Multiplexed Immunoassay System Differentiates Malaria Species
By LabMedica International staff writers
Posted on 03 Oct 2019
Over 40% of world’s population live in malaria-endemic areas, and malaria is found in more than 100 countries in Africa, Latin America, the Caribbean, Southeast Asia, the Eastern Mediterranean, the Western Pacific, and parts of Europe.Posted on 03 Oct 2019
Malaria continues to impose a tremendous burden in terms of global morbidity and mortality, yet even today, a large number of diagnoses are presumptive resulting in lack of or inappropriate treatment. Early diagnosis and prompt, appropriate treatment is essential for improving patient outcomes, preventing overuse of malarial drugs, and minimizing development and spread of anti-malarial drug resistance.
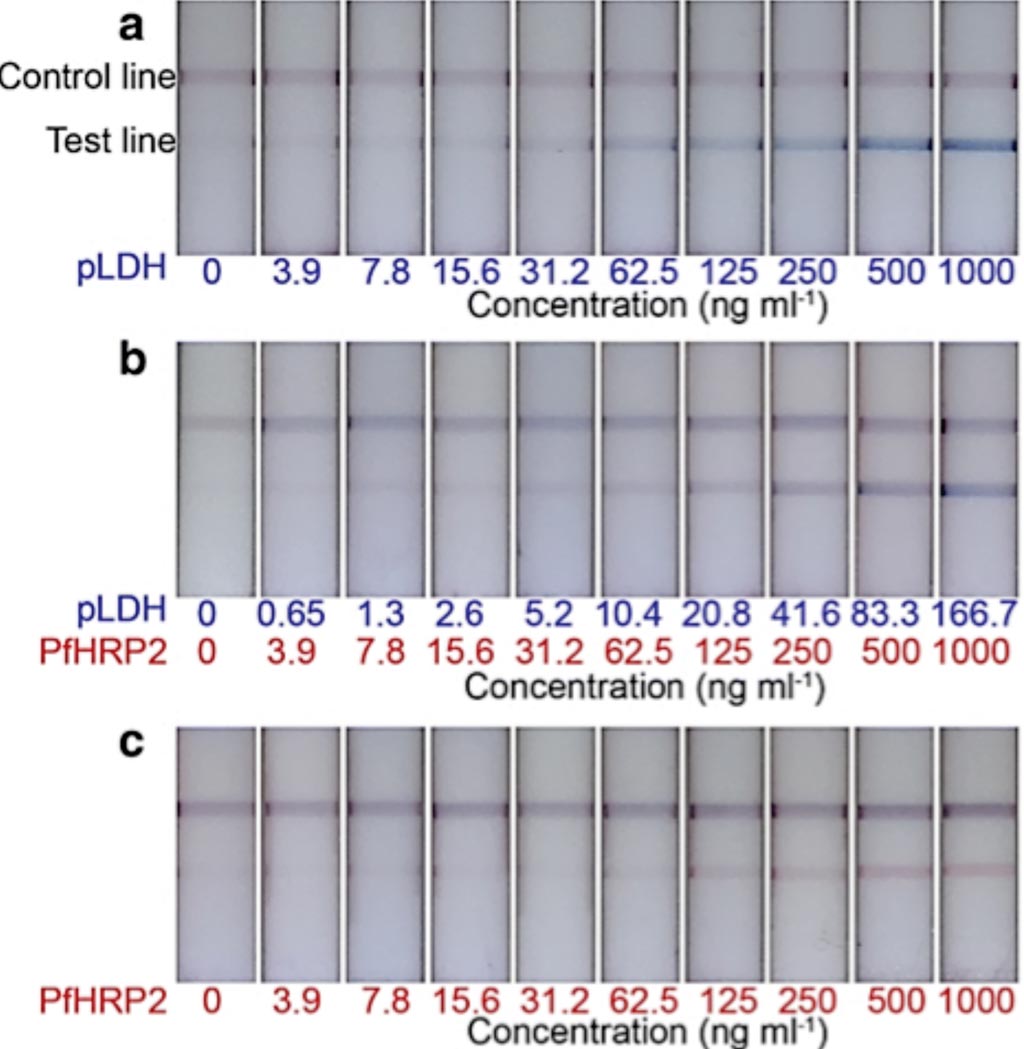
Image: Representative test strips of the two-color LFA. Distinct colors were observed at the test lines, corresponding to a type of antigen, such as a blue test lines for pLDH detection only (Photo courtesy of Cornell University).
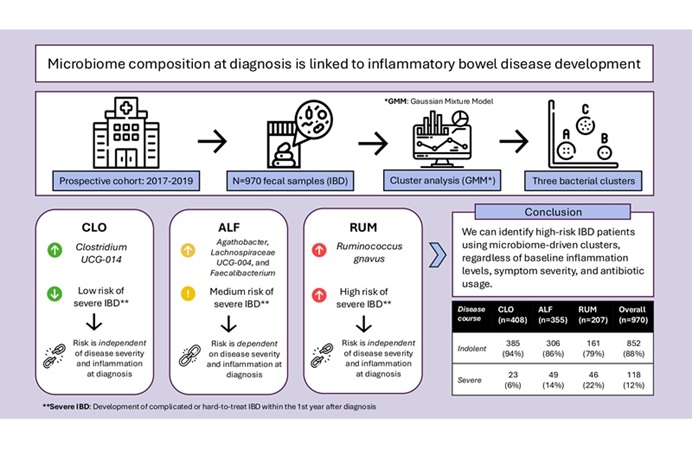
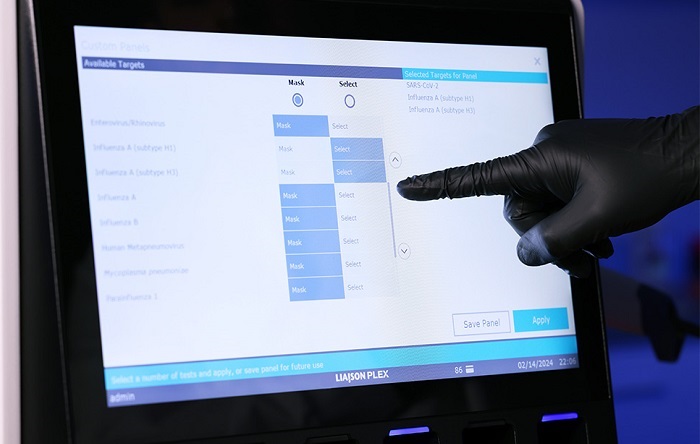
Scientists from Cornell University (Ithaca, NY, USA) and their colleagues developed a two-color lateral flow immunoassay (LFA) system to identify infections by Plasmodium spp. and differentiate Plasmodium falciparum infection from the other three human malaria species (Plasmodium vivax, Plasmodium ovale, Plasmodium malariae). To validate the performance of the assay, 25 clinical research samples in whole blood form were tested: 15 malaria negative samples, five P. falciparum positive samples, and five P. vivax positive samples.
The assay performance was first optimized and evaluated with recombinant malarial proteins spiked in washing buffer at various concentrations from zero to 1,000 ng mL−1. The color profiles developed on the single test line were discriminated and quantified: color types corresponded to malaria protein species; color intensities represented protein concentration levels. The samples were also tested using commercially available ELISA kits: Quantimal pLDH CELISA, and Quantimal Pf-HRP2 CELISA to get quantitative results to evaluate the performance of the LFA strips.
The team reported that the limit of detection (the lowest concentrations of malaria antigens that can be distinguished from blank samples) and the limit of color discrimination (the limit to differentiate pLDH from PfHRP2) were defined for the two-color assay from the spiked buffer test, and the two limits were 31.2 ng mL−1 and 7.8 ng mL−1, respectively. To further validate the efficacy of the assay, the 25 human whole blood frozen samples were tested and successfully validated against ELISA and microscopy results: 15 samples showed malaria negative; five samples showed P. falciparum positive; five samples negative for P. falciparum, but contained other malaria species.
The authors concluded that the assay provides a simple method to quickly identify and differentiate infection by different malarial parasites at the point-of-need and overcome the physical limitations of traditional LFAs, improving the multiplexing potential for simultaneous detection of various biomarkers. The study was published on September 18, 2019, in the Malaria Journal.
Related Links:
Cornell University