Genomic Test Detects Lynch Syndrome in Colorectal Cancer
By LabMedica International staff writers
Posted on 11 Apr 2018
Lynch syndrome (LS) affects approximately 3% of all patients with colorectal cancer (CRC), making it the most common hereditary syndrome that predisposes individuals to develop CRC. Universal tumor screening for LS in CRC is recommended and involves up to six sequential tests.Posted on 11 Apr 2018
Somatic gene testing is performed on stage IV CRCs for treatment determination. The diagnostic workup for patients with CRC could be simplified and improved using a single up-front tumor next-generation sequencing test if it has higher sensitivity and specificity than the current screening protocol.
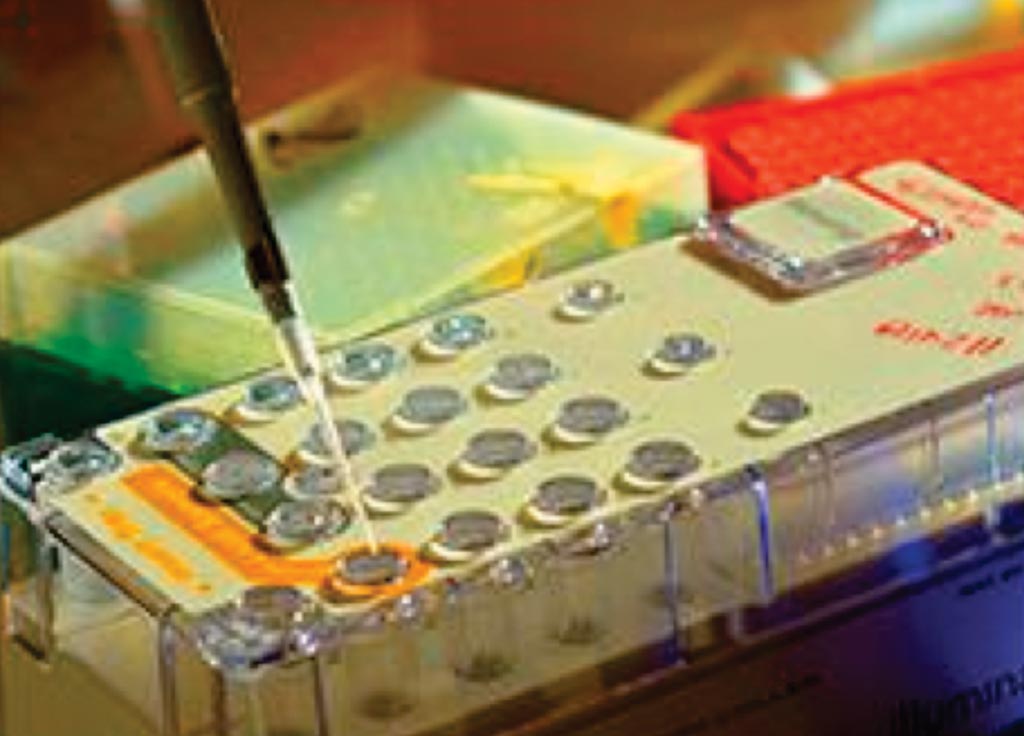

Image: UW-OncoPlex is a diagnostic tool that uses genetic sequencing to look for mutations that cause cancer (Photo courtesy of University of Washington).
Scientists at Ohio State University (Columbus, OH, USA) and their colleagues analyzed tumor samples from 419 CRC patients who participated in the Ohio Colorectal Cancer Prevention Initiative (OCCPI), a statewide study to screen newly diagnosed CRC patients and their biological relatives for Lynch syndrome. All OCCPI study participants had their tumor samples analyzed using the traditional multi-test genetic testing approach and the single, upfront genomic tumor-sequencing test approach in which a single tumor sample was analyzed for multiple mutations simultaneously. A total of 197 of the 419 prospective cases underwent germline genetic testing.
Tumor sequencing was performed using University of Washington (UW)-OncoPlex in the Clinical Laboratory Improvement Amendments–certified laboratory setting (Seattle, WA, USA). UW-OncoPlex is a clinically validated targeted next-generation sequencing (NGS) deep sequencing panel that sequences to 500 × mean depth all exons, introns, and flanking regions of MLH1, MSH2, and MSH6, and all exons of PMS2 in addition to assessing BRAF, KRAS, NRAS, and MSI status among other genes.
The team compared the results from the two screening methods and found that the upfront tumor-sequencing approach was more sensitive and more specific for detecting Lynch syndrome than the old, multiple-test model. Tumor sequencing resulted in a 10% improvement in Lynch syndrome detection rates while also providing important information about treatment options for the patients.
Heather Hampel, MS, CGC, the principal investigator and corresponding author of the study, said, “Testing methods of the past would just point to a suspicion of Lynch syndrome, but they could not confirm the diagnosis without multiple additional tests, which slows down the diagnostic process and adds costs. This new approach points to the exact mutation patients were born with and does so through a single test. The mutation will need to be confirmed using a blood test but this requires a single mutation test, which is less expensive than multi-gene panel testing. The previous method could sometimes require patients to get up to five individual tests before knowing if they had Lynch syndrome.” The study was published on March 29, 2018, in the journal JAMA Oncology.
Related Links:
Ohio State University
University of Washington