Multiplex Immunoassay Developed for Confirmation and Typing of HTLV Infections
By LabMedica International staff writers
Posted on 24 Nov 2021
Human T-Cell Lymphotropic Viruses (HTLV) type 1 and type 2 account for an estimated five to 10 million infections worldwide and are transmitted through breast feeding, sexual contacts and contaminated cellular blood components.Posted on 24 Nov 2021
HTLV-associated syndromes are considered as neglected diseases for which there are no vaccines or therapies available, making it particularly important to ensure the best possible diagnosis to enable proper counseling of infected persons and avoid secondary transmission. The virus can cause adult T-cell leukemia/lymphoma (ATL) and progressive nervous system condition known as HTLV-1-associated myelopathy or tropical spastic paraparesis.
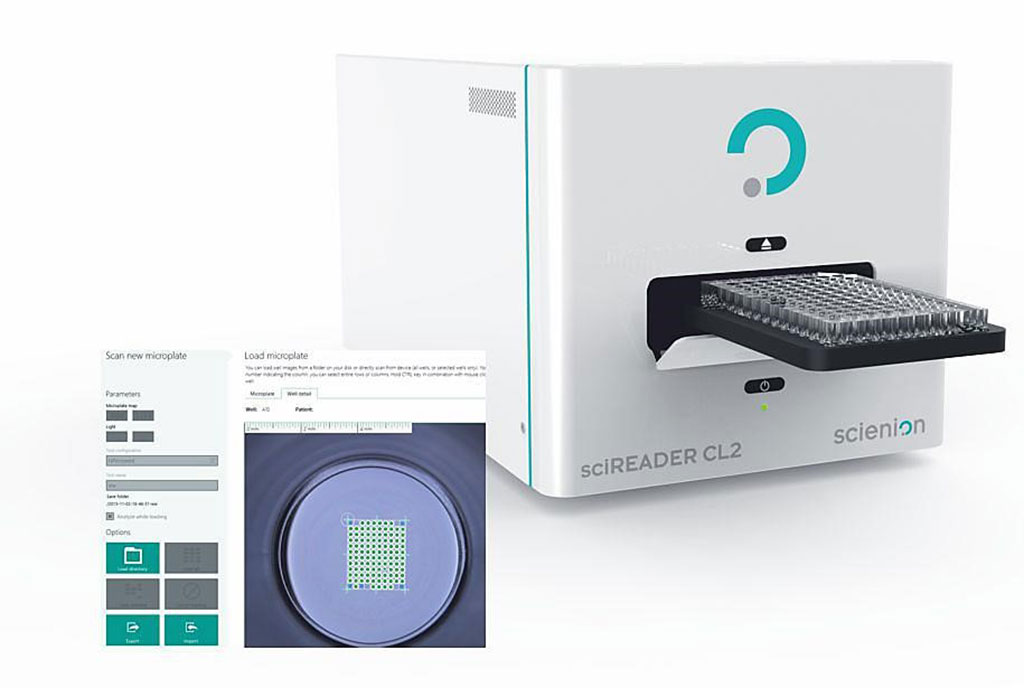

Image: The sciREADER CL2 enables high quality digital colorimetric imaging of various support formats (Photo courtesy of SCIENION)
A large team of scientists from the commercial company InfYnity Biomarkers (Lyon, France) collaborated with several academic institutions to develop and validate a new high-performance multiplex immunoassay for confirmation and discrimination of HTLV-1 and HTLV-2 strains. The team conducted a prospective study with samples from the USA-based cohort HTLV Outcomes Study (HOST) which includes samples from subjects recruited from five major US blood donation centers and 694 frozen serum specimens were obtained from the cohort repository. Included in the study was only the first sample tested of each participant, more specifically, 199 HTLV-positive samples from 199 participants (100 HTLV-1 from 100 patients and 99 HTLV-2 from 99 patients) and 198 negative samples from 198 participants. Other samples were collected from France and Canada.
The multiplex platform was used first as a tool to identify suitable antigens and in a second step for assay development. The Multi-HTLV assay is composed of (i) three confirmatory antigen spots for binding specific HTLV-1 and HTLV-2 antibodies and derived from common-type and type-specific immunodominant epitopes of env GP21, env GP46 and gag P19; and (ii) three discriminatory antigen spots derived from type-specific immunodominant epitopes of HTLV-1 env GP46, HTLV-1 gag P19 and HTLV-2 env GP46.
The Multi-HTLV test was performed very similarly to a standard ELISA assay enabling simultaneous detection of circulating antibodies to a set of selective and validated antigens. Each plate was read and analyzed both visually and using a specific microarray reader sciREADER CL2 (SCIENION, Berlin, Germany) which acquires high-resolution digital images and an integrated software calculates the pixel intensity for each spot.
The investigators reported that the Multi-HTLV assay demonstrated very high performance for confirmation and strain discrimination with 100% sensitivity, 98.1% specificity and 100% of typing accuracy in validation samples. The assay can be interpreted either visually or automatically with a colorimetric image reader and custom algorithm, providing highly reliable results. They observed in 35 of 60 patients a certain dynamic in antibody levels between the initial donation and 10–11 years of follow-up.
The authors concluded that the newly developed Multi-HTLV is very competitive with currently used confirmatory assays and reduces considerably the number of indeterminate results. The multiparametric nature of the assay opens new avenues to study specific serological signatures of each patient, follow the evolution of infection, and explore utility for HTLV disease prognosis. The study was published on November 1, 2021 in the journal PLOS Neglected Tropical Diseases.
Related Links:
InfYnity Biomarkers
SCIENION




.jpg)