-generation sequencing (NGS) technologies. A new method that simplifies and accelerates nanopore-based short-length DNA library preparation and sequencing has been applied to test a panel of normal and aneuploid genomic DNA samples.
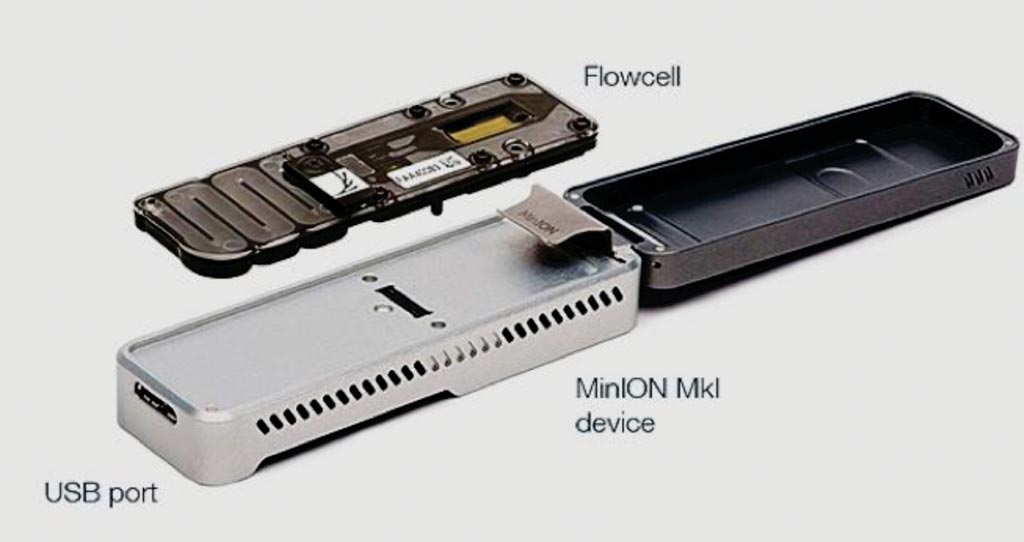

Image: The MinION is the only portable real-time device for DNA and RNA sequencing (Photo courtesy of Oxford Nanopore Technologies).
Scientists at the Columbia University Medical Center (New York, NY, USA) tested the new protocol on nine blinded samples. The nine samples included both diploid and aneuploid samples and one normal reference male sample. Trophoectoderm biopsy samples were taken from fresh, day-five embryos and sent to a reference laboratory for clinical preimplantation genetic screening (PGS) testing.
PGS screening was performed using the VeriSeq PGS assay with excess DNA used to run the MinION-based workflow. After sequencing, the scientists used only reads that were assigned to a barcode and uniquely matched to the reference genome for analysis. More than 70% of reads were assigned to a unique barcode, and of those, between 75% and 95%were uniquely assigned to the reference.
The investigators determined that to detect whole chromosome aneuploidy, 30,000 reads were needed. In the study, their results were concurrent with the VeriSeq PGS test. Of the nine samples, five were abnormal, including a female monosomy 22, female trisomy 19, female trisomy 22 and monosomy X, female trisomy 13 and monosomy 14, a male with an extra X chromosome and trisomy 15, and a female with trisomy 15 and monosomy 18 that was also mosaic for trisomy 6.
S. Zev Williams, MD, PhD, an Associate Professor of Obstetrics and Gynecology, and senior author of the study, said, “One limitation of nanopore sequencing has been its accuracy. However, since aneuploidy detection does not rely on calling point mutations, base-level accuracy is not as important. We want to have an accuracy of over 99%, and to test enough samples to be sure that's what it is will take time. Nonetheless, I think the potential is there.” The study was published in the April 2018 issue of the journal G3: Genes, Genomes, Genetics.
Related Links:
Columbia University Medical Center