Performance of Blood Culture Molecular Panels Evaluated
By LabMedica International staff writers
Posted on 06 Sep 2018
High accuracy of direct identification of pathogens from positive blood culture molecular panels is imperative, particularly for the detection of resistance determinants as it allows for antimicrobial optimization prior to conventional susceptibility testing.Posted on 06 Sep 2018
In general, when patients are suspected of having bacteremia or sepsis, clinicians immediately prescribe a broad-spectrum antibiotic aimed at killing most bacteria known to cause the infection. They start de-escalating use of the drugs when they deem it appropriate.

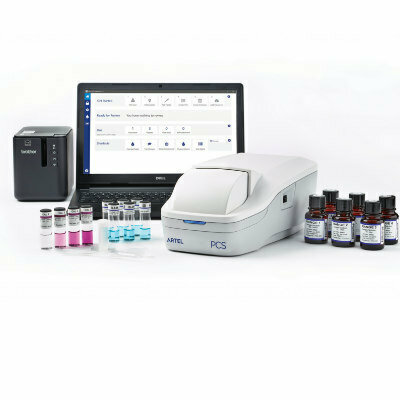
Image: The Verigene Gram-positive blood culture test detects 12 Gram-positive targets and three resistance markers (Photo courtesy of Luminex).
Scientists at the Children’s Hospital Los Angeles (Los Angeles, CA, USA) and their colleagues have provided extensive data of a five-year study since implementation of the Verigene Gram-positive blood culture panel in 2013.
Within five years, 1,636 blood culture bottles positive for a Gram-positive organism were tested on the BC-GP panel. The BC-GP panel identified 1,520 Gram-positive organisms in 1,636 (92.9%) blood cultures tested. For positive blood cultures, they observed 96.4% (806/834) concordance to the species level. Compared with conventional antimicrobial susceptibility testing, the positive percent agreement (PPA) of methicillin-resistant Staphylococcus aureus, SA (MRSA) (50) and methicillin-resistant Staphylococcus epidermidis, SE (MRSE) (365) was 100%.
The mecA gene was detected in two methicillin-susceptible Staphylococcus aureus (MSSA) and one methicillin-susceptible S. epidermidis (MSSE) with a negative percent agreement (NPA) of 99.1% (221/223) and 99.2% (120/121), respectively. The PPA and NPA for vancomycin-resistant Enterococcus faecium (VRE) was 100%. The BC-GP panel demonstrated excellent performance and clinicians can confidently de-escalate antimicrobial therapy in the absence of mecA and vanA/B gene. The panel also identifies coagulase-negative staphylococci that are usually contaminants obtained from a patient's skin at the time of a blood draw, and that can enable clinicians to discontinue antibiotics if the patient is stable and the clinician is comfortable that an infection is not present.
Jennifer Dien Bard, PhD, D(ABMM), F(CCM), an associate professor and co-author of the study, said, “Deescalating to a more targeted antibiotic, such as cefazolin or oxacillin, is efficacious because it optimizes treatment for patients with a methicillin-susceptible Staphylococcus aureus (MSSA. A clinician would switch, in this case, from prescribing vancomycin, a broad-spectrum antibiotic. Traditional susceptibility testing, however, can take longer than is optimal, requiring about 20 to 24 hours to identify the pathogen and an additional 18 to 24 hours to test for susceptibility. That's when the process is straight forward and we don't have to do repeat testing for some reason.” The study was published on August 11, 2018, in the European Journal of Clinical Microbiology & Infectious Diseases.
Related Links:
Children’s Hospital Los Angeles